Scheffler Immo E. Mitochondria
Подождите немного. Документ загружается.

Post - transcriptional mechanisms as well as transcriptional regulation must
be considered in yeast when the regulation of respiratory capacity and assem-
bly of the oxidative phosphorylation system in yeast mitochondria is under
discussion. One can speculate whether such a multitude of mechanisms is
required for fi ne - tuning, for establishing the overall range of the activities
involved, or for a rapid response to rapidly changing environmental conditions.
It will be important to keep these considerations in mind when similar ques-
tions are raised about various mammalian cell types in intact organisms.
4.2.2.3 Regulation of Nuclear Respiratory Genes in Mammalian Cells The
general considerations and the observations made with yeast have encouraged
hypotheses and experiments aimed at similar goals for mammalian cells: (a)
characterization of transcription factors and their targets on the relevant
promoters, which might explain the coordinate expression of the large number
of nuclear genes encoding mitochondrial proteins, and (b) the nuclear –
mitochondrial interactions, which are assumed to take place to make nuclear
gene expression match mitochondrial gene expression or vice versa. A genetic
approach is technically and logistically diffi cult, if not impossible, with diploid
mammalian cells, and no conditions are known in tissue culture which would
induce or repress the biogenesis of mitochondria to permit willful manipula-
tion of the experimental system. A priori one would not expect mammalian
cells in culture to be able to respond like yeast to changing carbon sources,
since they have become adapted in evolution to a very stable environment in
vivo and hence have no need regulate their mitochondria over a short term.
Experiments with Chinese hamster fi broblasts have shown that a large fraction
> 50% of the energy needs of these cells is satisfi ed by glycolysis (115) ; and
even when glucose is deliberately limited in culture, there is no induction of
transcripts of complex II, for example (80) , and only a modest increase in res-
piration. On the other hand, it has become apparent that different tissues and
cell types may have widely differing energy needs and hence may have to
adjust their level of mitochondrial respiration and oxidative phosphorylation
to these needs as part of the process of differentiation. From the study of
mitochondrial diseases and the discovery of the differential sensitivity of
different tissues to mutations in mtDNA, neurons and muscle cells were
deduced to be the most active tissues, as already anticipated from their physi-
ological functions. Cardiac muscle is a specifi c example of a tissue deriving
most of its ATP from respiration; and not surprisingly, cardiac myopathies are
prominent among the clinical manifestations associated with mutations in
mtDNA.
To repeat, one may not expect to fi nd rapid, short - term fl uctuations in
mitochondrial biogenesis in a given cell type, but one can anticipate that the
expression of a large number of genes encoding respiratory and other mito-
chondrial proteins may be expressed at a higher level in some cell types com-
pared to others, and thus the existence of common regulatory molecules may
be the simplest mechanism for a coordinate expression.
NUCLEAR GENES ENCODING MITOCHONDRIAL PROTEINS 91
92 BIOGENESIS OF MITOCHONDRIA
The number of mammalian genes investigated fully is still small. For practi-
cal reasons, cDNA clones are the fi rst to be isolated; and the step to the full
genomic clone, or at least the promoter region, is nowadays made possible by
the availability of whole genome sequences. From the examples investigated,
one may tentatively conclude that there is no single universal cis - acting
sequence element with its associated transcriptional activator which is common
to all such genes. However, at least two factors have been characterized which
play key roles in the regulation of entire subsets of genes. Which genes belong
to the set, and why, is not yet clear.
Evans and Scarpulla (116, 117) were the fi rst to defi ne a factor, designated
as NRF - 1 (nuclear respiratory factor 1), from their analysis of the promoters
of the mammalian (rat) cytochrome c and cytochrome oxidase subunit Vb
(COXVb). Sequence analysis of the cytochrome c promoter revealed the pres-
ence of several consensus sites for well - known transcription factors such as
Sp1, CCAAT, and the cAMP response element binding protein CREB. In
addition, a new cis - acting element was defi ned as a palindromic motif consist-
ing of tandemly repeated GC sequences over a single helical turn that interacts
with the NRF - 1. The case was strengthened when over the years several other
nuclear genes encoding mitochondrial proteins were found to have the
same element (COXVb, COXVIIaL, COXVIc, mtTFA, a ubiquinone - binding
protein, a mtRNA processing enzyme, aminolevulinate synthase) (118) ,
prompting Scarpulla and his colleagues to purify the human protein to near
homogeneity from cultured cells. Partial peptide sequences were used to syn-
thesize degenerate primers, which in turn led to the isolation of full - length
cDNA for NRF - 1 and the production of authenticated recombinant protein
in bacteria (119, 120) . NRF - 1 is a single peptide with a novel DNA - binding
domain (related to DNA - binding domains in two developmental regulatory
proteins found in Drosophila and the sea urchin) and a transactivating domain.
The fi nding of an NRF - 1 site in the promoter for the mitochondrial transcrip-
tional activator (mtTFA) is of special interest. Since mtTFA is required for
both replication and transcription of mtDNA (see Section 4.4.1 ), one can begin
to build a case for a role of NRF - 1 in the nuclear control of mtDNA copy
number and gene expression. Aminolevulinate synthase is also not an arbi-
trary, irrelevant enzyme for this discussion. It is the fi rst and rate - limiting
enzyme in the pathway of heme synthesis in the matrix. Although a role for
heme in the control of expression of mammalian cytochromes is not yet as
well - defi ned as in yeast, the connection is tantalizing.
When several other promoters of respiratory genes (e.g., COXIV) either
failed to show an NRF - 1 - binding site or could not be footprinted with pure
NRF - 1, the search for additional recognition sites and factors by Virbasius and
Scarpulla succeeded in the identifi cation of a second factor termed NRF - 2
(121, 122) . The core GGAA motif was somewhat familiar from the previous
characterization of the ETS - domain family of transcription factors, but when
the NRF - 2 site from COXIV was used in an affi nity column, a novel DNA
binding activity could be purifi ed (123) . The DNA - binding activity co - purifi ed
with fi ve distinct polypeptides ( α , β
1
, β
2
, γ
1
, γ
2
). The α subunit was subsequently
identifi ed as the DNA - binding peptide, but it is found as a heteromeric complex
with one of the other peptides ( αβ
1
, αβ
2
, αγ
1
, αγ
2
). These complexes differ in
their affi nity for the NRF - 2 site and to tandem NRF - 2 sites, a characteristic
that had already been observed for the related ETS - domain transcription
factor GABP of the mouse. Further cloning and sequencing of all fi ve human
NRF - 2 subunits established NRF - 2 as the homologue of the mouse GABP,
a widely expressed transcription factor fi rst defi ned by the study of viral
promoters. Consensus sequences for the NRFs are the following: NRF - 1: . . .
YGCGCA Y GCGC R . . . ; NRF - 2: . . . ACC GGAA GAG. . . .
Reminiscent of the Hap2/3/4 complex in yeast, NRF - 2 has DNA - binding
domains and transactivation domains on separate peptides. In addition, the
affi nity of some of the heterodimers for a promoter can be further enhanced
by cooperative binding to tandem NRF - 2 motifs, and overall a mechanism is
created to modulate promoter strength for different genes.
It was also fi rst found by Scarpulla and his co - workers that some genes
have both NRF - 1 and NRF - 2 motifs that contribute to the control of expres-
sion of these genes (124) . Another example was found in the promoter for
the iron – sulfur subunit of succinate dehydrogenase (and complex II) (125,
126) . In this promoter the two sites are within 200 nucleotides upstream from
the transcription start site, and mutagenesis of either site reduced the expres-
sion of a reporter gene from this promoter 5 - to 10 - fold. Both NRF - 1 and
NRF - 2 were shown to footprint the respective sites independently. Other
promoter elements were found further upstream (Sp1, CCAAT), but deleting
them had no signifi cant effect on the expression of a reporter gene in fi bro-
blasts, myoblasts, or differentiated myotubes. For a more detailed and expert
review the reader should consult (127, 128) . A highly reduced summary of
many studies in mammals is that environmental conditions infl uence the
expression of PGC - 1 family coactivators (PGC - 1alpha, PGC - 1beta, and PRC),
which, in turn, target specifi c transcription factors (NRF - 1, NRF - 2, and ERR
alpha) in the expression of respiratory genes (128a) . The notion of having a
limited, select number of transcription factors and promoter sequences exclu-
sively devoted to the transcription of respiratory genes and mitochondrial
proteins is apparently too simple. Several such genes have been found to lack
NRF - 1 or NRF - 2 sites altogether (based on sequence inspection), while other
genes unrelated to mitochondrial function contain such sites in their promoter
region.
Mammalian cytochrome oxidase (COX, complex IV) consists of 13 sub-
units, 10 of which are encoded in the nucleus. It has been intriguing that three
of the nuclear - encoded subunits (VIa, VIIa, and VIII) exist as multiple iso-
forms that are differentially expressed in different tissues. The L - form is
named after its identifi cation in liver, and this form is also found in many
other tissues. The H - form is found predominantly in skeletal and cardiac
muscle (heart). A detailed analysis of the COXVIaH promoter region with a
reporter construct in transgenic mice as well as in tissue culture has verifi ed
NUCLEAR GENES ENCODING MITOCHONDRIAL PROTEINS 93
94 BIOGENESIS OF MITOCHONDRIA
the highly specifi c elevation of expression in heart and muscle, with some
activity in the brain but virtually none in other tissues. Expression of this
gene was suppressed in myoblasts, but dramatically induced when these myo-
blasts were induced to differentiate into myotubes. A 300 - bp promoter
segment was entirely suffi cient to mimic the expression of this gene and its
dependence on muscle differentiation. No NRF - 1, NRF - 2, or other element
previously associated with respiratory genes (see above) was found in this
region. Instead, an MEF2 consensus sequence and an E - box element were
shown to be essential by mutational analysis (129) . The muscle enhancer
factor (MEF) comprises a family of four genes induced by myogenic HLH
proteins (myo - D, myogenin, Myf5, and MRF4), and MEF2 has been demon-
strated to be signifi cantly involved in muscle differentiation (see references
130 and 131 for summaries and examples). Thus, the activation of the
COXVIaH gene during muscle cell differentiation is understandable from a
consideration of its promoter, but the promoter does not betray that the
protein functions in mitochondria.
A very similar analysis was made with the SDH2 promoter (iron – sulfur
subunit of complex II) (126) . Transcripts of this gene are also signifi cantly
elevated in adult skeletal and cardiac muscle. In contrast to COXVIaH,
however, the expression of the SDH2 gene was not up - regulated upon myotube
formation, consistent with the absence of promoter elements responding to
myogenic transcription factors. If there is no general, coordinate induction of
respiratory chain peptides, one may wonder about the purpose of inducing
COXVIaH mRNA by at least an order of magnitude in myotubes compared
to myoblasts. It has been proposed from experiments with the homologous
subunit in yeast that this subunit is responsible for sensing ATP and phosphate
levels — that is, for the modulation of enzymatic activity of complex IV under
varying metabolic conditions (132) . For this factor to participate in the regula-
tion of cytochrome oxidase activity requires one of two possible mechanisms:
Either the subunit VIaH replaces the preexisting subunit VIaL directly, or
there is turnover of the entire complex, with newly made complex including
the muscle - specifi c VIaH subunit.
Thus, at fi rst sight, the well - controlled and already thoroughly investigated
rat or mouse myoblast/myotube system in tissue culture may serve as a good
model system in which to study mitochondrial biogenesis and the control of
respiratory gene expression. Much previous attention has been focused on
muscle specifi c proteins such as creatine kinase and the various cytoskeletal
proteins of the sarcomere. On the other hand, the preliminary lessons are that
more subtle mechanisms for modulating mitochondrial respiration may be at
work, requiring only a limited number of changes in the composition of the
electron transport complexes. A second and very interesting preliminary con-
clusion is that myotube differentiation in culture is incomplete, in the sense
that maturation, stimulation, and contractile activity of a myofi bril may be
required for further changes in respiratory gene expression and mitochondrial
capacity.
Two other characterizations of genes had once raised expectations about
fi nding more universal promoter elements that might be unique or specifi c for
mitochondrial proteins. The analysis of the promoters for the ATP/ADP trans-
locator and for the β subunit of the F1 - ATP synthase had identifi ed so - called
OXBOX and REBOX elements that were postulated to respond to nuclear
factors present at variable levels in myoblasts, myotubes, and HeLa cells (133,
134) . A generalization of these conclusions awaits the characterization of more
genes in this category, but at this time one can state that many other relevant
genes already analyzed do not contain these particular elements as a recogniz-
able consensus sequence.
4.2.2.4 Co-evolution of Nuclear and Mitochondrial Genomes From the
above description it is clear that several dozen nuclear - coded proteins interact
either with the mitochondrial genome (replication, transcription, etc.) or
directly with proteins encoded by the mt genome. The mt genome suffers
sequence substitutions at a signifi cantly faster rate than the nuclear genome.
If altered proteins result from mtDNA substitutions, compensatory changes
may have to occur in the nuclear genome to assure proper strong, functional
interactions between the gene products, in forming a complex of the electron
transport chain, for example. Thus the two genomes are expected to co - evolve,
and strong evidence for this concept has been deduced from studies with
interspecifi c hybrids and cybrids of mammalian cells. The fi rst experiments
addressing this issue were reported by Clayton et al. (135) from their analysis
of mtDNAs in human - mouse interspecies hybrids. Further pioneering studies
of Wallace and Eisenstadt (136) indicated quite convincingly that the nuclear
and mitochondrial genome must be compatible; that is, there is species specifi c-
ity, and a human mtDNA cannot coexist for long in a rodent cell, or vice versa.
More specifi cally, in the Scheffl er laboratory, it was found that nuclear muta-
tions in a hamster cell mutant causing a defect in complex I could be comple-
mented by a gene on a hamster or mouse X chromosome, but not by a human
X - linked gene (137) . This result was also interpreted in terms of the incompat-
ibility of heterologous gene products.
A very informative test of genome compatibility was recently reported by
Kenyon and Moraes (138) . By fusing enucleated cells from the hominoid apes
(chimpanzee, pigmy chimpanzee, gorilla, and orangutan) with ρ
0
(mtDNA -
less) human cells these authors created “ xenomitochondrial cybrids ” with ape
mtDNA and human nuclear DNA. Only the combinations of human nuclei
with mitochondria from the most closely related species (chipanzee, gorilla)
yielded cells capable of oxidative phosphorylation. Orangutan mitochondria
in such cells did not express a functional OXPHOS system. An even more
specifi c example of incompatibility between species was reported by Yadava
et al. (139) . The MWFE subunit of complex I is highly conserved in mamma-
lian species, and yet a human protein fails completely to complement a defi -
ciency in a hamster cell. A change of two amino acids (out of 70) in the human
protein restores the function of this protein in hamster cells.
NUCLEAR GENES ENCODING MITOCHONDRIAL PROTEINS 95
96 BIOGENESIS OF MITOCHONDRIA
4.3 REPLICATION AND MAINTENANCE OF
MITOCHONDRIAL DNA
4.3.1 DNA Replication in Mammalian Mitochondria
Transcription and replication of mtDNA are basic processes which are closely
related to both the function of the organelle and its biogenesis (75) . The
present section will deal with mtDNA replication, with an emphasis on animal
mitochondria. When mtDNAs were discovered almost 50 years ago and the
fi rst estimates of their size became available, it also became clear that the rela-
tively small size of animal mtDNAs made them attractive candidates for study.
Expectations were not disappointed, and our understanding of DNA replica-
tion and transcription in animal mitochondria is relatively far advanced.
Insights from such studies will be presented fi rst. An update on studies in other
organisms will summarize the progress, provide some contrasting or unique
characteristics, and emphasize some of the issues for which answers still have
to be sought.
Mitochondrial DNA replication can be investigated conveniently in tissue
culture, where intracellular mitochondrial proliferation is loosely coupled to
cellular division and multiplication. The total number of mtDNAs has to
double in each cell cycle. Replication intermediates can be found readily.
Before entering into the details of this process, one should be reminded that
the situation may be more complicated in an intact animal, and several aspects
of the process need to be discussed separately. It is necessary to distinguish
between the number of mitochondria per cell and the number of mtDNA
molecules per cell (estimated to be a few thousand). A priori it makes sense
for mitochondrial DNA replication to be controlled to maintain a roughly
constant number of mitochondria per cell, and thus there has to be coordina-
tion with DNA replication in the nucleus. However, during embryonic devel-
opment and differentiation in mammals the number of mitochondria per cell
may be altered, depending on the energy requirements of particular tissues. It
is possible that in adult tissue such as muscle and brain there is also turnover
of mitochondria and hence mtDNA replication which is not linked to nuclear
DNA replication and cell division. A particularly fascinating situation may
occur during oogenesis, where only a small subset of the total population of
mtDNA molecules may be involved in mtDNA replication to generate the
large number of mitochondria found in the mammalian egg. While there is as
yet no proof of this hypothesis, such a “ bottleneck ” has been invoked to
explain observations on mitochondrial inheritance and the phenotypic expres-
sion of mitochondrial mutations in heteroplasmic cells (140) . Alternatively, a
bottleneck can also be created if there is no replication of mtDNA during the
fi rst 10 – 15 zygotic divisions following fertilization. A more complete discussion
of this topic will be found in Chapter 7 .
In cells in tissue culture, mtDNA replication seems to be loosely coupled to
nuclear DNA replication, with the precise mechanism of the control of copy
numbers yet to be elucidated. On the other hand, the process of replication
itself is quite well understood (141 – 143) . Mammalian mtDNAs, and probably
most vertebrate mtDNAs, have two origins of DNA replication, one responsi-
ble for each strand. The asymmetric distribution of nucleotides (G + C) in
mammalian mtDNA allows one to distinguish a “ heavy ” and “ light ” strand on
alkaline CsCl gradients, and hence one can also speak of O
H
as the origin of
H - strand synthesis and O
L
as the origin of L - strand synthesis. O
H
is located
within the region devoid of genes referred to as the displacement loop, or D -
loop region, and at other times as the control region. Initiation and elongation
of the H - strand is the fi rst event, and it proceeds a considerable distance around
the circle; only after the second origin (O
L
) is displaced as a single - stranded
template will DNA replication of the other strand and in the opposite direction
begin. O
H
is therefore dominant, and O
L
is not an independent origin, since
DNA replication beginning at this origin without the prior activation of O
H
has
never been observed. It is signifi cant that O
L
is located within a cluster of fi ve
tRNA genes. When this region has been displaced and is found in single -
stranded form, it is likely that a secondary structure is formed which contrib-
utes to the functioning of the origin. Support for such a hypothesis comes from
the observation that in chickens the O
L
sequence itself (found in mammals) is
absent, but a similar cluster of fi ve tRNAs is still found at this location (144) .
Strand elongation by a mitochondrial DNA polymerase (polymerase γ )
proceeds in the usual 5 ′ – 3 ′ direction. The nature of this polymerase and its
relationship to the various bacterial and eukaryotic DNA polymerases is an
interesting one (see reference 51 for a more explicit discussion). Its low abun-
dance has made biochemical studies a challenge, but the enzyme has now been
characterized from several organisms including humans (63, 64, 145 – 148) . Pol
γ prefers ribohomopolymer templates, and therefore it was initially thought to
represent a cellular reverse transcriptase resembling the reverse transcriptase
of tumor viruses. Later it was recognized that nuclear preparations had been
contaminated with mitochondria. It is antigenically completely different from
the viral enzymes, and it is further distinguished by its inability to use natural
RNAs as templates. Other distinguishing characteristics include stimulation
by salt, resistance to aphidocolin, inhibition by N - ethylmaleimide, and by
dideoxynucleotide triphosphates (51) .
The most highly purifi ed preparations suggested that it consists of a holo-
enzyme of ∼ 125 – 140 kDa (POL
γ
A) that has both catalytic and exonuclease
activity, and it is associated with a smaller subunit of 35 – 54 kDa (POL
γ
B) that
functions as an essential processivity factor. When pol γ genes were cloned
(e.g., MIP1 from S. cerevisiae ), a 140 - kDa polypeptide was found to include
both polymerase and exonuclease domains with recognizable homology to
prokaryotic, A - type DNA polymerases — for example, E. coli DNA polymerase
I. The cloning of X. laevis and D. melanogaster pol γ genes confi rmed the con-
clusion that the polymerization and 3 ′ to 5 ′ exonuclease functions are com-
bined in one polypeptide (143) . The reconstitution of a minimal mtDNA
replisome in vitro has recently been achieved in the laboratory of M.
REPLICATION AND MAINTENANCE OF MITOCHONDRIAL DNA 97
98 BIOGENESIS OF MITOCHONDRIA
Falkenberg (149a) . Three proteins were defi ned to act in concert at the replica-
tion fork by a mechanism that resembles T4 or T7 replisomes: Polymerase γ ,
mtSSB (a single - strand binding protein), and TWINKLE, a mitochondrial
DNA helicase with 5 ′ to 3 ′ directionality. Single - stranded DNA molecules
could be generated at a rate of 180 bp/min, close to the rate reported for in
vivo DNA replication.
While there are no novel or unusual aspects of the mechanism of DNA
strand elongation in mitochondria, the same cannot be said about priming and
initiation of DNA replication. It became clear some time ago that DNA rep-
lication and transcription of mtDNA were directly and intimately linked. The
fi rst indication was that the two known promoters for transcription are both
located in the control region. They are ∼ 150 bp apart in mouse and human
mtDNA and control transcription in opposite directions; hence they can be
referred to as the light - strand promoter (LSP) and the heavy - strand promoter
(HSP). Their precise identifi cation became possible with the establishment of
a faithful in vitro system for transcription which will be described in further
detail in the section on transcription. Three key fi ndings were made when
detailed characterizations were made of the major nascent DNA chains at the
earliest stages in DNA replications, and of all the transcripts from the LSP.
First, all newly initiated DNA chains had a 5 ′ end that corresponded to the
sequence ∼ 200 nt downstream from the LSP. Second, LSP transcripts were of
two types: (a) the expected long transcripts that would later be processed to
yield eight tRNAs and one mRNA and (b) short transcripts with 3 ′ termini
precisely or closely adjacent to the 5 ′ termini of the nascent DNA chains. It
was a very plausible jump to the conclusion that the RNA transcripts might
serve as primers for the start of DNA replication from the O
H
origin. A satisfy-
ing confi rmation of this idea was provided by the third fi nding: LSP transcripts
still covalently attached to the nascent DNA (in the mouse). Transcriptional
activation is therefore closely linked to, and a prerequisite for, leading - strand
DNA replication. A current model incorporating many fi ndings is presented
in Figures 4.5 and 4.6 (143) .
Figure 4.5 shows the control region of the vertebrate mt genome between
the Phe - tRNA on the left and the Thr - tRNA on the right. Initiation of tran-
scription at the L - strand promoter generates an RNA that remains fi rmly
associated with the DNA in the region defi ned by the conserved sequence
blocks (CSBs), displacing the DNA strand to form a loop. Subsequent process-
ing on either side releases a long downstream transcript and a short upstream
fragment. The RNA segment remaining in the R - loop can then serve as primer
for DNA synthesis. The D - loop is formed when the extension of the newly
synthesized H - strand is arrested (see below). Initiation of mtDNA replication
in other vertebrates is likely to be quite similar, with some variations in the
size of the control region, the nature of the CSB elements (with CSB1 appar-
ently the most essential), and the spacing between them (143) .
Recent studies in vitro have provided additional support to this model
(150) . RNA of defi ned length made in vitro was capable of annealing to
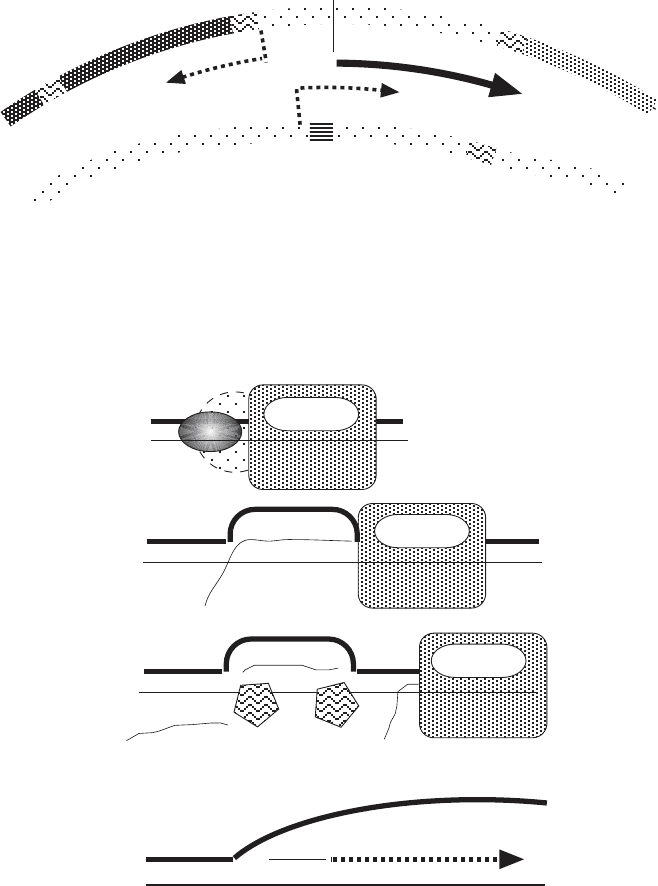
Figure 4.5 The D - loop region of mammalian mitochondrial DNA and the fl anking
genes. HSP and LSP are the heavy - strand and light - strand promoters. A processed
transcript from the LSP serves as the primer for DNA replication from the O
H
origin.
(Modifi ed after D. Clayton.)
O
H
cytochrome b
LSP
12s rRNA
Thr
Phe
Leu
5’
3’
HSP
Pro
CSBs
Figure 4.6 Detailed schematic representation of the generation of the RNA primer
for DNA replication. RNA polymerase and factor(s) generate an RNA transcript. This
transcript is modifi ed by a mitochondrial RNA processing enzyme (MRP), leaving a
short RNA tightly annealed to the DNA and creating an RNA loop. Priming from this
RNA of DNA replication leads to the generation of the larger D - loop. (Modifi ed after
D. Clayton.)
5’
5’
5’
R-loop
nascent H-strand (DNA)
D-loop
RNA pol
RNA pol
RNA pol
RNA primer
MRP
REPLICATION AND MAINTENANCE OF MITOCHONDRIAL DNA 99
100 BIOGENESIS OF MITOCHONDRIA
supercoiled mouse mtDNA under defi ned conditions, creating an “ R - loop. ”
This R - loop had been previously observed in in vitro transcription experi-
ments with purifi ed mitochondrial RNA polymerase activity, and it had been
shown to consist of a “ persistent ” RNA – DNA hybrid. It was also possible to
create the R - loop by in vitro transcription with SP6 polymerase from an SP6
promoter inserted into mouse mtDNA control region. Its formation was not
dependent on the presence of a particular RNA polymerase or additional
protein, but did require negative superhelical turns in the mtDNA.
The R - loop with the RNA – DNA hybrid must have unusual stability, since
it is stable to manipulation and purifi cation without branch migration, and
even linear restriction fragments containing the R - loop are stable for consider-
able time at 37 ° C without loss of one of the polynucleotide strands. The RNA
in the R - loop is not base paired with the complementary DNA strand along
its entire length. From a detailed examination and comparison of RNase H
and RNaseT1 cleavage sites in the R - loop with those of a double - stranded
hybrid made from the same RNA and its cDNA, it was concluded that a
unique structure must be formed. A model of the R - loop has emerged in which
intramolecular RNA – RNA interactions stabilize and protect distinct regions
against single - strand - specifi c nucleases (stem - loops?), while other segments
interact with unique sites on the DNA. An examination of the relevant DNA
sequence has revealed three conserved sequence blocks (CSB I, CSB II, CSB
III), and both CSB II and CSB III contain poly G tracts and are absolutely
required for R - loop formation. From the results of the probing with single -
strand - specifi c nucleases and RNase H, the whole structure cannot be repre-
sented by conventional Watson – Crick basepairing schemes. The CSBs
sequences attracted attention as potential cis - acting regulatory regions as early
as 1981, because they were short, highly conserved regions within an otherwise
highly divergent region (141, 143) .
Endonucleolytic processing of the RNA in this hybrid structure is postu-
lated to create a primer which is positioned in the correct position for exten-
sion by deoxyribonucleotides with DNA polymerase. A candidate enzyme
performing this function was isolated by Clayton ’ s laboratory in 1987 (142,
151, 152) . It was shown to be a site - specifi c endonuclease capable of cleaving
RNA transcripts from the D - loop region between CSB II and CSB III; and
most intriguingly, this RNase MRP (mitochondrial RNA processing) contains
an RNA component encoded by nuclear genes in mouse and humans. Base-
pairing interactions between this RNA and the substrate RNA for this nucle-
ase are therefore likely to guide substrate selection and cleavage site selection.
Undoubtedly, the secondary structure of the LSP transcript within the R - loop
is also a strong determinant of specifi city and activity. Identical ribonucleopro-
teins have been isolated from several mammals, the frog Xenopus laevis , and
even yeast. Cloning of these genes as well as the genes encoding the RNA
component has confi rmed a high degree of evolutionary conservation (143) .
As discussed in the review by Shadel and Clayton (143) , it was a challenge to
show that the MRP is indeed found in mitochondria, since a very similar activ-
