Scheffler Immo E. Mitochondria
Подождите немного. Документ загружается.

-
-
-
O
O
O
P
O
O
O
P
OCH
2
CH
2
CH
3
C
CH
2
O POP
O
POP
O
POP
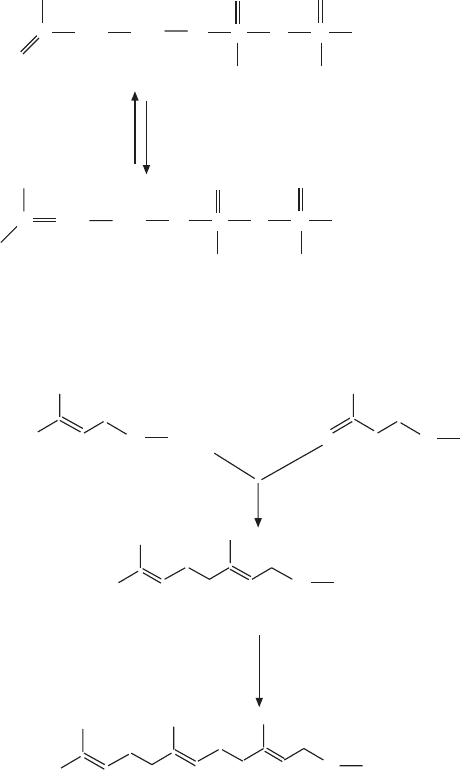
isopentenyl-pyrophosphate
dimethylallyl pyrophosphate
geranyl-pyrophosphate
farnesyl-pyrophosphate
CH
C
CH
CH CH
O
3
3
P
O
O
O P
O
O
O
-
-
-
2
O
POP
B
Figure 6.7 (Continued)
of the enzymes involved in the branch - point reactions have a high affi nity for
FPP, and fl ux into these branches can occur even at low FPP concentrations.
The branch - point enzyme squalene synthase, on the other hand, has a low
affi nity for FPP, and thus cholesterol biosynthesis will occur only when high
concentrations of FPP are achieved by an activation of HMG - CoA
reductase.
Protein isoprenylation with FPP or GGPP (see Figure 6.7C ) as isoprenoid
donors is now recognized to be widespread in eukaryotic cells. Proteins such
BIOSYNTHESIS OF UBIQUINOL (COENZYME Q) 321
322 METABOLIC PATHWAYS INSIDE MITOCHONDRIA
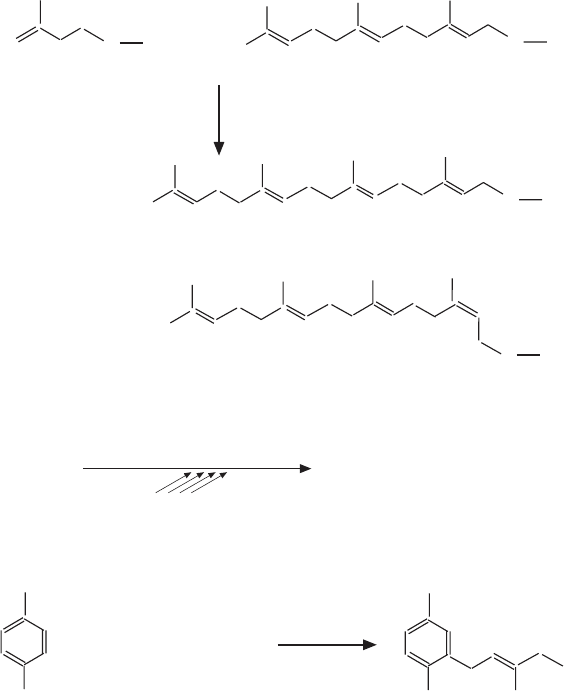
isopentenyl-PP (IPP)
farnesyl-PP (FPP)
all-trans-geranylgeranyl-PP (GGPP)
trans,trans,cis-geranylgeranyl-PP (GGPP)
O
POP
O
POP
O
POP
GGPP
Polyprenyl-PP
IPP
+
polyprenyl-PP
OH
COOH
O
POP
+
or
OH
COOH
()
n
C
Figure 6.7 (Continued)
as the ras protein can be derivatized with long hydrophobic prenyl side chains
that serve to anchor them on membranes at specifi c subcellular locations. A
full discussion of this subject is beyond the scope of this chapter. As shown in
Figure 6.7C , geranyl - geranyl - pyrophosphate (GGPP) is made from FPP and
IPP by enzymes named GGPP synthases (a subset of prenyltransferases).
There are at least two types, and the product can be either the all - trans - GGPP,
or the trans, trans, cis - GGPP. These enzymes differ in their intracellular distri-
bution; they may be cytosolic or membrane - associated; they have been found
in mitochondria, peroxisomes, and the ER; either Zn
2+
or Mg
2+
may be abso-
lutely required. The cytosolic enzyme appears to produce only all - trans - GGPP,
while trans, trans, cis - GGPP is produced by membrane - bound enzymes.
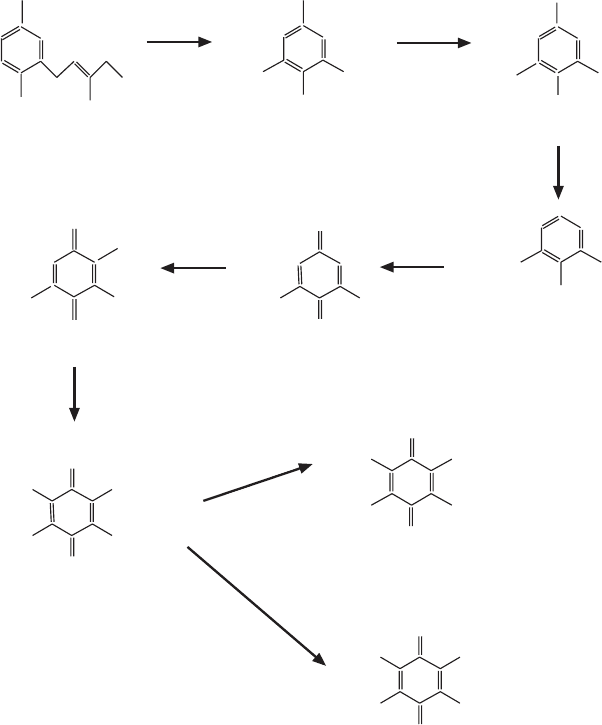
ubiquinone
rhodoquinone
R
HO
OH
COOH
COOH
OH
R
CH
3
O
CH
3
O
O
O
R
CH
3
R
O
O
CH
3
O
OH
R
CH
3
O
R
HO
O
O
CH
3
CH
3
O
R
O
O
CH
3
CH
3
O
CH
3
O
CH
3
O
CH
3
R
O
O
NH
2
(
)
n
OH
COOH
D
Figure 6.7 (Continued)
Additional prenyltransferase reactions with IPP as donor serve to elongate
the chain, but the all - trans - and the trans, trans, cis - GGPP have different fates.
The latter is used for the synthesis of dolichol, containing between 16 and 23
isoprenoid units in animal cells. Dolichol is phosphorylated and esterifi ed to
oligosaccharides to form an essential intermediate in glycoprotein syntheses
on the ER. This is another long story not to be pursued here. The all - trans -
GGPP has been clearly established as the substrate for protein isoprenylation;
and it is also likely to be the substrate for trans - prenyltransferase, leading to
the synthesis of all - trans - polyisoprenoid - PP. Consisting of about 9 or 10 units,
BIOSYNTHESIS OF UBIQUINOL (COENZYME Q) 323
324 METABOLIC PATHWAYS INSIDE MITOCHONDRIA
the polyisoprenyl group can be transferred to p - hydroxybenzoic acid in the
fi rst distinct step leading to ubiquinone (Figures 6.7C and 6.7D ).
The subcellular localization of the distinct prenyl transferase enzymes could
be used to control the destination of the various products, and also serve in
establishing separate regulatory mechanisms. Cell fractionation studies in con-
junction with genetic analyses are not yet complete to present a fi nal picture.
As long as ubiquinone was thought to be exclusively present in the inner
mitochondrial membrane, the presence of the biosynthetic enzymes in mito-
chondria could be rationalized. However, it has since been established that
ubiquinone is found in extra - mitochondrial membranes as well. Are there
multiple enzymes for ubiquinone synthesis, or can this molecule be exported
from mitochondria? The more abundant species is found to be the reduced
form (ubiquinol), and there appears to be agreement that ubiquinol has an
important function as an antioxidant (34) . Its abundance excedes α - tocopherol
(vitamin E) by a factor of 10 in animal cells.
An interesting problem presents itself in certain eukaryotic microorganisms
that have complex life cycles and may include a stage that exists in an aerobic
environment (carrying out normal respiration and oxidative phophorylation),
and another stage living in an environment requiring drastic adjustments in
metabolism and even anearobic electron transfer (35) . One such example of
a parasite, the liver fl uke, Fasciola hepatica , has been studied in some detail.
The free - living stage (aerobic) oxidizes succinate and transfers electrons via
UQ
10
to complex III of the electron transfer chain. The parasitic stage oxidizes
NADH via complex I and the reduction of fumarate to succinate by fumarate
reductase. The mobile carrier between complex I and FRD is another quinone,
rhodoquinone (RQ) (36, 37) . Experiments with [2 -
14
C] - mevalonate demon-
strated that UQ and RQ can be synthesized de novo in these organisms.
Rhodoquinone has an amino group substituted on the quinone ring, and this
modifi cation must be part of the developmental program of this organism. The
substitution alters the redox potential of the RQ/RQH
2
redox couple that
appears necessary to allow electron fl ow to occur in the reverse direction in
FRD compared to complex II (38) .
6.8 BIOSYNTHESIS OF Fe – S CENTERS
Iron – sulfur clusters were discovered in the early 1960s (39) , and for several
decades it was assumed that they formed spontaneously. Research during the
past decade has revealed that the process is surprisingly complex, and mito-
chondria were found to play an essential and central role in it. The studies were
greatly stimulated by research on the biosynthesis of Fe – S clusters in bacteria
(40) , as well as by the use of yeast as a model organism for genetic and bio-
chemical approaches. In turn, insights from studies in yeast can be extrapolated
to higher eukaryotic systems including humans with increasing confi dence.
Many of the components involved in the machinery have been highly con-
served in evolution. This has greatly facilitated the search for, and the identifi -
cation of, the relevant genes/proteins in higher eukaryotes from knowledge
about the corresponding genes in prokaryotes or in other eukaryotes.
Bacteria are capable of Fe – S cluster assembly by three distinct systems
(NIF, ISC, SUF), one or two of which may be present in the same cells depend-
ing on the strain. The NIF machinery is most specifi c and exclusively respon-
sible for the biosynthesis of nitrogenase required for nitrogen fi xation. The
SUF machinery may be induced (?) under iron - limiting or stress conditions,
but it may also be the only system present — for example, in cyanobacteria. The
majority of cellular Fe – S proteins are generated with the help of the ISC
machinery that may therefore be considered a “ housekeeping ” system. It is
the system inherited by eukaryotic cells from their bacterial symbionts, with
modifi cations and additions made in the course of evolution. In the following
the ISC system of eukaryotes will be the focus of attention.
Fe – S clusters in proteins can serve a variety of functions (40, 41) . In the
present context, their role in electron transfer and redox reactions is the most
prominent, but they also serve in enzymatic reactions in substrate binding and
activation (without electron transfers), or they may have a purely structural
role in stabilizing the tertiary structure of a protein. They may serve in iron
storage (ferredoxins) or in iron - sensing (cytoplasmic aconitase). As the list is
growing, it is important to recognize that Fe – S proteins are found in at least
three distinct compartments of a eukaryotic cell: mitochondria, cytosol, and
nucleus. Only one example of a nuclear Fe – S protein is known so far (DNA
glycosylase). A larger list of cytoplasmic Fe – S proteins includes a variety of
metabolic enzymes, and two that deserve special mention. The cytosolic aconi-
tase alias “ iron - regulatory protein 1 ” (IRP - 1) has already been introduced as
the sensor of intracellular iron (page 312). When intracellular iron becomes
defi cient, its Fe – S cluster dissociates, producing a protein with high affi nity for
an iron - responsive element (IRE) localized in the 5 ′ UTR or 3 ′ UTR of a subset
of mRNAs. Their stability or translatability is controlled by this mechanism,
and thus the level of key enzymes/proteins associated with uptake, storage,
and utilization of iron (e.g., heme biosynthesis) is regulated. The ABC protein
Rli1is essential for the biogenesis of ribosomes, RNA processing, and transla-
tion initiation in yeast (42, 43) . It is one of the most conserved proteins in
evolution, and under some conditions in yeast it may be the only indispensable
Fe – S protein (see below).
An authoritative and comprehensive reviews of Fe – S protein biogenesis
in eukaryotes by one of the pioneering laboratories in this fi eld has
appeared recently (41) . The reader is referred to this review and a comple-
menting review emphasizing the bacterial systems (40) , but the key features
of the eukaryotic system are also concisely summarized by Johnson et al. A
summary and some discussion is presented here with reference to Figure 6.8
(from reference 41 ).
The process can be decomposed into several distinct stages, beginning with
import of Fe
2+
into mitochondria. The ISC assembly machinery is made up of
BIOSYNTHESIS OF Fe – S CENTERS 325
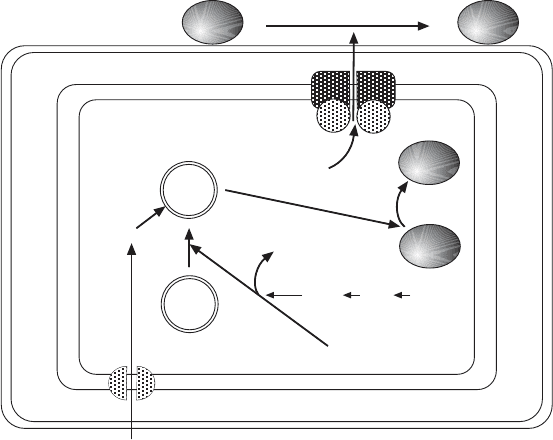
326 METABOLIC PATHWAYS INSIDE MITOCHONDRIA
Figure 6.8 Biosynthesis of [Fe – S] centers (adapted from reference 41) .
Yfh1
Fe
2+
Fe
S S
Fe
[2Fe-2S]
SSq1/ATP
Jac1/Mge1
Grx5
Ala
Cys
Nfs1
Yah1 Arh1
NADH
Isa1/2 ?
Nfu1 ?
Isu1/2
Atm1
Erv1
[2Fe-2S]
CIA machinery:
Cfd1, Nbp35, Nar1
ISC
assembly
machinery
CYTOSOL
INTERMEMBRANE SPACE
MATRIX
so - called scaffolding proteins (Isu1,2) to which the two constituents of a Fe – S
cluster are delivered. Yfh1 is the yeast homologue of human frataxin; its func-
tion in yeast is to deliver Fe
2+
to the ISC assembly machinery. Defects in this
protein lead to an impairment of Fe – S cluster biosynthesis, mitochondrial iron
accumulation, heme defi ciency, and oxidative stress. In humans, frataxin defi -
ciency causes a neurodegenerative disease, Friedreich ’ s ataxia (44) . The second
branch in the lower part of the fi gure is concerned with the production and
delivery of the sulfi de (S
2 –
). The sulfi de is produced from cysteine by a pyri-
doxal phosphate - dependent cystein desulfurase (Nsf1p). A more speculative
aspect of this pathway is the role of the Yah1 and Arh1 proteins. Lill and
Muhlenhoff propose an electron transport chain for electrons provided by
NADH and transferred to the desulfurase and to elemental sulfur (S
0
) to form
the sulfi de (S
2 –
) ion. Intriguingly, in yeast these proteins are also required,
together with Cox10p and Cox15p, for the synthesis of heme a in complex IV,
while homologous proteins in the mammalian adrenal gland are associated
with steroidogenesis from cholesterol.
The Fe – S cluster thus formed and bound to the scaffold (consisting of Isu1p,
Isu2p, and possibly other proteins) now has to be transferred to apoproteins
in the mitochondria, where they may remain as soluble proteins (e.g., aconi-
tase), or they will have to be incorporated into the various complexes of the
electron transport chain. The temporal sequence of these events is not clear.
For example, there are three Fe – S clusters in the SDHB subunit of complex
II, and it remains to be established whether these clusters are incorporated
before or after association of this subunit with the fl avoprotein, SDHA, and
before or after the association of SDHB with the two integral membrane
proteins SDHC and SDHD. Elucidation of the assembly pathway of the
peripheral subcomplex of complex I with 7 or 8 Fe – S clusters in several dif-
ferent subunits can be anticipated to be even more challenging. Genetic studies
in yeast have identifi ed additional components believed to act in the transfer
of the Fe – S cluster from the initial scaffold to diverse target proteins. SSq1p
and Jac1p are chaperone proteins, and Grx5 is glutaredoxin. A precise mecha-
nism for their involvement has not been formulated, and ATP hydrolysis is a
likely driving force for various association/dissociation steps. A role for the
gene products from the ISA1/2 and NFU1 genes in yeast can be inferred from
the defects in the activity and synthesis of various mitochondrial Fe – S proteins
resulting from a depletion of Isa1p or Isap2. Nfu1 - like proteins have been
demonstrated to be required for Fe – S protein biosynthesis in plant mitochon-
dria, chloroplasts and cyanobacteria. In yeast a deletion of Nuf1 has no phe-
notype, but synthetic lethality with other ISC genes (e.g., ISU1 ) suggests an
auxiliary function.
As mentioned above, Fe – S proteins are also found in the cytosol and even
in the nucleus. It appears now that Fe – S cluster formation is initiated in mito-
chondria and all mitochondrial ISC - assembly proteins are required for the
biosynthesis of the cytosolic and nuclear proteins. A current picture includes
an ISC - export machinery in the inner mitochondrial membrane and a CIA
machinery in the cytosol (Figure 6.8 ). Components of each have been identi-
fi ed in yeast by genetic approaches. A mutation in the ATM1 gene was found
to cause leucine auxotrophy due to the absence of an essential cytosolic
enzyme (and Fe – S protein) in the leucine maturation pathway. ATM1 encodes
an ATP binding cassette transporter in the inner mitochondrial membrane.
Further support for a role of this transporter comes from the discovery that
mutations in the corresponding human gene ABCB7 are the cause of X - linked
sideroblastic anemia and cerebellar ataxia, an iron storage disease. It is hypoth-
esized that an Fe – S cluster produced by the ISC - assembly machinery is trans-
ferred to a stabilizing factor that is then exported by the Atm1 transporter,
but the precise nature of the exported substrate remains to be established.
Indirect experiments have suggested that the exported Fe - cluster is stabilized
by a peptide. The Erv1p in the export machinery is a sulfhydryl oxidase local-
ized in the intermembrane space, and deletion of the ERV1 gene is lethal.
Another essential component of the export machinery is glutathione, but its
precise role is still unknown.
The cytosolic machinery for Fe – S protein biosynthesis (CIA) includes
several components identifi ed only recently. The highly conserved eukaryotic
proteins Cfd1p and Nbp1p belong to a class of soluble P - loop NTPases, and
the Nar1p is related to the bacterial iron - only hydrogenases. Many details of
the molecular mechanism for cytosolic and nuclear Fe – S protein assembly will
be the subject of future experiments.
BIOSYNTHESIS OF Fe – S CENTERS 327
328 METABOLIC PATHWAYS INSIDE MITOCHONDRIA
Lill and colleagues have raised the question, Why is Fe – S protein matura-
tion essential in yeast, even when glucose is abundant for fermentation and rich
media provide all (?) essential metabolites? They conclude that the only
known essential yeast Fe – S protein is the cytoplasmic Rli1p, required for ribo-
some biogenesis and function. It is one of the most highly conserved proteins
in evolution and likely to be equally essential in other eukaryotes. This leads to
the further proposal that the only absolutely indispensable function of mito-
chondria in yeast is Fe – S protein biogenesis. What about eukaryotes that lack
mitochondria? Such organisms ( Giardia intestinalis, Trichomonas vaginalis,
cryptosporidium parvum ; see chapter 2 ) have been shown to harbor subcellular
organelles called hydrogenosomes or mitosomes. When complete genomic
sequences were available, genes for oxidative phosphorylation and reactions of
the citric acid cycle were missing. However, numerous genes encoding proteins
of the mitochondrial ISC - assembly complex were found by amino acid sequence
comparisons, suggesting that an ISC - like complex is present in these organisms.
Furthermore, some of these proteins have now been localized in the hydro-
genosomes of T. vaginalis and in mitosomes of G. intestinalis , and it is a good
guess that these organelles are the site of Fe – S cluster assembly. Most, if not all,
of the Fe – S clusters will have to be exported for incorporation into cytosolic
proteins, since no Fe – S proteins are known in mitosomes. A consensus is
emerging that these organelles are derivatives of ancestral mitochondria. They
appear to have evolved independently several times during the evolution of
organisms adapting to anaerobic environments (45 – 47) . It may be concluded
that the biosynthesis of Fe – S clusters is the most important function of mito-
chondria that must be conserved under all conditions.
A highly succinct summary of the most signifi cant features of Fe – S cluster
biogenesis has also been provided by Johnson et al. (40) :
•
Participation of cysteine desulfurases in the mobilization of sulfur
•
A requirement for molecular scaffolds for assembly of [Fe – S] clusters
•
The participation of molecular chaperones
•
Intact cluster transfer from scaffolds to target proteins
•
Specialized [Fe – S] cluster biosynthetic machinery for specifi c targets
•
Specialized [Fe – S] cluster biosynthetic machinery to accommodate stress
conditions
•
Feedback regulation of [Fe – S] cluster biosynthesis and regulation in
response to environmental conditions
•
Compartmentalization of [Fe – S] cluster biosynthetic machinery
•
Intracellular traffi cking of preformed [Fe – S] clusters
•
Mechanisms for the protection or repair of [Fe – S] proteins (not discussed
in this review)
Many details of each of these mechanisms, along with the genes/proteins
involved, can be expected to emerge in the years to follow. It can hoped that
TRANSPORT OF SMALL SOLUTES INTO AND OUT OF MITOCHONDRIA 329
studies will succeed in reconstituting major steps in vitro with isolated and
pure components to gain insight into the precise interaction of various enzymes,
scaffolds, chaperones, and transporters.
6.9 TRANSPORT OF SMALL SOLUTES INTO
AND OUT OF MITOCHONDRIA
6.9.1 Introduction
Although numerous references to the transport of various ions across the
mitochondrial inner membrane have been made in the sections above, it seems
worthwhile to focus on this problem once more as a distinct problem. The
following ions and small solutes feature prominently in discussions of mito-
chondrial functions: H
+
, P
i
, ADP/ATP, Fe
2+
, Cu
+
, Zn
2+
, Ca
2+
, NAD
+
/NADH, and
a large variety of small organic compounds that are metabolites associated
with biochemical reactions. The fi rst three have received a maximum of atten-
tion because of their role in oxidative phosphorylation and the Mitchell
hypothesis. Fe
2+
, Cu
+
, and Zn
2+
ions have been encountered as constituents in
the electron transport chain (cytochromes, iron – sulfur centers, complex IV)
and in metallo - proteases. Mitochondria are prominently mentioned in the
context of the regulation of intracellular calcium; they may constitute a reser-
voir or buffer, and/or their activities may themselves be regulated by Ca
2+
ions
(48, 49) . NAD
+
/NADH are not transported freely across mitochondrial mem-
branes, and most biochemistry textbooks emphasize the importance of shuttle
systems in the transport of reducing equivalents into mitochondria. Never-
theless, there is an intra - mitochondrial pool of NAD
+
/NADH, and concentra-
tions may or may not be similar to those in the cytosolic pool; a limited
transport of this cofactor must be possible. Finally, amino acids, fatty acids,
acetate, glycerol - phosphate, keto acids, and many other metabolites must be
able to enter or exit mitochondria. It seems appropriate to consider these
transport systems and shuttles in the present chapter .
The fl ux of protons through the inner membrane has been discussed exten-
sively in the previous sections. Protons are pumped out of the matrix by the
operation of the electron transport chain, and they return via complex V and
through various “ leakage ” channels, uncouplers, symporters, and antiporters.
In in vitro systems, protons can be exchanged for potassium ions by the use
of the ionophore Nigericin. The elucidation of the precise path of a proton
through the various complexes remains a challenging problem. An interesting
mitochondrial proton pump that has received less attention (at least in text-
books) is the nicotinamide nucleotide transhydrogenase. The bovine enzyme
is an integral membrane protein made up of two identical monomers of
109,065 kDa. An N - terminal domain (430 residues) and a C - terminal domain
(200 residues) fl ank a hydrophobic central domain that can be predicted to
have 14 transmembrane helices. The hydrophilic end - domains extend into the
330 METABOLIC PATHWAYS INSIDE MITOCHONDRIA
matrix, where the N - terminal domain binds NAD(H), and the C - terminal
domain binds NADP(H). In the process of transhydrogenation involving a
hydride transfer from one cofactor to the other, protons are pumped across
the inner membrane.
NADH NADP nH NAD NADPH nH
out
+
in
+
++⇔+ +
As discussed in more detail in a review by Hatefi and Yamaguchi (50) , this
transhydrogenase is a unique proton pump and a potentially powerful model
system to understand how a simple protein can translocate protons across a
membrane driven by conformational transitions that in turn are induced by a
chemical reaction on one side of the membrane and by the different binding
energies for substrates and products.
6.9.2 Porin Alias VDAC
It is generally assumed that pores in the outer membrane formed by an abun-
dant mitochondrial protein named VDAC (or porin) make the outer mem-
brane highly permeable to most small molecules ( < 4000 – 5000 kDa). Thus, the
inner mitochondrial membrane is the major permeability barrier between
the cytosol and the mitochondrial matrix, and the outer membrane is
usually ignored when considering small molecules. VDAC stands for “ voltage -
dependent anion channel, ” and this term immediately becomes confusing if
one has the image of “ pores ” with a fi xed diameter pore size. There is a generic
defi nition of porins as membrane proteins that form a channel through which
molecules of limited size can diffuse passively. They are found prominently in
bacterial outer membranes forming β - barreled structures. Most of the porin
types form trimeric structures, and they exhibit some specifi city so that a variety
of porins are typically found in a bacterial outer membrane. In many discussions
a β - barreled protein (VDAC) in the mitochondrial outer membrane is consid-
ered to be very similar, but it should be pointed out that the mitochondrial
porin - like protein has no primary sequence homology to bacterial porins. In
contrast to the bacterial form, it is believed that a single monomer can form the
functional channel, although aggregation in the membrane has been observed
(see reference 51 ). There are still arguments over whether association of mono-
mers (dimers or tetramers) is necessary for forming a channel.
Crystal structures for bacterial porins have strongly infl uenced the specula-
tions about the VDAC channel and its selectivity. The latest interpretation
based on protease studies, peptide - specifi c antibodies, and site - directed muta-
genesis suggests a channel composed of 13 β strands and one α helix (51) . The
authors call the single α helix “ aesthetically unappealing ” but rationalize it by
the need to have a gating mechanism associated with a large conformational
change.
VDAC proteins have been reconstituted into planar lipid bilayers for elec-
trophysiological studies of single - channel conductance, ion selectivity, and
