Klipp E., Herwig R., Kowald A., Wierling C., Lehrach H. Systems Biology in Practice: Concepts, Implementation and Application
Подождите немного. Документ загружается.

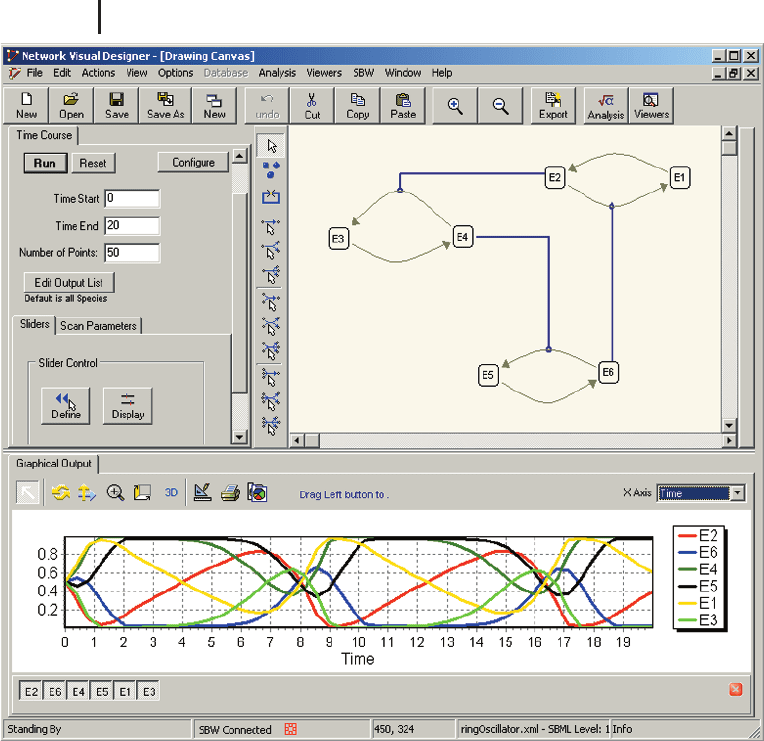
six species (E1 to E6) that are connected via six reactions. JDesigner allows changing
the shape, color, and position of the graphical elements to achieve a visually pleasing
result. But if we are actually interested in the dynamics of the system, we have to de-
fine initial concentrations for the species and kinetic laws for the reactions. JDe-
signer has a list of predefined kinetic laws that can be assigned to a reaction, but it
also allows defining arbitrary custom kinetics. The model can then be saved in
SBML Level 1 or 2. However, the current version of JDesigner, 1.934, seems to have
subtle problems with the produced SBML. Unfortunately, none of the supplied
430
14 Modeling Tools
Fig. 14.5 JDesigner is an SBW-enabled applica-
tion that allows the visual construction of reac-
tion networks. The upper right part of the
screenshot shows a network of six species cycli-
cally connected by six reactions. Connection to
the time course service of the Jarnac module
brings up the control panel on the upper left
part. After Jarnac has finished the simulation,
the results are passed back to JDesigner, where
they are displayed in the bottom part of the win-
dow.

SBML example models passes the online syntax validation that is available at http://
sbml.org/tools/htdocs/sbmltools.php, which prevents, for instance, the transfer of
models from JDesigner to CellDesigner.
Like all SBW-enabled programs, JDesigner has a special menu called SBW that
lists the services provided by other SBW modules. Services provided with the stan-
dard distribution are stochastic time course simulations, deterministic time course
and steady-state calculations, and the generation of a Matlab ODE function file of
the current model. However, JDesigner also offers another, tighter, integration with
Jarnac, a network simulation module. The menu entry “Actions/Connect Simulator”
establishes a direct connection to Jarnac, which in turn enables several entries in the
Analysis menu. The left side of Fig. 14.5 shows the control panel that appears if the
Time Course Simulation is chosen from this menu. After the simulation is finished,
a graphical representation of the solution can be examined (bottom of Fig. 14.5). The
graphical output is very flexible and allows selection of the variables to be plotted, of
the axes labels and scaling, and of titles and legends. Furthermore, the result can be
saved in different graphic formats and the numerical values can be exported as text,
XML, or Excel format. Finally, a good tutorial on how to use JDesigner is available
from http://public.kgi.edu/~hsauro/sysbio/papers/JDBooklet.pdf.
14.1.5.2 CellDesigner
CellDesigner, another network creation tool, might be an alternative to JDesigner for
people who do not use Microsoft Windows (Funahashi et al. 2003). The application
is written in Java and therefore also runs under MacOS X and Linux. The current
version is 2.0 and can be downloaded from http://systems-biology.org. As with JDe-
signer, networks can be constructed from compartments, species, and reactions.
However, CellDesigner comes with a large number of predefined shapes that can be
used for different types of molecules such as proteins, receptors, ion channels, small
metabolites, etc. It is also possible to modify the symbols to indicate phosphoryla-
tions or other modifications (Fig. 14.6, center). The program also provides several
icons for special reaction types such as catalysis, transport, inhibition, and activa-
tion.
Reading and writing of the models is SBML-based (Level 1 and 2), and the models
written by CellDesigner pass the online validation at http://sbml.org/tools/htdocs/
sbmltools.php and thus conform to the SBML standard. A nice feature in this re-
spect is the ability to display the SBML model structure as a tree (Fig. 14.6, left side).
A click on a species or reaction in this tree highlights the corresponding elements in
the graphics canvas and in the matching tab on the right side showing further de-
tails. This tab is also the place where initial concentrations and reaction details are
entered. Like JDesigner, CellDesigner allows entering arbitrary kinetic equations,
but unfortunately it has no list of standard kinetics (mass action or Michaelis-Men-
ten) that can be applied. For each reaction, the rate law has to be typed in by hand.
A connection to SBW is realized via the standard SBW menu and provides the ser-
vices described together with JDesigner. For a further introduction to CellDesigner, a
startup guide can be obtained from the download page at http://systems-biology.org.
431
14.1 Modeling and Visualization
432
14 Modeling Tools

14.1.6
Petri Nets
A Petri Net is a graphical and mathematical modeling tool for discrete and parallel
systems. The mathematical concept was developed in the early 1960s by Adam Petri.
The basic elements of a Petri Net are places, transitions, and arcs that connect places
and transitions. When represented graphically, places are shown as circles and tran-
sitions as rectangular bars (see Fig. 14.7). Places represent objects (molecules, cars,
machine parts), and transitions describe whether and how individual objects are in-
ter-converted. Places can contain zero or more tokens, indicating the number of ob-
jects that currently exist. The tokens are shown as dots in Fig. 14.7a (one in place P1
and two in place P3). Whether a transition can take place (can fire) or not depends
on the places that are connected to the transition by incoming arcs to contain
enough tokens. If this condition is fulfilled, the transition fires and changes the state
of the system by removing tokens from the input places and adding tokens to the
output places. The number of tokens that are removed and added depends on the
weight of the arcs. The movement of tokens during successive firing events is called
the token game.
Petri Nets are not only an optically pleasing representation of a system but they
can also be described mathematically in terms of integer arithmetic. For simple types
of Petri Nets, certain properties can thus be calculated analytically, but often the net
has to be run to study the long-term system properties. Over the years many exten-
sions to the basic Petri Net model have been developed for different simulation pur-
poses (Bernardinello and de Cindio 1992):
1. Hybrid Petri Nets that add the possibility to have places that contain a continuous
token number instead of discrete values.
2. Timed Petri Nets that extend transitions to allow for a specific time delay between
the moment when a transition is enabled and the actual firing.
3. Stochastic Petri Nets that go one step further and allow a random time delay
drawn from a probability distribution.
4. Hierarchical Petri Nets, in which modularity is introduced by representing whole
nets as a single place or transition of a larger net.
5. Colored Petri Nets that introduce different types (colors) or tokens and more com-
plicated firing rules for transitions.
With these extensions Petri Nets are powerful enough to be used for models in
systems biology. Biochemical pathways can be modeled with places representing me-
433
14.1 Modeling and Visualization
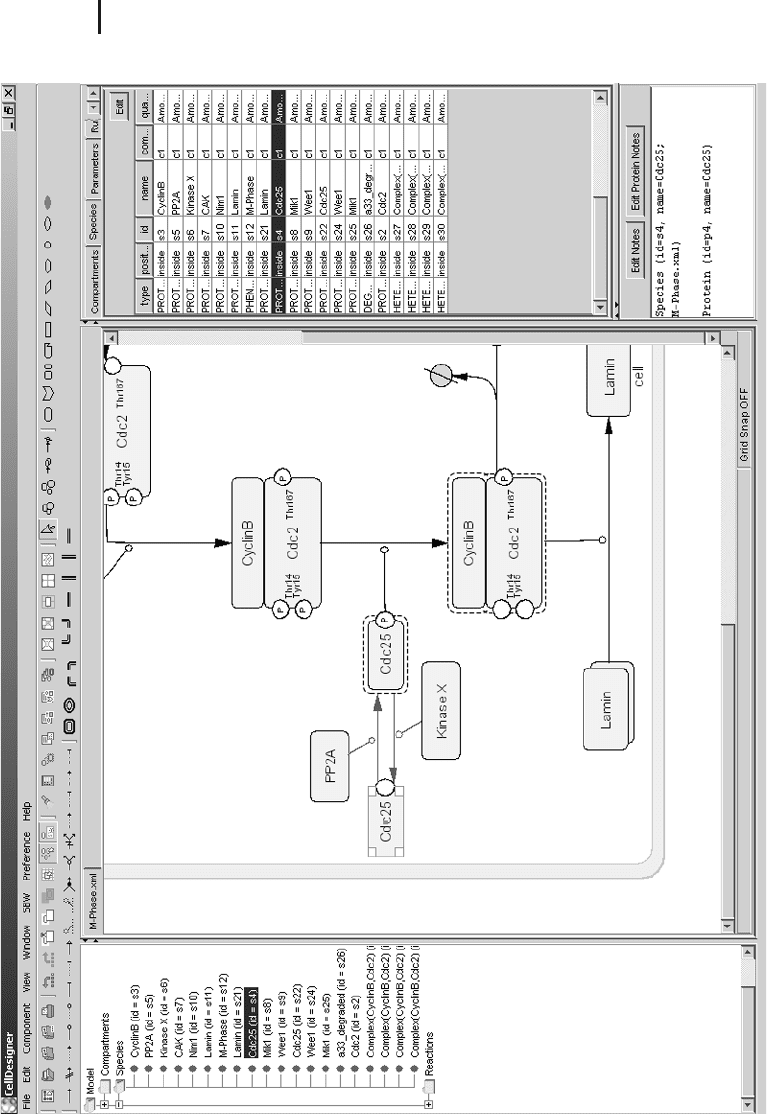
3 Fig. 14.6 CellDesigner 2.0 is another program
that is SBW aware and can be used to construct
reaction networks that can be analyzed by other
SBW modules or saved in SBML format. The
special strength of CellDesigner is its advanced
graphical representation skills. Different shapes
are available for different types of molecules,
such as ions, proteins, receptors, genes, DNA,
or RNA. Another highlight is the model tree (left
side) that provides a convenient overview of the
SBML structure and components.
434
14 Modeling Tools
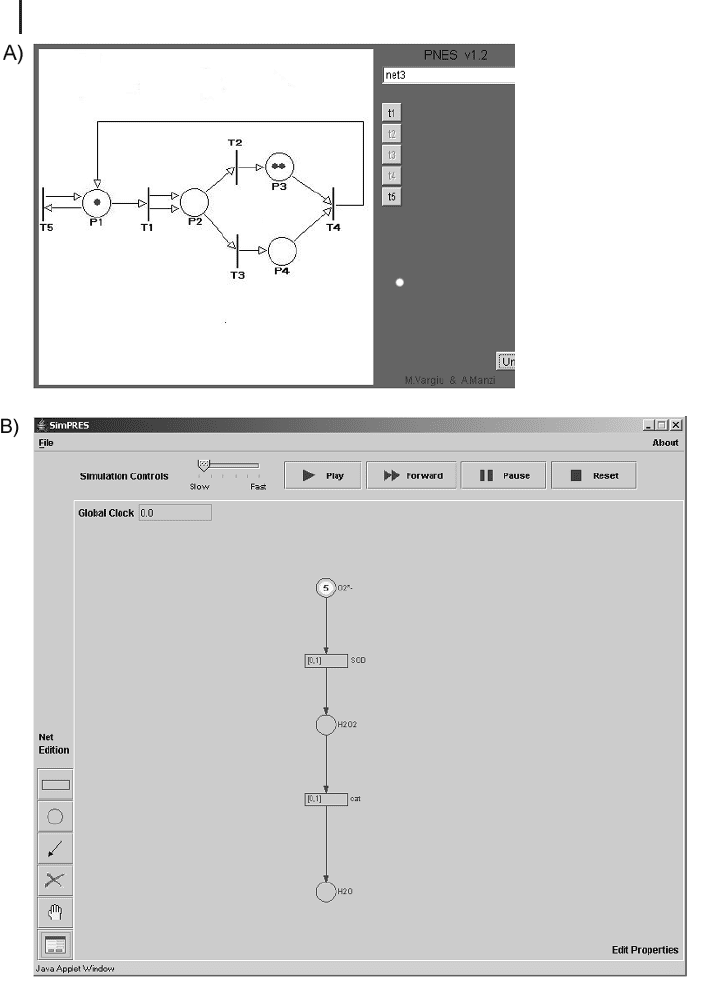
Fig. 14.7 Several Java applets exist to get
started with Petri Nets. (a) For a very first con-
tact, PNES at http://web.tiscali.it/marconthe-
net/Pnes.Pnes.html is a good choice. It contains
numerous predefined nets, and by firing enabled
transitions manually, the network behavior can
be studied. (b) A simple applet that also allows
users to construct and run their own Petri Nets
can be found at http://www.ida.liu.se/~luico/
SimPRES. It allows one to specify different time
delays for transitions, and thus simple Timed
Petri Nets can be created.

tabolites and transitions representing reactions, and stoichiometric coefficients are
encoded as different weights of input and output arcs. Consequently, Petri Nets have
been used to model metabolic networks (Reddy et al. 1996; Küffner et al. 2000) and
signal transduction pathways (Matsuno et al. 2003). Many free and commercial tools
are available to explore the behavior of Petri Nets. The Petri Nets World homepage
(http://www.daimi.au.dk/PetriNets) is an excellent starting point for this purpose. It
contains a large amount of information about tutorials, software, and conferences as
well as a large bibliography regarding Petri Nets. In addition to comprehensive stan-
dalone packages, there are also a number of Petri Nets available that have been im-
plemented as Java applets. The Petri Net Educational Simulator (PNES) at http://
web.tiscali.it/marconthenet/Pnes.Pnes.html is very limited because it displays only a
collection of predefined nets, but by stepping through the net (playing the token
game) one can get a first impression. Figure 14.7 a shows the user interface of
PNES. A more advanced applet (can also be installed as application) that also allows
the creation and modification of Petri Nets is SimPRES (http://www.ida.liu.se/
~luico/SimPRES). Figure 14.7 b shows the applet with our well-known reaction sys-
tem from superoxide to water (see Section 14.1.1) implemented as Petri Net.
However, these applets can run only simple Petri Nets, which are restricted in
their computational capabilities. Consequently, the radical detoxification reactions
shown in Fig. 14.7b lack realism. The number of superoxide radicals that is con-
verted into hydrogen peroxide is constant per unit time. A more realistic assumption
is that the reaction follows the mass action (as done in Sections 14.1.1 and 14.1.2).
But for such calculations larger packages are needed, e.g., CPN Tools (http://wiki.-
daimi.au.dk/cpntools), which is an actively developed Colored Petri Net tool devel-
oped at the university of Aarhus, Denmark, or the commercially available Cell Illus-
trator (http://www.gene-networks.com/ci), which is based on a Hybrid Petri Net.
14.1.7
STOCKS 2
The stochastic simulation tool STOCKS 2 was developed by Andrzej Kierzek and Ja-
cek Puchalka and is available free of charge under the GNU GPL license (http://
www.sysbio.pl/stocks/). Versions are available for Microsoft Windows and Linux.
Simulations using differential equations assume that the number of molecules in
the described system is so large that it can be treated as a continuous variable. A sec-
ond assumption of ODE modeling is that the system is completely deterministic. Ran-
dom fluctuations do not occur. The smaller the number of molecules, the more unrea-
listic those assumptions become. Most molecules in a cell exist in large numbers, but
some are very rare. Transcription factors, for example, exist in notoriously low num-
bers in cells. There are only approximately 10 molecules of the Lac repressor in an
E. coli cell (Levin 1999). Proteins involved in signal transduction pathways are also
very rare, as are defective mitochondria that might be relevant for the aging process
(see Section 7.3.2). Under those circumstances it becomes important that four or five
molecules might be in a cell, but not fractional amounts like 4.325 (as assumed by
ODEs). Of special importance can be the difference between zero and one items if this
435
14.1 Modeling and Visualization

item is a self-reproducing object like a defective mitochondrion. If modeled with
ODEs it is practically impossible to obtain zero defective mitochondria; a small
amount (well below one) will always remain. Because of their self-reproducing prop-
erty, a population of defective mitochondria could always regrow from this artifact. If
modeled stochastically, all defective organelles disappear (zero concentration) if the
last one is destroyed, and thus they cannot regrow. If the simulated system has more
than one possible steady state, there can also be qualitative differences between a deter-
ministic simulation with differential equations and a stochastic simulation that takes
random effects into account. In an ODE model the system will settle into one of the
possible steady states and remain there forever. If modeled stochastically, however, the
system can jump from one steady state to the other if they are close enough together.
As discussed in Section 3.6, the basic algorithms used for stochastic simulations
are Gillespie’s first reaction method and direct method (Gillespie 1977) or the more
efficient, recently developed, next reaction method (Gibson and Bruck 2000). How-
ever, all these methods have problems to model systems that contain both intensive
reactions involving molecule species present in large numbers and reactions invol-
ving rare molecules. The majority of computation time is used for the intensive reac-
tions, which occur in very short time intervals and thus are called fast reactions. It
therefore often requires an unacceptably long computation time before the interest-
ing slow reactions are simulated.
STOCKS 2 (Puchalka and Kierzek 2004) is based on the idea of modeling fast reac-
tions with an approximate method (Gillespie 2001) and the slow reactions with the ex-
act next reaction method. The assignment of reactions into slow and fast is done auto-
matically and dynamically. This means that if a molecular species increases in num-
ber during the course of a simulation, which in turn increases the reaction rate,
STOCKS 2 automatically moves it from the list of slow reactions to the group of fast
reactions. The program uses as input a model description in SBML (Section 14.2.2)
and generates as output text files containing the molecule numbers of the different
variables at user-specified time points. SBML is a very bulky and verbose format, and
hence writing an SBML model description by hand is a painful process. Luckily an in-
put editor is provided to create the model and the SBML file (Fig. 14.8a). The docu-
mentation is, at the time of this writing, rather sparse, but the buttons and associated
dialog boxes roughly have to be processed from top to bottom. “Model Parameters” al-
lows one to enter several details relevant for the actual simulation, such as the simula-
tion time, the number of repetitions of an experiment (to get a feeling for the spread
of individual trajectories), the time interval for saving output, and whether to save de-
bug information (very handy during model development). “Parameters” leads to the
definition of constants for the model, the next button allows one to define compart-
ments of different sizes, and in “Species” the used variables have to be declared. The
most important part is finally the definition of the reactions. It is possible to choose
from a variety of different kinetics types or to define an arbitrary kinetics. However,
this has to be entered in MathML format, which is discussed in Section 14.2.3. The
last two buttons are for advanced topics and will not be discussed here.
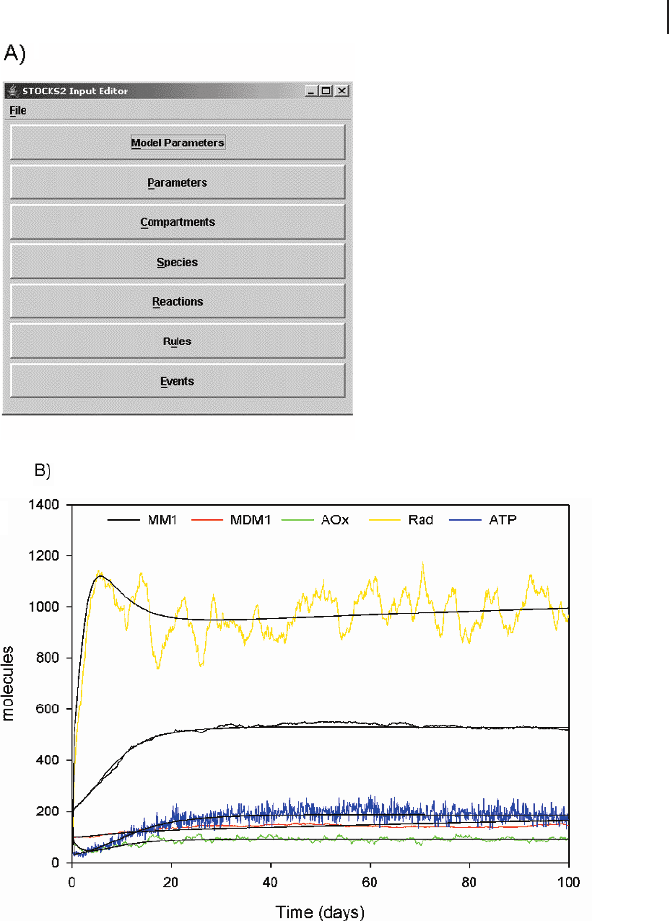
The output of STOCKS 2 is a plain text file, but for illustrative purposes Fig. 14.8b
compares the processed stochastic results of an example model (accumulation of de-
436
14 Modeling Tools
437
14.1 Modeling and Visualization
Fig. 14.8 STOCKS 2 is a simulation tool for sto-
chastic models. (a) Models can be constructed
with the Input Editor, which generates the SBML
format used by STOCKS 2. Each of the seven
buttons opens dialog boxes that allow one to en-
ter simulation parameters, variables, reactions,
compartments, etc. (b) Comparison of the sto-
chastic trajectories generated by STOCKS 2 and
the corresponding ODEs of an example model
describing the population dynamics of mito-
chondria.

fective mitochondria, similar to Section 7.3.2) with the solutions of a corresponding
differential equations model. In this case there is excellent agreement between the
two types of simulation. We would like to recommend this dual modeling approach
since the ODEs can be used to derive the stochastic reactions, a simulation of the ODE
system can give a quick and rough feeling for the model behavior, and disagreement
between the two approaches can often be used to spot implementation errors of the
stochastic model. If there are differences they have to be understandable by the special
above-mentioned differences between the deterministic and stochastic approaches.
14.1.8
Genetic Programming
Genetic programming (Koza 1992) is a variation of genetic algorithms (Holland
1975). This means that it uses techniques borrowed from evolution. A solution to a
problem is obtained by letting many different versions of a program compete against
each other. From one generation to the next, they can undergo mutations and recom-
bination and propagate according to their fitness. The better a variant solves the
problem at hand, the higher its fitness is. The main difference between genetic pro-
gramming (GP) and genetic algorithms lies in the way the solution is presented. GP
produces a ready-to-use program in a Lisp-like presentation, while genetic algo-
rithms create a string of numbers that represents the solution.
As an example, assume that the problem at hand is to find the mathematical func-
tion that most accurately approximates the data points shown in Fig. 14.9a. The criti-
cal point is to find a way to represent the solution (a mathematical function) as a tree-
like structure as seen in Fig. 14.9b. Such a tree consists of internal nodes, consisting
of problem-specific functions (here mathematical operators), and terminal nodes,
consisting of the input for the functions. Each tree represents one individual of the
competing population. An appropriate measure of fitness is another important point
for successfully applying GP. In this case the sum of squares of the distance between
the data points and the corresponding function values could be used. New individuals
can be generated by mutating nodes (replacement of one operator by another or modi-
fication of a numerical constant) or by recombination between two individuals (ex-
change of sub-branches). If those modifications follow certain syntactical constraints,
it can be guaranteed that syntactically correct mutants are always generated.
Genetic programming has been used successfully in such diverse areas as the
automatic creation of the topology and sizing of analogue electrical circuits (Koza
et al. 1999), the design of antennas according to high-level specifications (Comisky
et al. 2000), and the reverse engineering of metabolic pathways (Koza et al. 2000,
2001). The last example is especially interesting because it relates directly to systems
438
14 Modeling Tools
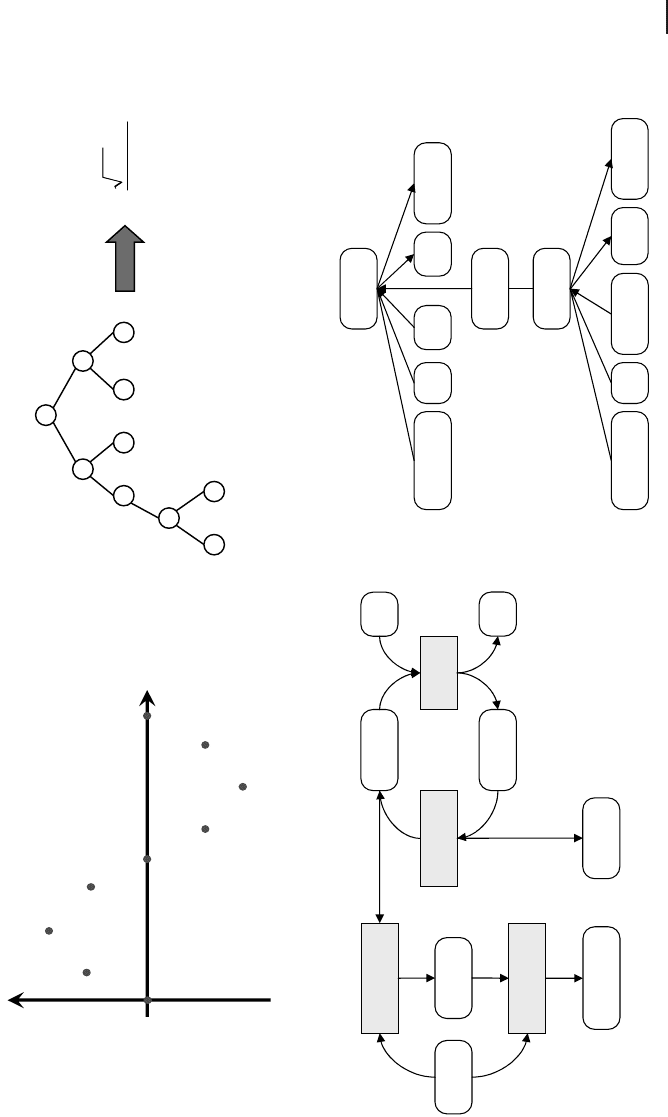
Fig. 14.9 (a) Function to data fitting as simple
example of Genetic Programming (GP). (b) Pos-
sible solutions to the problem (mathematical
functions) are represented as a tree. (c) The
structure and numerical constants of this meta-
bolic reaction network are to be reverse engi-
neered by GP using only concentration values of
diacyl-glycerol for different enzyme concentra-
tions. (d) Tree-like representation of two of the
four enzyme reactions seen in (c).
"
439
14.1 Modeling and Visualization
2x 5
2x
y
-
*
2
x
2
x
*
5
acylglycerol
lipase
triacylglycerol
lipase
glycerol-1
phosphatase
glycerol
kinase
fatty acid
diacyl-glycerol
monoacyl
glycerol
glycerol
glycerol-3
phosphate
ATP
ADP
ortho
phosphate
glycerol-1
phosphatase
1.19
glycerol-3
phosphate
glycerol
ortho
phosphate
Rtype-1-2
first product
glycerol
kinase
1.69 ATP ADP
glycerol-3
phosphate
Rtype-2-2
A)
B)
C)
D)
x
