Klipp E., Herwig R., Kowald A., Wierling C., Lehrach H. Systems Biology in Practice: Concepts, Implementation and Application
Подождите немного. Документ загружается.


In a DAG, each node can have multiple child nodes, as well as multiple parent
nodes. Cyclic references, however, are forbidden. The combination of vocabulary and
relationship between nodes is referred to as ontology. At the root of the GO are the
three top-level categories, molecular function, biological process, and cellular compo-
nent, which contain many levels of child nodes (GO terms) that describe a gene pro-
duct with increasing specificity. The GO consortium, in collaboration with other da-
tabases, develops and maintains the three top-level ontologies (the set of GO terms
and their relationship) themselves, creates associations between the ontologies and
the gene products in the participating databases, and develops tools for the creation,
maintenance, and use of the ontologies.
Let’s look at a practical example to see how the concept works. The enzyme superox-
ide dismutase, for instance, is annotated in FlyBase (the Drosophila melanogaster data-
base) with the GO term “cytoplasm” in the cellular component ontology, with the GO
terms “defense response” and “determination of adult lifespan” in the biological pro-
cess ontology, and with the terms “antioxidant activity” and “copper, zinc superoxide
dismutase activity” in the molecular function ontology. The GO term cytoplasm itself
has the single parent “intracellular,” which has the single parent “cell,” which is finally
connected to the cellular component. The other GO terms for superoxide dismutase
are connected in a similarly hierarchical way to the three top categories.
The following table gives the number of gene products that have been annotated
to the top-level categories of the GO for several popular databases. The table dates
from January 2004 and excludes annotations that are based exclusively on electronic
inferences.
Database Biological process Molecular function Cellular component
SGD
Saccharomyces cerevisiae
6455 6438 6441
FlyBase
Drosophila melanogaster
4974 7084 4291
MGI
Mus musculus
6634 7127 8175
TAIR
Arabidopsis thaliana
13,962 6449 11746
WormBase
Caenorhabditis elegans
3903 410 756
RGD
Rattus norvegicus
1436 1833 745
Gramene
Oryza sativa
5459 2770 16,628
ZFIN
Danio rerio
413 315 296
DictyBase
Dictyostelium discoideum
410 426 5383
GO Annotations@EBI
Human
8109 7585 7037
400
13 Databases and Tools on the Internet

To use the Gene Ontology effectively, many different tools have been developed
that are listed on the GO Web site (http://www.geneontology.org/GO.tools.html).
The repertoire encompasses Web-based and standalone GO browsers and editors,
microarray-related tools, and programs for many specialized tasks. In the remainder
of this section, we will have only a quick look at three such tools. Our first candidate
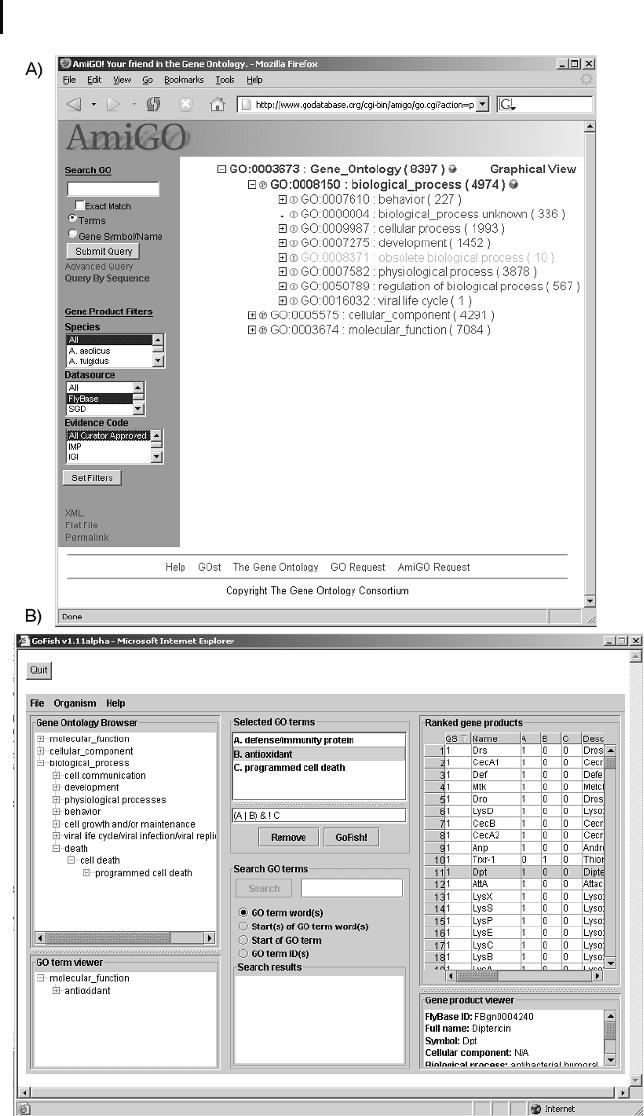
is AmiGO (http://godatabase.org/), which is a Web-based GO browser maintained
by the GO consortium (Fig. 13.1 a). First of all, AmiGO can be used to browse the
terms of the ontologies. The numbers in parenthesis behind the GO terms show
how many gene products in the currently selected database are annotated to this
term. The seven-digit number behind GO is the GO-ID that links each GO term to a
unique identifier. One or more species and one or more data sources can be selected
to restrict the results of the search. A click on a leaf of the GO hierarchy (such as bio-
logical_processes unknown) brings up a window that lists the genes in the selected
databases that have been annotated to this term. Instead of browsing the GO tree,
one can also search for specific GO terms and get to the gene products associated
with these terms or search for gene products and find the connected GO terms. Fi-
nally, it is also possible to get a graphical view, which shows where the selected term
is located within the ontology tree.
AmiGO’s search options are quite limited since it is only possible to search for
several terms that are connected via the OR function (under advanced query). The
Java applet GoFish v1.11 (http://llama.med.harvard.edu/~berriz/GoFishWelco-
me.html) is a good alternative for such cases (Fig. 13.1b). Different GO terms can be
selected from a GO tree (left side), which can then be combined into complex Boo-
lean expressions (window in the middle). In the case shown here, we are searching
for gene products in the FlyBase that are antioxidants or are involved in defense/im-
munity but are not concerned with programmed cell death. When the user selects a
specific database, the applet downloads the GO terms and associations for this data-
base, and therefore the response time of GoFish is normally faster than for AmiGO.
Recently developed high-throughput techniques such as DNA chips enable re-
searchers to measure the expression profile of thousands of genes in parallel. Often,
one is interested in whether the genes that are over- or underexpressed share biologi-
cal functions. The Web-based program GOstat (http://gostat.wehi.edu.au/) makes
use of the Gene Ontology to test whether certain GO terms are statistically over- or
underrepresented in a group of genes of interest. GOstat can compare the list of
genes either against a user-supplied control list or against the complete list of genes
in a selected database. It then uses Fisher’s exact test or the Chi-square test to deter-
mine whether the observed differences of the frequencies of GO terms are signifi-
cant or not (cf. Section 9.4). The output is a list of P-values that state how specific the
associated GO terms are for the list of genes provided. The list can then be obtained
as text or as an HTML file.
This was only a brief introduction of three of the many available programs that
make use of GO annotations. Furthermore, it is always possible to develop one’s
own programs, since all GO-related files such as lists of GO definitions or database
annotations can be downloaded at http://www.geneontology.org.
401
13.1 Gene Ontology
402
13 Databases and Tools on the Internet

13.2
KEGG
KEGG (Kyoto Encyclopedia of Genes and Genomes; http://www.genome.ad.jp/
kegg/) is a reference knowledgebase offering information about genes and proteins,
biochemical compounds, reactions, and pathways. The data are organized in three
parts: the gene universe (consisting of the GENES, SSDB, and KO databases), the
chemical universe (with the COMPOUND, GLYCAN, REACTION, and ENZYME da-
tabases which are merged as the LIGAND database), and the protein network con-
sisting of the PATHWAY database (Kanehisa et al. 2004). In addition, the KEGG da-
tabase is hierarchically classified into categories and subcategories at four levels. The
five topmost categories are metabolism, genetic information processing, environ-
mental information processing, cellular processes, and human diseases. Subcate-
gories of metabolism are, e. g., carbohydrate, energy, lipid, nucleotide, or amino acid
metabolism. These are subdivided into the different pathways, such as glycolysis, ci-
trate cycle, purine metabolism, etc. Finally, the fourth level corresponds to the KO
(KEGG Orthology) entries. A KO entry (internally identified by a K number, e. g.,
K00001 for the alcohol dehydrogenase) corresponds to a group of orthologous genes
that have identical functions.
The gene universe offers information about genes and proteins generated by gen-
ome sequencing projects. Information about individual genes is stored in the GENES
database, which is semiautomatically generated from submissions to GenBank, the
NCBI RefSeq database, the EMBL database, and other publicly available organism-
specific databases. K numbers are further assigned to entries of the GENES database.
The SSDB database contains information about amino acid sequence similarities be-
tween protein-coding genes computationally generated from the GENES database.
This is carried out for many complete genomes and results in a huge graph depicting
protein similarities with clusters of orthologous and paralogous genes.
The chemical universe offers information about chemical compounds and reac-
tions relevant to cellular processes. It includes more than 11,000 compounds (intern-
ally represented by C numbers, e. g., C00001 denotes water), a separate database for
carbohydrates (nearly 11,000 entries; represented by a number preceded by G, e.g.,
G10481 for cellulose), more than 6000 reactions (with R numbers, e.g., R00275 for
the reaction of the superoxide radical into hydrogen peroxide), and more than 4000
enzymes (denoted by EC numbers as well as K numbers for orthologous entries). All
these data are merged as the LIGAND database (Goto et al. 2002). Thus, the chemi-
cal universe offers comprehensive information about metabolites with their respec-
tive chemical structures and biochemical reactions.
403
13.2 KEGG
3 Fig. 13.1 (a) AmiGO, a Web-based GO browser
developed by the GO consortium. It allows
browsing the GO hierarchy and searching for
specific GO terms or gene products in different
databases. The numbers in brackets behind the
GO terms indicate how many gene products
have been annotated to this term in the selected
database. (b) GoFish, a Java applet, can also
connect to several databases and allows the user
to search for gene products using complex Boo-
lean expressions of GO terms.

KEGG’s protein network provides information about protein interactions compris-
ing pathways and protein complexes. The 235 KEGG reference pathway diagrams
(maps) offered on the Web site give clear overviews of important pathways. Organ-
ism-specific pathway maps are automatically generated by coloring of organism-spe-
cific genes in the reference pathways.
The KEGG database can be queried via the Web interface, e.g., for genes, proteins,
compounds, etc. Access to the data via FTP (http://www.genome.ad.jp/anonftp) as
well via a SOAP server (http://www.genome.ad.jp/kegg/soap) is possible for aca-
demic users, too (SOAP is a protocol for the exchange of messages between compu-
ter software based on XML; see Section 14.2.1).
13.3
BRENDA
High-throughput projects, such as the international genome sequencing efforts, ac-
cumulate large amounts of data at an amazing rate. These data are essential for the
reconstruction of phylogenetic trees and gene-finding projects. However, for kinetic
modeling, which is at the heart of systems biology, kinetic data of proteins and en-
zymes are needed. Unfortunately, this type of data is notoriously difficult and time-
consuming to obtain, since proteins often need individually tuned purification and
reaction conditions. Furthermore, the results of such studies are published in a large
variety of journals from different fields.
In this situation, BRENDA aims to be a comprehensive enzyme information sys-
tem (http://www.brenda.uni-koeln.de). Basically, BRENDA is a curated database that
contains a large amount of functional data for individual enzymes. These data are
gathered from the literature and made available via a Web interface. The table on the
next page gives an overview of the types of information that is collected and the
number of entries for the different information fields (as of June 2004). For instance,
enzymes representing 4379 different EC numbers and over 50,000 different K
m
va-
lues are contained in the database.
One of BRENDA’s strengths is the multitude of ways the database can be
searched. It is easy to find all enzymes that are above a specific molecular weight, be-
long to C. elegans, or have a temperature optimum above 308C. If desired, the list of
results can then be downloaded as a tab-separated text file for later inspection. Using
the Advanced Search feature, it is possible to construct arbitrarily complex search
queries involving the information field shown in the table.
Sometimes one wants to search for all enzymes that are glycosylases without know-
ing the corresponding EC number or to find all enzymes that are found in horses
without knowing the exact scientific name. In this situation, the ECTree browser and
the TaxTree search are helpful by providing a browser-like interface to search down the
hierarchy of EC number descriptions or taxonomic names. A similar browser is also
available for Gene Ontology terms,which were discussed in Section 13.1.
BRENDA is also very well connected to other databases that can provide further
information about an enzyme in question. Associated GO terms are directly linked
404
13 Databases and Tools on the Internet

to the AmiGO browser; substrates and products of the catalyzed reactions can be dis-
played as chemical structures; links to the taxonomic database NEWT (http://www.e-
bi.ac.uk/newt) exist for information of the organism; sequence data can be obtained
from Swiss-Prot; and if crystallographic data exist, a link to PDB (see Section 13.8) is
provided. Finally, literature references (including PubMed IDs) from where the pro-
tein data originated are provided.
13.4
Databases of the National Center for Biotechnology
The National Center for Biotechnology (NCBI) (http://www.ncbi.nlm.nih.gov/) pro-
vides several databases that are widely used in biological research. Most important
are the molecular databases, offering information about nucleotide sequences, pro-
teins, genes, molecular structures, and gene expression. Besides this, several data-
bases comprising scientific literature are available. The NCBI also provides a taxon-
omy database that contains names and lineages of more than 130,000 organisms.
For more than 1000 organisms, whole genomes (either already completely se-
quenced or for which sequencing is still in progress) and corresponding gene maps
are available, as well as tools for their inspection. A full overview of the databases
405
13.4 Databases of the National Center for Biotechnology
Information Field Entries Information Field Entries
Enzyme nomenclature Functional parameters
EC number 4379 K
m
value 52,343
Recommended name 4376 Turnover number 9035
Systematic name 3469 Specific activity 22,108
Synonyms 30,475 pH range and optimum 5063/18,866
CAS registry number 4005 Temperature range and optimum 1396/8014
Reaction 3864 Molecular properties
Reaction type 6326 pH stability 3899
Enzyme structure Temperature stability 9124
Molecular weight 18,725 General stability 5847
Subunits 11,666 Organic solvent stability 468
Sequence links 65,249 Oxidation stability 487
Post-translational modifications 2069 Storage stability 8249
Crystallization 1741 Purification 15,102
3D structure, PDB links 11,236 Cloned 6666
Enzyme-ligand interactions Engineering 6190
Substrates/products 82,194 Renatured 346
Natural substrates 33,619 Application 1641
Cofactor 9967 Organism-related information
Activating compound 11,780 Organism 99,502
Metals/ions 16,162 Source-tissue, organ 28,181
Inhibitors 75,584 Localization 11,711
Bibliographic data
References 73,406

provided by the NCBI can be found under http://www.ncbi.nlm.nih.gov/sitemap/in-
dex.html. All these databases are searchable via the Entrez search engine accessible
through the NCBI homepage.
Among the nucleotide sequence databases, the Genetic Sequence database (Gen-
Bank), the Reference Sequences database (RefSeq), and UniGene can be found. Gen-
Bank (Release 143.0, from August 2004) comprises 41.8 billion nucleotide bases
from more than 37 million reported sequences. The RefSeq database (Maglott et al.
2000; Pruitt et al. 2000; Pruitt and Maglott 2001) is a curated, non-redundant set of
sequences including genomic DNA, mRNA, and protein products for important
model organisms. In UniGene, expressed sequence tags (ESTs) and full-length
mRNA sequences are organized into clusters, each representing a unique known or
putative gene of a specific organism. (For molecular biological analyses, e.g., sequen-
cing or expression profiling, the mRNA of expressed genes is usually translated into
a complementary DNA [cDNA, copy DNA], since this is more stable and feasible for
standard biotechnological methods. An EST is a short, approximately 200–600 bp
long, sequence from either side of a cDNA clone that is useful for identifying the
full-length gene, e. g., for locating the gene in the genome.)
In addition to nucleotide sequences, protein sequences can also be searched for at
the NCBI site via Entrez-Proteins. Searches are performed across several databases,
including RefSeq, Swiss-Prot, and PDB.
Finally, the LocusLink database (http://www.ncbi.nlm.nih.gov/LocusLink) offers
diverse information about specific genetic loci (the location of a specific gene). Thus,
LocusLink provides a central hub for accessing gene-specific information for a num-
ber of species, such as human, mouse, rat, zebrafish, nematode, fruit fly, cow, and
sea urchin.
Among the literature databases are PubMed and OMIM (Online Mendelian In-
heritance in Man). PubMed is a database of citations and abstracts for biomedical lit-
erature. Citations are from MEDLINE (http://medline.cos.com) and additional life
science journals. OMIM is a catalog of human genes and genetic disorders with tex-
tual information and copious links to the scientific literature.
Thus, the databases at the NCBI are one of the major resources for sequence data,
annotations, and literature references. They can be used to determine what is known
about a specific gene or its protein or to get information about the sequences and
their variants or polymorphisms. In addition to this, the NCBI also offers a database
on gene expression data (Gene Expression Omnibus, GEO).
Besides all these databases, the NCBI also provides tools mostly operating on se-
quence data. These include programs comparing one or more sequences with the
provided sequence databases.
13.5
Databases of the European Bioinformatics Institute
The European Bioinformatics Institute (EMBL-EBI) also offers several biologically
relevant databases (http://www.ebi.ac.uk/Databases). These include databases on
406
13 Databases and Tools on the Internet

nucleotide sequences, genes, and genomes (EMBL Nucleotide Database, Ensembl
automatic genome annotation database), a database on alternative splicing sites
(ASD), a database of protein modifications (RESID), a database on protein families
and protein domains (InterPro), a database on macromolecular structures (E-MSD),
and a database on gene expression data (ArrayExpress). The protein databases Swiss-
Prot, TrEMBL, and UniProt as well as the Reactome database on pathways and pro-
cesses relevant for humans will be discussed in separate sections below.
13.5.1
EMBL Nucleotide Sequence Database
The EMBL Nucleotide Sequence Database (http://www.ebi.ac.uk/embl) incorpo-
rates, organizes, and distributes nucleotide sequences from public sources and syn-
chronizes its data in a daily manner with the DNA Database of Japan (DDBJ) and
GenBank, which are the two other most important nucleotide sequence databases
worldwide (Kulikova et al. 2004).
13.5.2
Ensembl
The Ensembl project (http://www.ensembl.org/) is developing and maintaining a
system for the management and presentation of genomic sequences and annotation
for eukaryotic genomes (Hubbard et al. 2002; Birney et al. 2004a, 2004b; Hammond
and Birney 2004). What does annotation mean in this context? Annotation is the
characterization of features of the genome using computational and experimental
methods. In the first place, this is the prediction of genes, including structural ele-
ments like introns and exons, from the assembled genome sequence and the charac-
terization of genomic features, like repeated sequence motifs, conserved regions, or
single-nucleotide polymorphisms (SNPs). SNPs (pronounced „snips“) are common
DNA sequence variations among individuals, where a single nucleotide is altered.
Furthermore, annotation includes information about functional domains of the
proteins encoded by the genes and the roles that the gene products fulfill in the or-
ganism.
The central component of Ensembl is a relational database storing the genome se-
quence assemblies and annotations produced by Ensembl’s automated sequence-an-
notation pipeline, which utilizes the genome assemblies and data from external re-
sources for this purpose. In September 2004, Ensembl provided genomic annota-
tions for several vertebrates (human, chimp, mouse, rat, puffer fish, zebrafish, and
chicken), arthropods (mosquito, honeybee, and fruit fly), and nematodes (Caenorhab-
ditis elegans and Caenorhabditis briggsae). Annotations, such as genes with their in-
tron/exon structure, SNPs, etc., can be viewed along the assembled sequence contigs
using the Ensembl ContigView, which is accessible via the organism-specific Web
pages (e.g., for humans, http://www.ensembl.org/Homo_sapiens/).
407
13.5 Databases of the European Bioinformatics Institute

13.5.3
InterPro
InterPro (http://www.ebi.ac.uk/interpro/) is a protein signature database comprising
information about protein families, domains, and functional groups (Biswas et al.
2002; Mulder et al. 2003). It combines many commonly used protein signature data-
bases and is a very powerful tool for the automatic and manual annotation of new or
predicted proteins from sequencing projects. In addition, InterPro entries are
mapped to the Gene Ontology (GO, see Section 13.1) and are linked to protein en-
tries in UniProt (see Section 13.6).
13.6
Swiss-Prot,TrEMBL, and UniProt
In addition to several nucleotide sequence databases, a variety of protein sequence
databases also exist, ranging from simple sequence repositories to expertly curated
universal databases that cover many species and provide a great deal of additional in-
formation. One of the leading protein databases is Swiss-Prot (http://www.ebi.ac.uk/
swissprot/). As of August 2004 (release 44.4), it contains 158,010 protein sequence
entries. Swiss-Prot is maintained by the Swiss Institute of Bioinformatics (SIB) and
the European Bioinformatics Institute (EBI) and offers a high level of annotation
comprising information about the protein origin (gene name and species), amino
acid sequence, protein function and location, protein domains and sites, quaternary
structure, references to the literature, protein-associated disease(s), and many other
details. In addition, Swiss-Prot provides cross-references to several external data col-
lections such as nucleotide sequence databases (DDBJ/EMBL/GenBank), protein
structure databases, databases providing protein domain and family characteriza-
tions, disease-related databases, etc. (Boeckmann et al. 2003).
Since the creation of fully curated Swiss-Prot entries is a highly laborious task, an-
other database called TrEMBL (Translation from EMBL), which uses an automated
annotation approach, was introduced. TrEMBL (http://www.ebi.ac.uk/trembl/) con-
tains computer-annotated entries generated by in silico translation of all coding se-
quences (CDS) available in the nucleotide databases (DDBJ, EMBL, GenBank). The
entries offered at TrEMBL do not overlap with those found in Swiss-Prot.
The world’s most comprehensive catalog providing protein-related information is
the UniProt database (http://www.uniprot.org). UniProt is composed of information
of Swiss-Prot, TrEMBL, and PIR (http://pir.georgetown.edu). One part of UniProt,
UniParc, is the most comprehensive, publicly accessible, non-redundant protein se-
quence collection available (Apweiler et al. 2004).
408
13 Databases and Tools on the Internet

13.7
Reactome
Sequence and annotation databases such as RefSeq (Pruitt et al. 2000; Pruitt and
Maglott 2001), GeneCards (Safran et al. 2002), or YPD (Costanzo et al. 2000), provide
a gene-by-gene view of the genome. These databases present all the information
known about the structure and function of a gene and its protein(s) in a single re-
cord. However, important for biology – especially systems biology – is information
about the complex interactions among proteins, nucleic acids, and other molecules,
e.g., metabolites that carry out complex biological processes. Such data about biolo-
gical processes and pathways are not covered by the above-mentioned gene-centric
databases but often are found in the scientific literature. Reactome (formerly known
as Genome Knowledgebase, Joshi-Tope et al. 2003), is an open, online database of
fundamental human biological processes that tries to narrow this gap. The Reac-
tome project is managed as a collaboration of the Cold Spring Harbor Laboratory,
the European Bioinformatics Institute (EBI), and the Gene Ontology Consortium.
A screenshot of the Reactome homepage (http://www.reactome.org) is shown in
Fig. 13.2. The database is divided into several modules of fundamental biological
processes that are thought to operate in humans. Each module of the database has
one or more primary authors and is further peer reviewed by experts in the specific
fields. Each module can also be referenced by its revision date and thus can be cited
like a publication.
On one hand the Reactome database is intended to offer valuable information for
wet-lab scientists who want to know, e.g., more about a specific gene product they
are unfamiliar with. On the other hand the Reactome database can be used by com-
putational biologists to draw conclusions from large datasets such as expression data
gained by cDNA chip experiments. For the latter purpose, the database can be down-
loaded and locally installed as a MySQL database (DuBois 2002). Due to its clear,
hierarchical, object-oriented design, it can be queried very flexibly by Structured
Query Language (SQL) statements. (SQL is a popular computer language used to
create, modify, and query databases.)
The data model of Reactome consists of three fundamental data types: physical
entity, reaction, and pathway. A physical entity is any kind of molecule or complex
that participates in a biological process. A reaction consumes one or more physical
entities as input and produces one or more physical entities as output. Reactions can
be further grouped into pathways. With the help of these data types, the topology of
a reaction network can be described. Kinetic data (e. g., rate constants or concentra-
tion gradients) that are required for true physiological modeling are not accounted
for by the database. Gene products mentioned in the database are linked to the gene-
specific databases SwissProt, Ensembl, and RefSeq.
Another tool offered by Reactome is the “Pathfinder.” This utility enables the user
to find the shortest path between two physical entities, e.g., the shortest path be-
tween the metabolites d-fructose and pyruvate, or the steps from the primary mRNA
to its processed form. The computed path can be shown graphically. The pathfinder
also offers the possibility to exclude specific entities, such as the metabolites ATP or
409
13.7 Reactome
