Westermeier R., Naven T., H?pker H.-R. Proteomics in Practice: A Guide to Successful Experimental Design
Подождите немного. Документ загружается.


3 Mass Spectrometry270
Fig. 3.33: The workflow for DeCyder MS. DeCyder MS uses
data derived from an electrospray MS experiment.
Fig. 3.34: DeCyder MS PepMatch module (matching module)
showing the different expression of one particular peptide in
three repeats (courtesy of A. Parbel GE Healthcare).
3.6 MS Strategies 271
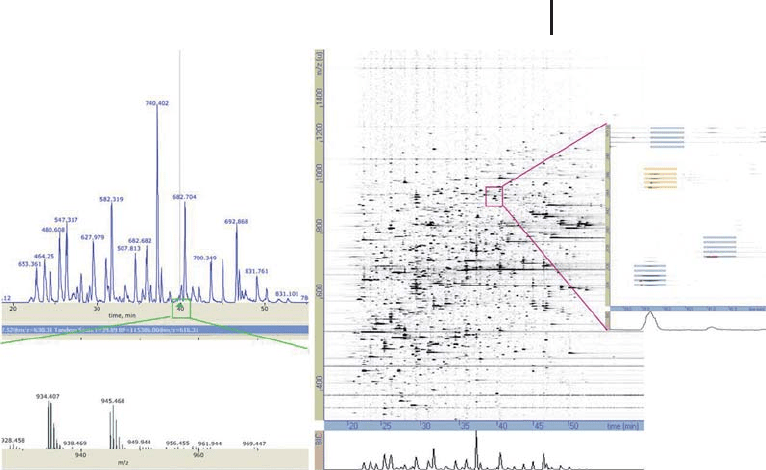
Fig. 3.35: Comparison of traditional LC-MS profile to that of
DeCyder MS, whereby peptide components can be clearly detected
with DeCyder MS. This visualization of an LC-MS profile in this
fashion is useful for a range of troubleshooting and monitoring needs.
(courtesy of A.Parbel GE Healthcare).
3.6
MS Strategies
Effectively there are two strategies, bottom up and top down. The dif-
ference, essentially, is what is the state of the sample when it enters
the mass spectrometer i.e. is it a peptide or a protein?
3.6.1
Bottom up Approach
In this approach, separated proteins or complex protein mixtures are
digested and the resultant peptides analyzed by MS in order to iden-
tify the native protein. The MS analysis is carried out either by pep-
tide mass fingerprinting or by sequence analysis using tandem mass
spectrometry. This is probably historical: 2D PAGE separated pro-
teins can only be extracted from the gel by digestion; high resolution,
nanoscale RPC of peptides supports high sensitivity MS analysis and
the performance and specification of mass spectrometers with
respect to peptides was higher than that for proteins. Fragmentation
of peptides, rather than whole proteins, has been more practical with

3 Mass Spectrometry272
mass spectrometers and mass accuracy greater for peptide analysis.
However, it does have its limitations. Acquiring complete or extensive
sequence coverage can be an issue, which can make protein charac-
terization problematic. Furthermore, the digestion process is largely
limited to trypsin, due to the fact that it locates basic residues to the
C-terminus of the peptide which is beneficial for fragmentation and
sensitivity, using CID tandem mass spectroscopy.
3.6.2
Top down Approach
This approach utilizes whole proteins and not digested peptides. This
approach is performed with mass spectrometers equipped with ETD
and ECD fragmentation. These techniques enable fragmentation of
large polypeptides and proteins.
Using this approach, accurate measurement of the molecular
weight and extensive sequence information can be acquired in one
analysis. Additionally, PTM information can be generated.

273
4
Functional Proteomics: Studies of Protein–Protein
Interactions
It is well known that the biological function cannot be linked to a sin-
gle protein. In fact proteins act in complexes and interact with other
proteins. Therefore the next step after expression proteomics is the
analysis of protein interactions. A few selected approaches are
described in brief in the following section. Many of these technolo-
gies, including SPR, are described in much more detail in the book
“Protein microarray technology” edited by Kambhamti (2003).
4.1
Non-immunological Methods
4.1.1
Separation of Intact Multi-protein Complexes
Because it provides a higher resolution than sucrose gradient centri-
fugation, the electrophoretic separation of intact protein complexes
with attached Coomassie brilliant blue dye in native polyacrylamide
gels has become well established in many proteomics laboratories.
Blue native PAGE had been developed by Schgger and von Jagow
(1991) for the separation of mitochondrial membrane proteins and
complexes in the mass range of 1,000 kDa to 10,000 kDa. Details on
the technique can be found in Section 1.2.3. In an alternative and
complementary approach multi-protein complexes are analysed by
electrospray ionization mass spectrometry (Lamond and Mann 1997).
4.1.2
Probing with Interaction Partners
“Far Western blotting” according to Burgess et al. (2000) is a non-anti-
body-based detection method for specific complex partners with a
purified bait protein after a native or a denaturating electrophoretic
separation.
Proteomics in Practice. A Guide to Successful Experimental Design 2
nd
Ed.
Reiner Westermeier, Tom Naven, and Hans-Rudolf Hçpker
Copyright 2008 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim
ISBN: 978-3-527-31941-1
Kambhampti, D (ed) Protein
Microarray Technology. Wiley-
VCH, Weinheim (2003)
Schgger H, von Jagow G. Anal
Biochem. 199 (1991) 223–
231.
Lamond AI, Mann M. Trends
Cell Biol 7 (1997) 139–142.
Burgess R, Arthur TM, Pietz
BC. Methods Enzymol. 328
(2000) 141–157.
4 Functional Proteomics: Studies of Protein–Protein Interactions274
4.1.3
Surface Plasmon Resonance
Surface plasmon resonance (SPR) measures biomolecular binding
events in real time without the use of labels by detecting mass con-
centration changes on a sensor surface. SPR has become an estab-
lished technique for measuring biomolecular interaction, in particu-
lar protein interactions. SPR occurs when surface plasmon waves are
excited at a metal/liquid interface (the sensor surface). Light is directed
at, and reflected from, the side of the surface not in contact with sam-
ple. SPR causes a reduction in the reflected light intensity at a speci-
fic combination of angle and wavelength (generating a refractive index-
dependent SPR signal). Molecules binding to the sensor surface cause
changes in the refractive index close to the surface which are detected
as changes in the SPR signal. Specifically, in an SPR experiment a
reactant (ligand) is immobilized on a surface and its interaction with
a second component (the analyte) is measured. Effectively the asso-
ciation and dissociation of the ligand–analyte interaction is measured
(Fgerstam et al. 1992; see Figure 4.1). An SPR detector is capable of
measuring real time label-free biomolecular interactions.
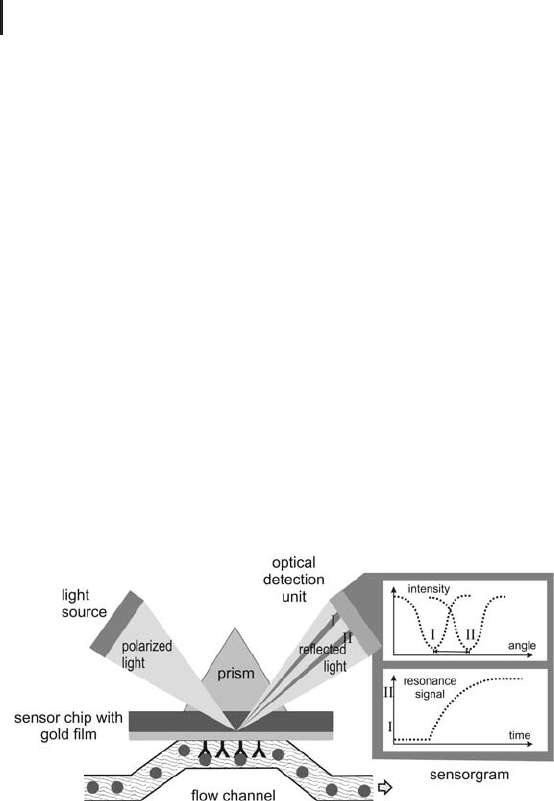
Fig. 4.1: SPR principal of detection. SPR is used to measure
the change in refractive index as the analyte binds to the ligand
(from Biacore seminar).
A number of applications have been developed for the SPR analysis
of proteins with the objective of further characterizing protein inter-
actions. In particular, real time binding data is often essential to
understand the dynamic interactions between proteins and other bio-
molecules that drive and regulate biological processes. Specific ques-
tions may include what does the protein of interest bind with or to,
where does it bind, when does it bind and how does it bind? Interest-
ingly, studies with SPR can elucidate binding strength (affinity) speci-
Fgerstam LG, Frostell-Karlsson
, Karlsson R, Persson, B,
Rçnnberg, I. J Anal Chromato-
graphy 597 (1992) 397–410.
4.2 Antibody-based Techniques 275
ficity of interaction and the rates at which interactants bind and dis-
sociate or kinetics (on and off rates).
Using SPR, complex association and dissociation are detected in
real time and data are presented graphically in an interaction profile
known as a sensorgram, which is rich in information on dynamics of
molecular interactions (see Figure 4.2). A typical experiment takes
about 30 minutes to immobilize or capture a ligand protein, 3 min-
utes for sample injection and kinetic binding measurement, and 1
minute for regeneration. The latter two steps are repeated for a series
of analyses.
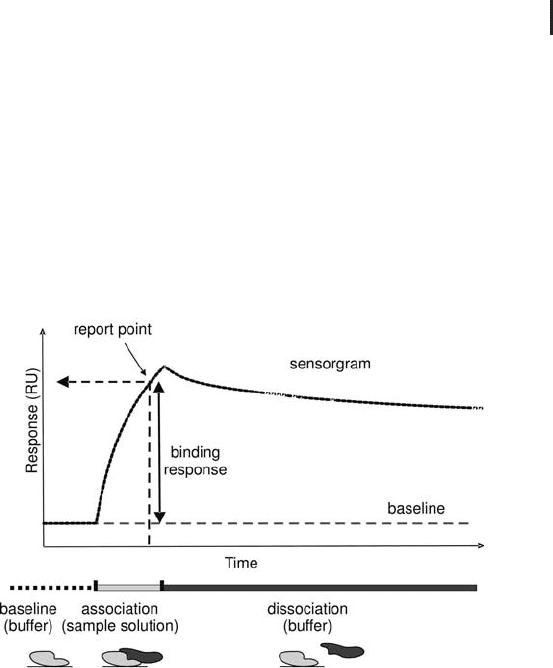
Fig. 4.2: Typical sensorgram of a label-free protein interaction analysis
with SPR. The signal height (measured in RU) is also prortional to the
mass of protein binding to the ligand.
Application areas in proteomics include the studies of protein com-
plexes, cell adhesion, signal transduction pathways, transcription,
regulation, and antibody recognition.
4.2
Antibody-based Techniques
4.2.1
Western Blotting and Dot Blots
The procedure of “Western blotting” is intensely described in Section
1.3. In many cases the efforts of electrophoretic separation of the
samples and transfer of the separated proteins onto the blotting
The major benefit of SPR tech-
nology is the information
gained from the kinetics of the
interaction.
However without pre-fractiona-
tion of the proteins cross-reac-
tions during the detection
procedure cannot completely
be excluded.

4 Functional Proteomics: Studies of Protein–Protein Interactions276
membrane is not necessary: The sample mixture possibly containing
the protein to be detected can also be applied directly on a blotting
membrane as a dot using dot blot or slot blot manifolds. The probing
for proteins is performed as described in Section 1.3.2. These dot
blots allow parallel screening of many samples.
4.2.2
Protein Microarrays
The concept described above, also called a “macroarray”, has mean-
while evolved into microarray technologies, using microscope slide
format (2575 mm). The basic technology for microarrys had been
developed for nucleic acid analysis in genomics and transcriptomics
studies. Similar methodologies are applied on expression profiling of
proteins in non-fractionated mixtures: Proteins are arrayed on mem-
branes or on microscope glass slides (protein chips) in very low
volumes in tiny spots using instruments similar to inkjet printers.
Either antibodies are printed on the carrier to capture specific ana-
lytes from body fluids, cell and tissue lysates; or – vice versa – the
sample lysates are immobilized on the carrier and antibodies are
used to detect specific sample components. Detection is usually per-
formed with a secondary antibody labeled with a fluorescent tag, like
cyanine dyes as used in the DIGE approach. In this way multiplexed
protein expression profiling can be carried out in a single assay; and
relative quantification is enabled. The spotting is fully automated by
employing pipetting robots, which apply the solutions from 384-well
plates. The images require high resolution multifluorescence scan-
ners.
Protein microarrays allow studying many biological events simulta-
neously with very small sample volumes, resulting in molecular fin-
gerprints on the protein level. Protein microarrays show still some
methodological limitation, however there is a huge potential for a
wide area of applications from high-throughput proteomics to clinical
diagnostics. Figure 4.3 shows a typical microarray image after scan-
ning with different wavelength fluorescence detection.
A critical step is the immobili-
zation of the proteins onto the
support. Random orientation of
the proteins can cause lossof
biological activity and/ or
recognition of its epitope by the
antibody.
Also their ease of use and their
possibility of automation
protein microarrays pledge to
be more suitable for diagnostic
applications than the classic
proteomic separation technolo-
gies.
4.2 Antibody-based Techniques 277
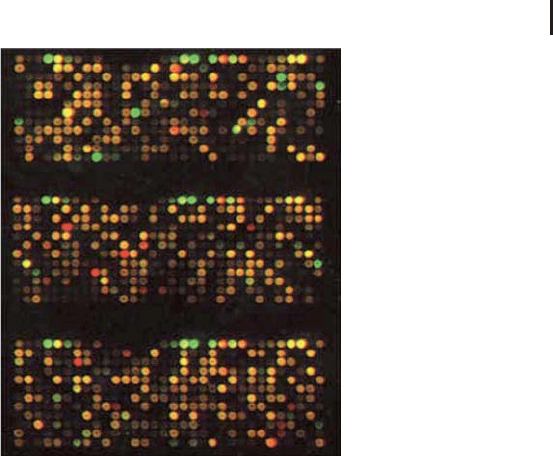
Fig. 4.3: Typical microarray image using Cy3 and Cy5
fluorescent labels after scanning with a high-resolution
version Typhoon multifluorescence scanner.

279
Part II:
Practical Manual of Proteome Analysis
Equipment, Consumables, Reagents 281
Step 1: Sample Preparation 287
Step 2: Fluorescence Difference Gel Electrophoresis 299
Step 3: Isoelectric Focusing 309
Step 4: SDS Polyacrylamide Gel Electrophoresis 323
Step 5: Scanning of Gels Containing Pre-labeled Proteins 357
Step 6: Staining of Gels 361
Step 7: Image Analysis and Evaluation of DIGE Gels 373
Step 8: Spot Excision 383
Step 9: Sample Destaining 387
Step 10: Protein Digestion 389
Step 11: Microscale Desalting and Concentrating of Sample 393
Step 12: Chemical Derivatization of the Peptide Digest 397
Step 13: MS Analysis 399
Step 14: Calibration of MALDI-ToF MS 403
Step 15: Preparing for a Database Search 407
Proteomics in Practice. A Guide to Successful Experimental Design 2
nd
Ed.
Reiner Westermeier, Tom Naven, and Hans-Rudolf Hçpker
Copyright 2008 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim
ISBN: 978-3-527-31941-1
