Klipp E., Herwig R., Kowald A., Wierling C., Lehrach H. Systems Biology in Practice: Concepts, Implementation and Application
Подождите немного. Документ загружается.

tages of federated databases is that they do not require that component databases be
physically combined into one database. A database federation system does not mod-
ify the primary data resources. The databases are connected through a specialized
network service shared by applications (databases and tools) and users to create a vir-
tual integrated database (Fig. 11.1).
In the data warehouse approach, the component databases are mapped to a single
database called the data warehouse. Creating a data warehouse is a process that con-
sists of two steps (Karp 1995). First the schemas of the component databases are
mapped onto the global schema (i. e., the schema of the data warehouse). Second,
data is extracted from the component databases and used to populate the data ware-
house. A data warehouse stores the core data (a set of relational databases) in a cen-
tral store. The warehouse is connected to other relational databases from which a
subset of the data can be selectively extracted and loaded for analysis. Slightly differ-
ent are data marts, which store specialized data derived from a data warehouse. The
emphasis here is on content, presentation, and ease of use in formats familiar to the
specialized user.
11.1.2
SRS
One of the first database integration platforms developed was SRS (Sequence Retrie-
val System). SRS was originally developed as a tool for providing easy access to biolo-
gical sequence databases (Etzold and Argos 1993; Etzold et al. 1996) and has been ex-
tended to other data sources as well. SRS was acquired by LION Bioscience AG in
1998 and further developed as a commercial product. SRS version 6 is freely avail-
able for academic research. Currently, there are more than 100 servers worldwide
(http://downloads.lionbio.co.uk/publicsrs.html). The EBI SRS server contains wrap-
pers for more than 400 databases and 20 applications connected to the system. Appli-
cations include, for example, sequence-matching tools such as BLASTand FASTA.
The federated databases are stored locally with a query integration engine acting
as middleware. SRS is based on the indexing of flat files. Text files are still the de
facto standard for biological databases such as EMBL, SWISS-PROT, etc. XML files
370
11 Data Integration
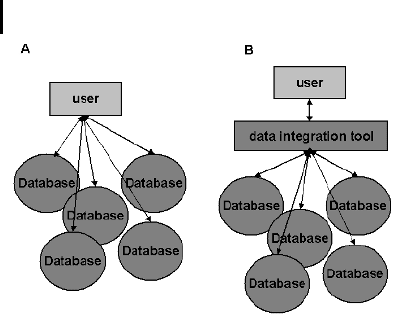
Fig. 11.1 Basic schema illustrating
the concept of database integration.
The common situation is that the user
works with different heterogeneous and
autonomous databases (a). This re-
quires the interaction with each data-
base, and each query has to be inserted
into each database separately. (b) In a
database integration setup the user in-
teracts with the integration tool.
Queries will then be translated by the
integration tool and sent to the indivi-
dual database and a consistent output
will be provided.

have a more structured format but are still considered as text files. In this system,
each database record in any component database is processed separately as a collec-
tion of data fields. An index is created for each data field (data field index). This index
classifies each record in the database according to a set of keywords from a con-
trolled vocabulary. Each database will have one index per data field. A different type
of index (link index) is used to link individual databases. A link index is created for
each pair of databases that includes cross-references to each other and these links
are bidirectional. Databases that do not directly reference each other can still be con-
nected by traversing links through intermediate databases. SRS has a unique ap-
proach to addressing syntactic and semantic heterogeneities.
SRS has an object-oriented design. It uses metadata to define a class for a database
entry object and rules for text-parsing methods, coupled with the entry attributes.
SRS incorporates a proprietary parsing language (Icarus) for generating the database
wrappers and another language (SRS query language) for the formulation of queries
(Zdobnov et al. 2002). Data in SRS can be subdivided into sections that correspond
to the main contents of the integrated databases such as DNA sequences, protein se-
quences, mapping data, SNPs, and metabolic processes.
SRS is a keyword-based system. Queries can be combined using logical operators
such as “&” (AND), “|” (OR) and “!” (BUT NOT). An HTML interface is available for
the formulation of queries and for viewing the results of the data retrieval. Thus,
SRS can be used as a front end to independently query multiple data sources.
There are some APIs of SRS version 6 to most widely used programming lan-
guages such as C++ and Java. This allows the development of customized interfaces
to proprietary analysis tools.
11.1.3
EnsMart
EnsMart (http://www.ensembl.org/EnsMart) provides a generic data warehouse sys-
tem (Kasprzyk et al. 2004). The system organizes data from individual databases into
one query-optimized system by the incorporation of the data-warehousing technique
for descriptive data. Currently, it is focused on Ensembl and thus primarily entails
data with genomic annotation such as genes and SNPs, functional annotation, and
expression data. Data are available for nine different species annotated in Ensembl
(Homo sapiens, Mus musculus, Rattus norvegicus, Danio rerio, Fugu rubripes, Anopheles
gambiae, Drosophila melanogaster, Caenorhabditis briggsae, and Caenorhabditis elegans)
(Birney et al. 2004).
EnsMart data are organized around central objects, so-called foci, and additional
satellite data. Currently, two foci exist, gene and SNP, and all additional data are pre-
sented in relation to these foci. EnsMart comes up with three different user inter-
faces. The MartView is an Internet user interface in the “wizard” style. It allows navi-
gating through pages to specify user input. Furthermore, it is used to specify the out-
put format and to handle data export. The MartExplorer is a local database. It is in-
stalled as a program and has a graphical user interface that allows displaying the
371
11.1 Database Networks

query as a tree in an interactive manner. The MartShell is a command line shell that
uses a query language specifically designed for mart queries.
EnsMart has powerful querying facilities. The querying process is organized into
three steps: start, filter, and output. The start stage specifies the organism and the fo-
cus for the query. The filter step offers the possibility of narrowing the search accord-
ing to several criteria. For example, chromosomes and specific regions on the chro-
mosome can be specified as an attribute using the region filter. Other filters restrict
the search to specific gene classes (“novel genes,” “disease genes,” etc.) or genes that
are mapped to a particular identifier (e.g., HUGO gene symbols). The output stage
defines the attributes that are used for exporting the data. This is dependent on the
focus that has been selected previously. Queries can be chained, i.e., the file output
from previous queries can be used as filters for the current query.
11.1.4
DiscoveryLink
DiscoveryLink was developed by IBM and is another example of a federated database
approach. It has been the result of the fusion of two products: IBM’s Garlic federated
database prototype for multimedia information and DataJoiner, a federated database
management product based on the DATABASE 2 (DB2) relational database product.
It supports SQL (standard query language) for queries and comes up with wrappers
in C++. DiscoveryLink requires a DB2 instance. Wrappers are available for most rela-
tional database management engines, e. g., ORACLE. The system consists of a logi-
cal data model for the federated data resources in the DB2 instance and a series of
“nicknames.” A nickname in IBM’s terminology is a reference to a relational table
that belongs to one of the federated data resources. For example, the nickname “pro-
tein” associated with a set of attributes would correspond to a “protein” table in a re-
lational database specifically designed to store protein-associated information.
Queries can be sent containing any combination of nicknames in the DB2 instance
as if they were relational tables. The set of nicknames defined for the federated data
sources forms a global schema. This schema determines the types of searches that
can be formulated, similar to a relational data model. DiscoveryLink is essentially a
middleware engine and applications and user interfaces must be developed exter-
nally.
11.1.5
Data Exchange
eXtensible Markup Language (XML) is a widespread standard for formatting text
files in a hierarchical order. XML consists of key words (or tags) and attributes asso-
ciated with these key words (cf. Section 14.2). The set of key words can be defined
flexibly, which distinguishes XML from, for example, HTML. This feature has given
rise to multiple XML conform data-exchange languages for specific purposes so that
XML is now the working standard for storing and exchanging information with an
inherent hierarchical structure (Achard et al. 2001). Most databases are available in
372
11 Data Integration

XML-compatible format and come up with tools to parse the information. Table 11.1
gives an overview of several relevant XML conform markup languages.
The recent advent of high-throughput technologies in genomics and proteomics
has given rise to large consortia that attempt to define common schemas for data sto-
rage and data exchange mostly based on XML. One example is the MIAME consor-
tium, which attempts to unify descriptions of microarray experiments (Brazma et al.
2001). MIAME requires detailed annotation about experimental conditions, materi-
als, and procedures to be captured. The MIAME-required information can be en-
coded in MAGE-ML.
MAGE-ML is a formal XML language designed to describe and communicate in-
formation about microarray-based experiments such as microarray designs, microar-
ray manufacturing information, microarray experiment setup and execution infor-
mation, gene expression data, and data analysis results.
A similar approach has been attempted in proteomics for sophisticated algorithms
and methods on data standardization, data integration, and data exchange in order
to gain improvements in data reproducibility and data quality control. This attempt
has been targeted in recent publications (Taylor et al. 2003; Hermjakob et al. 2004).
Proteomics technologies (MS, 2D gels, protein-protein interactions etc.) generate
large and diverse datasets that require standardized schemas for interchanging data.
For example, different MS manufacturers store data in different proprietary formats,
which limits to a large extent data analysis, exchange of raw datasets, and software
development. Furthermore, many strategies exist for assigning peptides to mass
spectra that end up in partly conflicting results. Protein-protein interactions and the
construction of networks suffer from high false-positive and false-negative rates (in-
correct folding, inadequate subcellular localization, etc.), which tremendously down-
grade the reproducibility and comparability of these approaches. The exchange, sto-
rage, and standardization of proteomics data based on XML-like schemas are a cen-
tral goal of the above-mentioned projects and consortia (PEDRo, HUPO).
373
11.1 Database Networks
Tab. 11.1 Some available XML conform markup languages.
Language Description Use
BSML Bioinformatic Sequence Markup Language Genomic sequences and
biological function
MATHML Mathematical Markup Language Mathematical formulas
BioML Biopolymer Markup Language Complex annotation for
protein and DNA sequences
MAGEML Microarray and Gene Expression Markup Language Microarray data exchange
SBML Systems Biology Markup Language Biochemical networks and
models
CML Chemical Markup Language Managing chemical
information
CellML Cell Markup Language Mathematical models

11.2
Information Measurement in Heterogeneous Data
Chapter 9 introduced the main analysis tools for DNA array data, basically targeting
measurements of the transcriptome. System-wide approaches will measure data on sev-
eral levels of cellular information, e.g., adding textual data in the form of gene annota-
tion, gene regulation, interaction data, etc. Aparticularly interesting question is whether
heterogeneous data can be integrated by analytical concepts in order to give a coherent
view of the objects under analysis across these data. Essentially, this is a problem of sui-
table similarity functions that allow the measurement of correlation in the different
data sources. Commonly used similarity measures are mathematical functions based
on the feature matrices of two datasets that are based on geometric considerations such
as distances and correlations. Such measures are less appropriate since datasets are het-
erogeneous and it is not straightforward to add a topological meaning to some data
sources, e.g., to categorical data or data derived from binary classifications. In contrast,
measures based on information concepts can be applied here. In this section we will in-
troduce such concepts, namely, entropy and mutual information. Entropy and mutual
information have been applied in the analysis of DNA sequences and gene expression
(Herwig et al. 1999). Furthermore, in Section 9.6 we showed the application of mutual
information to derive interaction rules in the reverse engineering of Boolean networks
(Liang et al. 1999). We will highlight a recent example from the literature where this
concept was used to combine gene expression data with functional annotations.
11.2.1
Information and Entropy
The concept of information was originally introduced by Shannon (1948) in the con-
text of communication theory. Given a source of information that sends discrete
symbols from an alphabet A ={a
1
,…,a
K
} with probabilities p
1
,…,p
K
, i. e.,
P
K
i1
p
i
=1,
the information content of a single letter of the alphabet, a
i
, is defined as I(a
i
)=
–log
2
(p
i
). This definition is motivated through the following considerations:
. Non-negativity: The information content of a single letter is nonnegative since we
have p
i
B [0,1] and thus –log
2
(p
i
)6 0.
. Monotony: The idea here is that rare letters should have a high information con-
tent since they are unexpected because of their low probability. Conversely, fre-
quent symbols should have a low information content since they correspond to
highly probable events. Thus, the information content of single letters should in-
crease reciprocally to their probability.
. Additivity: The amount of information of K independent letters a
1
,…,a
K
with the
same probability, p, should scale linearly with the amount of information of single
symbols. This is implied by the functional characteristics of the log function
Ia
1
; :::; a
K
log
2
pa
1
; :::; a
K
K log
2
p
KI a
1
: (11-1)
374
11 Data Integration

This concept extends straightforwardly if more than one information source is
used independently. Consider A and B to be two alphabets, A ={a
1
,…,a
K
} and
B ={b
1
,…,b
L
}. The composed alphabet, A6 B, consists of KL letter pairs (a
k
, b
l
). In-
tuitively, since the sources are independent of each other, the information content of
a letter pair should be the sum of the information content of the individual letters.
This is assured by the functional characteristic of the log function:
Ia
k
; b
l
log
2
p
kl
log
2
p
k
p
l
log
2
p
k
log
2
p
l
Ia
k
Ib
l
:
The base of the logarithm is equal to two, which is motivated by information the-
ory where information is stored in bits. The more bits that are stored in a memory
unit, the higher the information content is. For example, a unit of n bits can store 2
n
binary-coded digits, which have an information content of –log
2
1
2
n
= n.
To characterize the source completely, the information content of single letters is
not sufficient. Therefore, the entropy concept has been introduced. Entropy is de-
fined as the average (or expected) information content of the source with respect to
all letters, i.e.,
HA
P
K
i1
p
i
log p
i
: (11-2)
Entropy is commonly denoted by the symbol H. This points back to early concepts
of entropy in thermodynamics in the 19th century introduced by Boltzmann (defin-
ing the H function) to characterize systems in high-dimensional phase spaces in or-
der to locate these systems between order and chaos.
Alternatively to the motivation from information theory, entropy is sometimes in-
troduced in statistics in order to describe the grade of uncertainty of a probability dis-
tribution (compare Section 3.4). Let P ={p
1
,…,p
n
} be a finite probability distribu-
tion, i. e., a set of n nonnegative numbers that sum up to one,
P
n
i1
p
i
= 1. The entropy
of P is defined, similarly to Eq. (11-2), as
HP
X
n
i1
p
i
log p
i
: (11-3)
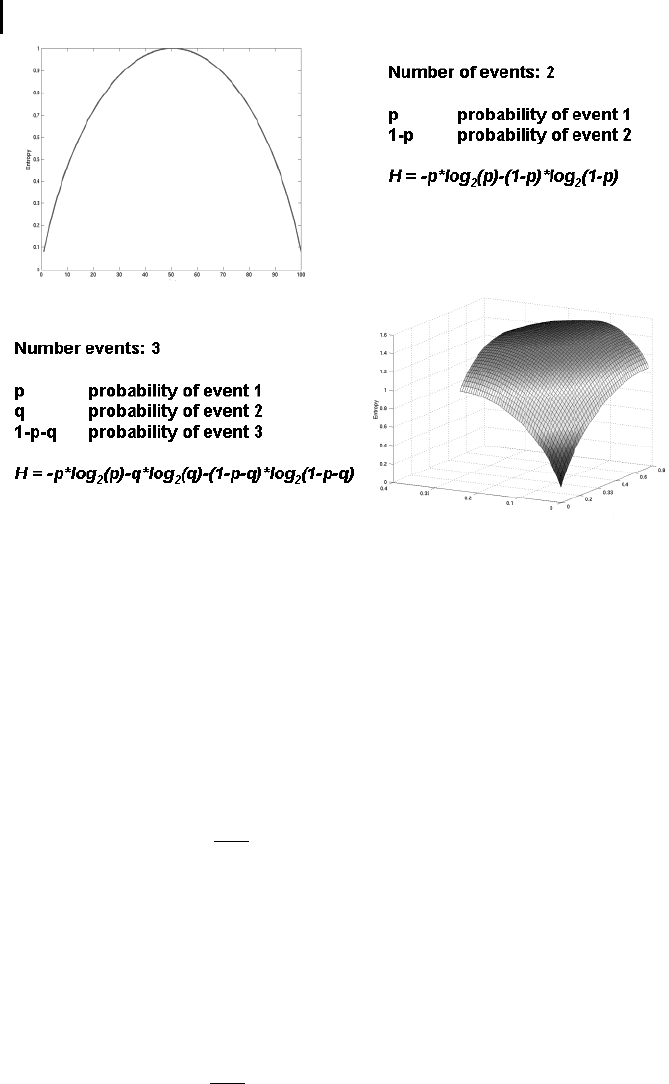
Figure 11.2 displays the entropy as a function of the individual probabilities in the
case of n = 2 and n = 3. If all probability weights are concentrated on a single event,
then entropy is zero. If the events have equal weights, then entropy is maximal. En-
tropy is a measure of the mean information inherent in the probability distribution
and can be considered as the expectation of the probability distribution under a log
transformation of the data.
375
11.2 Information Measurement in Heterogeneous Data
11.2.2
Mutual Information
Mutual information is a concept introduced by Shannon (1948) in order to measure
information coherence in two datasets. Consider an alphabet A6B composed of
two discrete alphabets, A ={a
1
,…,a
K
} and B ={b
1
,…,b
L
}, that are not necessarily in-
dependent of each other. The mutual information of A and B is defined as
HA;B
X
k;l
p
kl
log
2
p
kl
p
k
p
l
: (11-4)
Here, p
kl
is the probability of the letter pair (a
k
, b
l
) and p
k
and p
l
are the probabil-
ities of the letters a
k
and b
l
, respectively.
Similarly, mutual information can be defined with respect to two random vari-
ables, x and y, with corresponding probability distributions. If we denote by {p
kl
} the
set of probabilities of the joint probability distribution of x and y, then mutual infor-
mation is defined, similarly to Eq. (11-4), by
Hx;y
X
k;l
p
kl
log
2
p
kl
p
k
p
l
: (11-5)
376
11 Data Integration
Fig. 11.2 Entropy as a function of the individual probabilities of the
probability distribution for the case of distributions of size two (upper
part) and three (lower part).

Mutual information measures the amount of information of x inherent in y.
Using the characteristic of the log function, it can be shown that
Hx;y
HxHy
Hx; y
; (11-6)
where H(x,y) is the entropy of the joint probability distribution of x and y.
Several properties of mutual information can be derived (Cover and Thomas
1991):
0 Hx;y
min Hx; Hy
(11-7)
Hx;y
Hy;x
(11-8)
Hx;x
Hx;y
(11-9)
Hx;y
0 ; if and only if x and y are stochastically independent. (11-10)
The first inequality states that mutual information cannot be higher than the en-
tropy of individual sources. This follows intuition because we cannot gain informa-
tion from x through y that is not inherent in either x or y. Furthermore, if both vari-
ables are independent of each other, we cannot gain any information from x about y
and vice versa (Eq. (11-10)).
Practically, mutual information is used to evaluate contingency tables. Suppose
that we have two data measurements that partition the data in K and L groups, re-
spectively. The coherence of the measurements can be described by a KxL contin-
gency table of the following form:
x/y 1 2 … L Total
1 n
11
n
12
… n
1L
n
1.
2 n
21
n
22
… n
2L
n
2.
… ……………
K n
K1
n
K2
… n
KL
n
K.
Total n
.1
n
.2
… n
.L
n
Here, n
k.
and n
.l
refer to the kth and lth marginal of the row and column classes,
respectively, and n is the overall number of measured features. Mutual information
is calculated according to the frequency of the cell entries:
Hx;y
X
k;l
n
kl
n
log
2
n
kl
n
n
k:
n
: l
: (11-11)
Mutual information is an indicator of the coherence of information in the two da-
tasets. It should be noted that mutual information does not distinguish between cor-
relation (high agreement in the diagonal of the contingency table) and anti-correla-
tion (high agreement in the anti-diagonal of the table [see Example 11-1]).
377
11.2 Information Measurement in Heterogeneous Data

Example 11-1
Consider the binary data vectors x = (0000011111), y = (0000011111), and
z = (1111100000). Mutual information will result in similar values when judging
the pairwise similarities, i.e., H(x; y)=H(x; z), although the former two vectors
are identical and the latter two vectors are completely diverse. This might be a de-
sired effect. For example, if the vectors describe the state vectors of two compo-
nents of a Boolean network, y and z, in relation to a third, x, then an immediate
interaction rule would retrieve that x is an activator of y and an inhibitor of z.
However, sometimes this is an undesired effect that identifies the need for cor-
recting mutual information for anti-correlation. By applying the correction below,
we will get a negative result for the latter pair, i.e., H (x; z)=–H(x; y), which
takes into account anti-correlation.
For simplicity we will assume a binary categorization of the data. Thus the above
contingency table reduces to the following table:
x/y 1 2 Total
1 n
11
n
12
n
1.
2 n
21
n
22
n
2.
Total n
.1
n
.2
n
Intuitively, one would assign results that have more weights in the anti-diagonal than
in the diagonal a negative result, similar to the Pearson correlation coefficient, where
anti-correlation is measured with negative values close to –1. Thus, the exact threshold
should be a function of the diagonal elements and should take into account the margin-
als of the table. A reasonable indicator for anti-correlation is the occurrence of fewer
weights in the diagonal than one would get from a random assignment of class attri-
butes. Consider two n-dimensional vectors that carry information for two categories. Let
n
.2
and n
2.
be the marginals of the occurrences of the second category (we have to con-
sider only one category since the other values are then uniquely determined). We can ask
for the probability that we will observe k times an agreement of that category by chance
in the two vectors. This probability is given by the hypergeometric distribution, i.e.,
Pk
n
2:
k
n n
2:
n
:2
k
n
n
:2
: (11-12)
The expectation of this distribution in dependence on the marginals is
E
n
2:
n
:2
n
: (11-13)
378
11 Data Integration
The expectation value is a natural indicator for anti-correlation of the two meas-
urements, in the sense that any observed agreement should have significantly more
matches. Thus, in the binary case a correction of mutual information would read
Hx;y
sign n
22
n
2:
n
:2
n
X
2
k;l1
n
kl
n
log
2
n
kl
n
n
k:
n
:l
: (11-14)
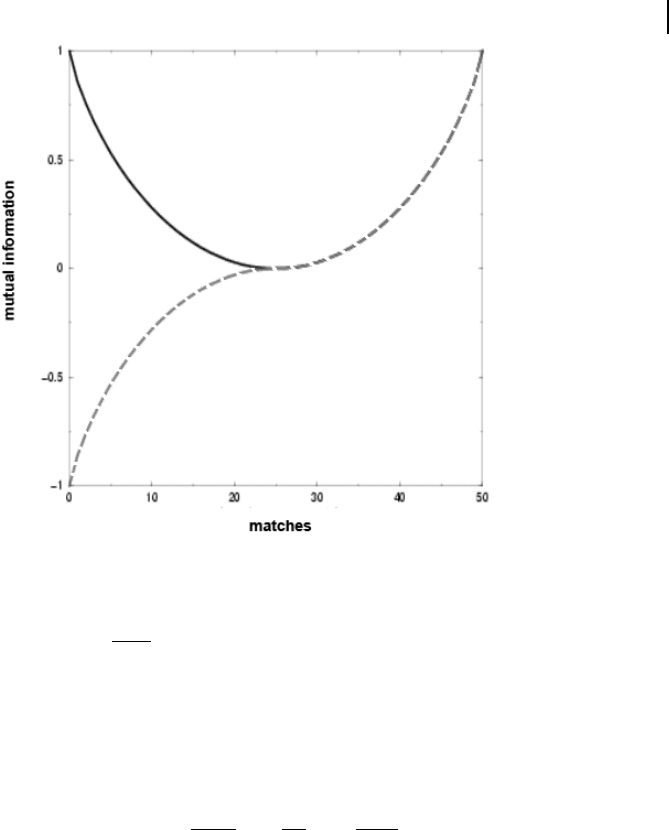
Figure 11.3 displays the effect of correction for anti-correlation of mutual informa-
tion. It should be pointed out that this correction cannot be straightforwardly ex-
tended to the case of more than two classes.
11.2.3
Information Correlation: Example
In this subsection we discuss the application of mutual information as a method for
validating the results of clustering gene expression profiles with functional annotation
(Gibbons and Roth 2002). These authors addressed the question of how clustering re-
379
11.2 Information Measurement in Heterogeneous Data
Fig. 11.3 Mutual information as a function of the diagonal element
in a 262 contingency table derived from a binary classification from
two different measurements (solid line). In this example, n = 100 and
n
2.
= n
.2
= 50. Mutual information is minimal if n
22
equals its expec-
tation, i.e., n
22
=
n
2:
n
:2
n
. This quantity can be used to adjust the
measure for anti-correlation (dashed line).
