Klipp E., Herwig R., Kowald A., Wierling C., Lehrach H. Systems Biology in Practice: Concepts, Implementation and Application
Подождите немного. Документ загружается.


is termed exocytosis when proteins produced by the cell are secreted to the exterior and
endocytosis or phagocytosis when extracellular substances are taken up by the cell.
There are two different kinds of exocytoses. The first one is a constitutive secretion:
synthesized proteins packed into transport vesicles at the Golgi complex move to the
plasma membrane and fuse with it, thereby delivering their payload to the exterior.
This happens, e.g., with proteins intended for the extracellular matrix. In the second
case termed regulated exocytosis, the proteins coming from the Golgi complex via
transport vesicles are enriched in secretory vesicles that deliver their content usually
due to an external signal recognized by a receptor and further transmitted via second
messengers (e.g., Ca
2+
). This pathway is common, for example, to neurotransmitters
secreted by neurons or digestive enzymes produced by acinus cells of the pancreas.
Vesicular transport is important for large molecules such as proteins. For smaller
molecules (e. g., ions or glucose) there are alternative mechanisms. In the case of
passive transport, the flux takes place along an osmotic or electrochemical concentra-
tion gradient and requires no expenditure of cellular energy. Therefore, either the
molecules can diffuse through the membrane or, since especially polar and charged
substances cannot pass this hydrophobic barrier, transport is mediated selectively by
integral transmembrane proteins. Other transmembrane proteins enable an active
transport against a concentration gradient that requires cellular energy (e.g., ATP).
Sensing of exterior conditions and communication with other cells are often
mediated by receptors of the cell membrane that tackle the signal transmission. Al-
ternatively, mostly hydrophobic substances like steroid and thyroid hormones can
cross the cell membrane directly and interact with receptors in the cell’s interior. Sig-
nal transduction is discussed in more detail in Chapter 6. A general overview of bio-
chemistry of signal transduction is given, e.g., in Krauss (2003).
Besides the plasma membrane, plant cells are further surrounded by a cell wall
with cellulose, a polysaccharide, as the main polymer forming the fundamental scaf-
fold. Prokaryotes also often have a cell wall where different monosaccharides act as
building blocks for the polymer.
2.3.2
Nucleus
Prokaryotes store their hereditary information – their genome – in a single circular,
double-stranded DNA (located in a subregion of the cell’s interior called the nu-
cleoid) and optionally in one or several small, circular DNAs (the plasmids), which
code for further genes. The genome of eukaryotes is located in the cell nucleus and
forms the chromatin that is embedded into the nuclear matrix and has dense regions
(heterochromatin) and less dense regions (euchromatin). The nucleus occupies
about 10% of the cellular volume and is surrounded by the nuclear envelope formed
by an extension of the ER that creates a double membrane. The nuclear envelope
has several protein complexes that form nuclear pores and that are responsible for
the traffic between the nucleus and the cytosol. A subregion of the chromatin in
which many repeats of genes encoding ribosomal RNAs are located appears as a
roughly spherical body called the nucleolus.
40
2 Biology in a Nutshell
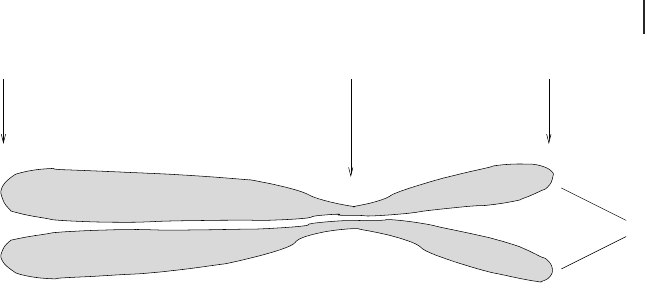
The structure of the chromatin usually becomes optically clearer during cell divi-
sion, when the DNA strands condense into chromosomes, each consisting of two
DNA double strands called chromatids. Both chromatids are joined at the centro-
mere (Fig. 2.10). The ends of the chromatids are called telomeres. At the molecular
level, the DNA of a chromosome is highly ordered. The double strand is wound
around protein complexes, the histones, and each DNA/histone complex is called a
nucleosome.
2.3.3
Cytosol
The cytosol fills the space between the organelles of the cytoplasm. It represents
about half of the cell volume and contains the cytoskeletal framework. This fibrous
network consists of different protein filaments that constitute a general framework
and are responsible for the coordination of cytoplasmatic movements. These activ-
ities are controlled by three major types of protein filaments: the actin filaments
(also called microfilaments), the microtubules, and the intermediate filaments.
The long, stretched actin filaments, with a diameter of about 5–7 nm, are built
up of globular actin proteins. One major task of actin filaments is the generation
of motility during muscle contraction. For the generation of movement, actin fila-
ments slide along another filament type called myosin. This ATP-consuming pro-
cess is driven by a coordinated interaction of these proteins. Together with other
proteins involved in the regulation of muscle activity, these filaments form very reg-
ular structures in muscle cells. Furthermore, in many animal cells, actin filaments
associated with other proteins are often located directly under the plasma mem-
brane in the cell cortex and form a network that enables the cell to change its shape
and to move.
Another filament type found in eukaryotes are the microtubules. They consist of
heterodimers of the proteins a- and b-tubulin, which form unbranched cylinders of
41
2.3 Structural Cell Biology
TelomereTelomere Centromere
Chromatid
Fig. 2.10 The chromosome consists of two identical chromatids
that are connected at the centromere. Both ends of the linear DNA
of a chromatid are called telomeres. Each chromatid has a short and
a long arm. The DNA is wrapped around histones and coiled further.

about 25 nm in diameter with a central open channel. These filaments are involved,
e.g., in rapid motions of flagella and cilia, which are hair-like cell appendages. Fla-
gella are responsible for the movement of, e.g., sperm and many single-celled eukar-
yotic protists. Cilia occur, for instance, on epithelial cells of the human respiratory
system. The motion of cilia or flagella is due to the bending of a complex internal
structure called the axoneme. Almost all kinds of cilia and eukaryotic flagella have
nearly the same characteristic structure of the axoneme. This is called the 9 + 2
structure, because of its appearance: nine doublets that look like two condensed mi-
crotubules form a cylinder together with other associated proteins, the center of
which contains two further single microtubules. The flexibility of the axoneme is
also an ATP-consuming process that is further assisted by the protein dynein.
The third major filament type of the cytoskeleton is the intermediate filament. In
contrast to actin filaments and microtubules, which are built of globular proteins, in-
termediate filaments consist of fibrous proteins. Several subtypes of these filaments
are known, e. g., keratin filaments in the cytosol of epithelial cells, which make these
cells resistant to mechanical influence, or lamin filaments, which are involved in the
formation of the nuclear lamina.
Furthermore, the cytosol contains ribosomes responsible for protein synthesis
(see Section 2.4.3), and it is filled with thousands of metabolic enzymes. A central
metabolic pathway that is catalyzed by some of these enzymes is glycolysis. Sub-
strates of this pathway are glucose or some similar six-carbon derivatives of it. These
substrates are converted by several reactions into two molecules of the three-carbon
compound pyruvate. Each metabolized glucose molecule generates two molecules of
ATP, and one NAD
+
(the oxidized form of nicotinamide adenine dinucleotide) is re-
duced to NADH. But via this pathway – which does not involve molecular oxygen –
only a small amount of the energy that can be gained through oxidation of glucose is
made available. In aerobic organisms the bulk of ATP is produced from pyruvate in
the mitochondria (see Section 2.3.4).
2.3.4
Mitochondria
Mitochondria have a spherical or elongated shape and are about the size of a bacter-
ium. Their interior is surrounded by two membranes: a highly permeable outer
membrane and a selective inner membrane. Therefore, mitochondria have two inter-
nal compartments, the intermembrane space and the matrix. The outer membrane
is permeable for ions and most of the small molecules due to several transmem-
brane channel proteins called porins. The inner membrane’s surface area is strongly
increased by numerous folds and tabular projections into the mitochondrial interior,
which are called cristae. Mitochondria are partially autonomous: they possess their
own DNA and enzymatic complexes required for protein expression (such as ribo-
somes and mRNA polymerase). Nevertheless, they depend on symbiosis with their
cell since most genes of mitochondrial proteins are encoded by the nuclear DNA.
These mitochondrial proteins are synthesized in the cytoplasm and are then im-
ported into the organelle.
42
2 Biology in a Nutshell

As mentioned above, the bulk of ATP (34 out of 36 molecules per metabolized glu-
cose molecule) is gained in mitochondria; thus, they can be termed the “power
plants” of eukaryotic cells. The underlying oxidative process that involves molecular
oxygen and yields CO
2
and ATP is driven mainly by pyruvate from the glycolysis and
fatty acids. Both pyruvate and fatty acids can be converted into acetyl-CoA molecules.
Acetyl-CoA has an acetyl group (CH
3
CO–, a two-carbon group consisting of a methyl
group and a carbonyl group) that is covalently liked to coenzyme A (CoA). Cytosolic
pyruvate can pass the outer mitochondrial membrane and enter the mitochondrial
matrix via a transporter of the inner membrane. Pyruvate is then converted into
acetyl-CoA by a huge enzyme complex called pyruvate dehydrogenase. Acetyl-CoA
reacts with oxaloacetate and thus enters the citrate cycle, a sequence of several reac-
tions during which two CO
2
molecules and energetic reduction equivalents (mainly
NADH, but also FADH
2
) are produced. Finally, oxaloacetate is regenerated and thus
the cycle is closed. The electrons delivered by the reduction equivalents are further
transferred step by step onto O
2
, which then reacts together with H
+
ions to form
water. The huge amount of energy provided by this controlled oxyhydrogen reaction
is used subsequently for the transfer of H
+
ions out of the mitochondrial matrix,
thus establishing an H
+
gradient across the inner membrane. The energy provided
by this very steep gradient is used by another protein complex of the inner mitochon-
drial membrane – the ATP synthetase – for the production of ATP inside the mito-
chondrial matrix by a flux of H
+
from the intermembrane space back into the matrix.
This coupled process of an oxidation and a phosphorylation is called oxidative phos-
phorylation. The complete aerobic oxidation of glucose produces as many as 36 mo-
lecules of ATP:
C
6
H
12
O
6
+6O
2
+36P
i
–
+ 36ADP
3–
+36H
+
? 6CO
2
+36ATP
4–
+42H
2
O
2.3.5
Endoplasmatic Reticulum and Golgi Complex
The endoplasmatic reticulum (ER) is a widely spread cytosolic membrane system
that forms tubular structures and flattened sacs. Its continuous and unbroken mem-
brane encloses a lumen that stays in direct contact with the perinuclear space of the
nuclear envelope. The ER occurs in two forms: the rough ER and the smooth ER.
The rough ER forms mainly flattened sacs and has many ribosomes that are at-
tached to its cytosolic surface; the smooth ER lacks ribosomes and forms mostly tub-
ular structures. Proteins destined for secretion but also intended for the ER itself,
the Golgi complex, the lysosomes, or the outer plasma membrane enter the lumen
of the ER directly after being synthesized by ribosomes of the rough ER. The total
number of ER membranes of a cell and the ratio of smooth and rough ER vary
strongly depending on species and cell type. All enzymes required for biosynthesis
of membrane lipids, such as phosphatidylcholine, phosphatidylethanolamine, or
phosphatidylinositol, are located in the ER membrane, their active centers facing the
cytosol. Membrane lipids synthesized by these enzymes are integrated into the cyto-
solic part of the ER bilayer. Since this would result in an imbalance of lipids in the
43
2.3 Structural Cell Biology

two layers of the membrane, phospholipid translocators can increase the flip-flop
rate for specific membrane lipids; thus, the lipid imbalance can be compensated and
the membrane asymmetry concerning specific membrane lipids can be established.
Furthermore, the ER can form transport vesicles responsible for the transfer of
membrane substances and proteins to the Golgi complex.
The Golgi complex (also called Golgi apparatus), usually located in the vicinity of
the nucleus, consists of piles of several flat membrane cisternae. ER transport vesi-
cles enter these piles at its cis-side. Substances leave the Golgi complex at the oppo-
site trans-site. Transport between the different cisternae is mediated by Golgi vesi-
cles. Some modifications of proteins by the addition of a specific oligosaccharide
happen in the ER, but further glycosylations of various types take place in the lumen
of the Golgi complex. Since such modified membrane proteins and lipids point to
the organelles’ inner space, they will be exposed to the cell’s outer space when they
are transported to the plasma membrane. The synthesis of complex modifications by
several additions of carbohydrates requires a special enzyme for each specific addi-
tion. Therefore, these reaction pathways become very complex.
2.3.6
Other Organelles
Eukaryotic cells have further compartments for certain functions. Some of these or-
ganelles and their major functions will be mentioned briefly here.
Lysosomes are responsible for the intracellular digestion of macromolecules.
These vesicular organelles contain several hydrolyzing enzymes (hydrolases), e.g.,
proteases, nucleases, glycosidases, lipases, phosphatases, and sulfatases. All of them
have their optimal activity at pH 5. This pH value is maintained inside the lysosomes
via ATP-dependent H
+
pumps (for comparison, the pH of the cytosol is about 7.2).
Peroxisomes (also called microbodies) contain enzymes that oxidize organic sub-
stances (R) and therefore use molecular oxygen as an electron acceptor. This reaction
produces hydrogen peroxide (H
2
O
2
).
RH
2
+O
2
? R+H
2
O
2
H
2
O
2
is used by peroxidase to further oxidize substances such as phenols, amino
acids, formaldehyde, and ethanol, or it is detoxified by catalase (2 H
2
O
2
? 2H
2
O+
O
2
).
In contrast to the ER, the Golgi cisternae, lysosomes, peroxisomes, and vesicles,
which are surrounded by a single membrane, chloroplasts as well as mitochondria
have a double membrane of which the inner one is not folded into cristae as in mito-
chondria. Instead, a chloroplast has a third membrane that is folded several times
and forms areas that look like piles of coins. This membrane contains light-harvest-
ing complexes and ATP synthases that utilize the energy of the sunlight for the pro-
duction of cellular energy and reduction equivalents used for the fixation of carbon
dioxide (CO
2
) into sugars, amino acids, fatty acids, or starch. Chloroplasts, as well as
mitochondria, have their own circular DNA and ribosomes.
44
2 Biology in a Nutshell

2.4
Expression of Genes
Classically, a gene is defined as the information encoded by the sequence of a DNA
region that is required for the construction of an enzyme or – more generally – of a
protein. We will see that this is a simplified definition, since, e.g., mature products
of some genes are not proteins but RNAs with specific functions; eukaryotic gene se-
quences in particular also contain non-coding information. The term gene expres-
sion commonly refers to the whole process during which the information of a parti-
cular gene is translated into a particular protein. This process involves several steps
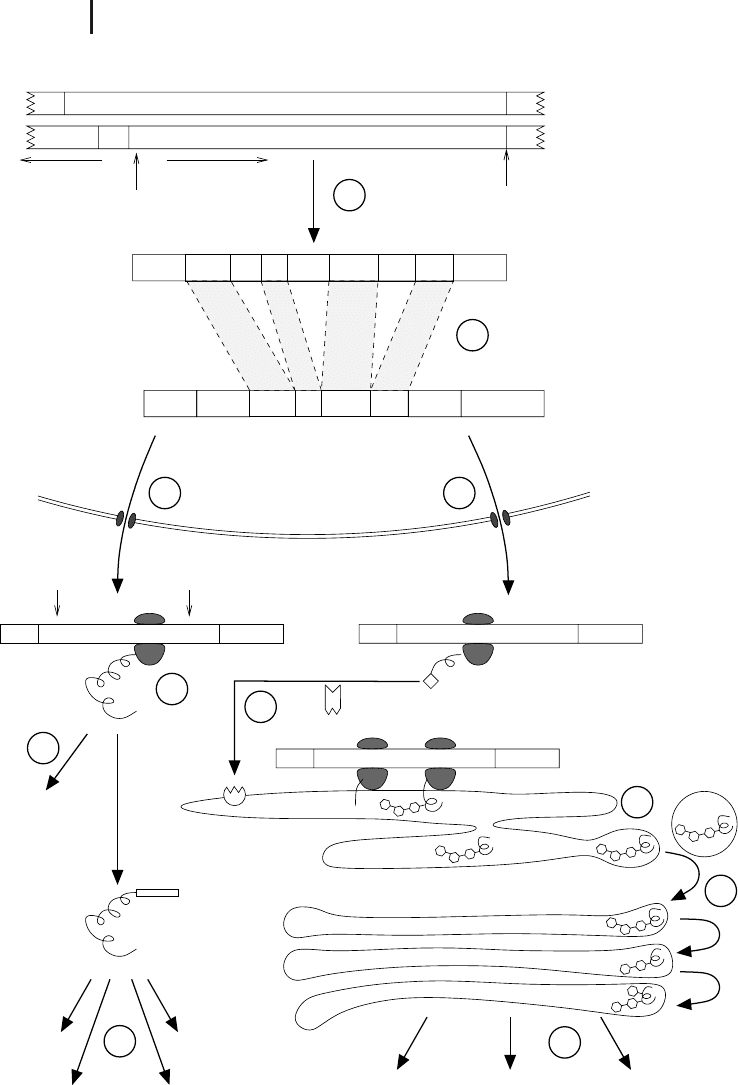
(cf. Chapter 8). First, during transcription (Fig. 2.11 1), the DNA region encoding
the gene is transcribed into a complementary messenger RNA (mRNA). In eukaryo-
tic cells this mRNA is further modified (2) inside the nucleus and transferred to the
cytosol (3). In the cytosol, the mRNA binds to a ribosome that uses the sequence as
a template for the synthesis of a specific polypeptide that can fold into the three-di-
mensional protein structure (4). In prokaryotic cells the mRNA is not further modi-
fied and ribosomes can bind to the nascent mRNA during transcription.
In eukaryotic cells the synthesized proteins can either remain in the cytosol (5)
or, if they have a specific signaling sequence, be synthesized by ribosomes of the
rough ER and enter its lumen (7). However, there are several mechanisms of direct-
ing each protein to its final destination. During this sorting, proteins are often modi-
fied, e.g., by cleavage of signaling peptides or by glycosylation.
All the genes of a single organism make up its genome. But only a subset of these
genes will be expressed at a particular time or in a specific cell type. Some genes ful-
fill basic functions of the cell and are always required; these are called constitutive or
housekeeping genes. Others are expressed only under certain conditions. The
amount of a gene product, e.g., a protein, depends mainly on its stability and the
number of its mRNA templates. The number of the latter depends on the transcrip-
tion rate, which is influenced by regulatory regions of the gene and transcription fac-
tors that control the initialization of transcription. Thus, quantitative changes in
gene expression can be monitored by mRNA and protein concentrations (see Chap-
ter 4 on experimental techniques used for this purpose). Rate changes in any produc-
tion or degradation step of a specific gene, which might happen in different cell
types or at developmental stages, can lead to differential gene expression.
The whole procedure of gene expression, protein sorting, and posttranslational
modifications is summarized in Fig. 2.11 and will be described in more detail in the
following section.
2.4.1
Transcription
The synthesis of an RNA polymer from ATP, GTP, CTP, and UTP employing a DNA
region as a template is called transcription. RNA synthesis is catalyzed by the RNA
polymerase. In eukaryotic cells there are different types of this enzyme that are re-
sponsible for the synthesis of different RNA types, including messenger RNA
45
2.4 Expression of Genes
46
2 Biology in a Nutshell
5’ UTR
pre−mRNA
Mature mRNA
Nuclear pore
5’ cap
Transcription end
upstream downstream
DNA sense strand
5’ UTR
Start codon
Nascent protein
Stop codon
5’ cap AUG Stop
5’ cap AUG
5’ 3’
2
1
10
3’ UTR
Endoplasmatic
reticulum (ER)
StopStop
5’
3’
Transcription start
3’
5’
Nuclear envelope
poly(A) tail
Signal peptide
7
poly(A) tail
poly(A) tail
8
9
Ribosome
5’ cap AUG
Signal peptides
Golgi complex
SRP
6
5
4
3
3
3’ UTR poly(A) tail
DNA antisense strand
E
2
E
1
E
4
E
3
E
3
E
2
I
2
I
1
E
1
I
3
E
4
Mature
cytosolic
protein
Plasma
membrane
Chloroplast
LysosomeSecretory vesicles
Peroxisome
MitochondrionNucleus
PR

(mRNA), ribosomal RNA (rRNA), or transfer RNA (tRNA). In prokaryotic cells all
these different RNA types are synthesized by the same polymerase. This enzyme has
an affinity to a specific DNA sequence, the promoter, that also indicates the first base
to be copied. During initiation of transcription, the RNA polymerase binds to the pro-
moter with a high affinity that is supported by further initiation factors. Complete for-
mation of the initiation complex causes the DNA to unwind in the promoter region.
Now the enzyme is ready to add the first RNA nucleoside triphosphate to the template
strand of the opened DNA double strand. In the subsequent elongation phase, the
RNA polymerase moves along the unwinding DNA and extends the newly developing
mRNA continuously with nucleotides complementary to the template strand. During
this phase a moving transient, double-stranded RNA-DNA hybrid is established. As
the polymerase moves along, the DNA rewinds again just behind it. As RNA synthesis
always proceeds in 5'?3' direction, only one of the DNA chains acts as a template,
the so-called antisense (–) strand. The other one, the sense (+) strand, has the same se-
quence as the transcribed RNA, except for the thymine nucleotides that are replaced
by uracil nucleotides in RNA. As much as the promoter is responsible for initiation of
transcription, the terminator – another specific DNA sequence – is responsible for its
termination. For the bacterium E. coli two different termination mechanisms are de-
scribed: the Rho-independent and Rho-dependent terminations. In Rho-independent
termination, the transcribed terminator region shows two short GC-rich and self-com-
plementary sequences that can bind to each other and thus form a so-called hairpin
structure. This motif is followed by a block of uracil residues that bind the comple-
mentary adenine residues of the DNA only weakly. Presumably, this RNA structure
causes the RNA polymerase to terminate and release the RNA. In Rho-dependent ter-
mination, a protein – the Rho factor – can bind the newly synthesized RNA near the
terminator and mediate the RNA release. Termination in eukaryotic cells shows both,
similarities to and differences from the mechanisms found in bacteria.
2.4.2
Processing of the mRNA
In eukaryotic cells the primary mRNA transcript (precursor mRNA or pre-mRNA) is
further processed before being exported into the cytosol and entering translation (Fig.
2.11 2). The protein-coding sequence lies internally in the mRNA and is flanked on
both sides by nucleotides that are not translated. During processing, a so-called 5' cap
is attached to the flanking 5' untranslated region (5' UTR, about 10 to 200 nucleotides)
preceding, or lying upstream of, the coding sequence. This 5' cap consists of three nu-
cleotides that are further modified. The 3' untranslated region (3' UTR) of most
mRNAs is also modified after transcription by addition of a series of about 30 to 200
47
2.4 Expression of Genes
3 Fig. 2.11 Gene expression in eukaryotic cells
comprises several steps from the DNA to the
mature protein at its final destination. This in-
volves the 1 transcription of the gene, 2 spli-
cing and processing of the pre-mRNA, 3 export
of the mature mRNA into the cytosol, 4 trans-
lation of the genetic code into a protein, and
5–0 several steps of sorting and modification.
More details are given in the text.

adenine nucleotides that are known as the poly(A) tail. Furthermore, the pre-mRNA is
often much longer than the mature RNA because the coding sequence is often inter-
rupted by one or several intervening sequences called introns, which do not occur in
the mature mRNA exported to the cytosol. These intron sequences are removed during
processing by a mechanism called splicing. The remaining sequences are called exons.
The final coding sequence thus consists of a series of exons joined together. It starts
with AUG, which is the first triplet translated into an amino acid, and it stops with a
stop codon (UGA, UAA, or UAG). Via the pores of the nuclear envelope, the mature
mRNA is finally exported to the cytoplasm,where the translation process takes place.
2.4.3
Translation
Translation of the genetic information encoded by the mRNA into the amino acid se-
quence of a polypeptide is done by ribosomes in the cytosol. To encode the 20 differ-
ent amino acids occurring in polypeptides, at least three bases out of the four possi-
bilities (G, U, T, C) are necessary (4
3
= 64 > 20). During evolution, a code developed
that uses such triplets of exactly three bases, which are called codons, to code the
amino acids and signals for start and end of translation. By using three bases for
each codon, more than 20 amino acids can be coded, and hence some amino acids
are encoded by more than one triplet. The genetic code is shown in Tab. 2.3. It is
48
2 Biology in a Nutshell
Tab. 2.3 The genetic code. Each codon of the genetic code – read in the 5'?3' direction along
the mRNA – encodes a specific amino acid or a starting or termination signal of translation.
Position 1 Position 2 Position 3
(5' end) (3' end)
UCAG
Phe Ser Tyr Cys U
U Phe Ser Tyr Cys C
Leu Ser Stop Stop A
Leu Ser Stop Trp G
Leu Pro His Arg U
C Leu Pro His Arg C
Leu Pro Gln Arg A
Leu Pro Gln Arg G
Ile Thr Asn Ser U
A Ile Thr Asn Ser C
Ile Thr Lys Arg A
Met Thr Lys Arg G
Val Ala Asp Gly U
G Val Ala Asp Gly C
Val Ala Glu Gly A
Val Ala Glu Gly G
highly conserved across almost all prokaryotic and eukaryotic species except for
some mitochondria or chloroplasts. For translation of the genetic information, adap-
ter molecules are required. These are the transfer RNAs (tRNAs). They consist of
about 80 nucleotides and are folded into a characteristic form similar to an “L”. Each
tRNA can recognize a specific codon by a complementary triplet, called an antico-
don, and it can also bind the appropriate amino acid. For each specific tRNA, a cer-
tain enzyme (aminoacyl-tRNA synthetase) attaches the right amino acid to the
tRNA’s 3‘ end. Such a loaded tRNA is called an aminoacyl-tRNA.
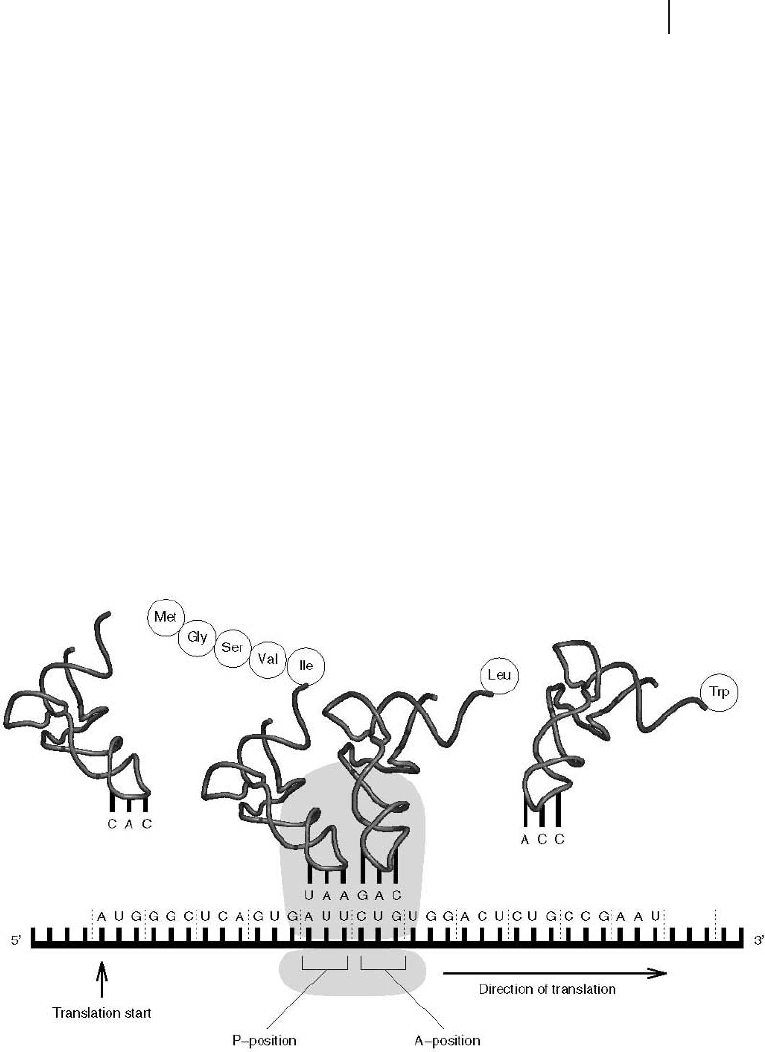
During translation (Fig. 2.12), the genetic information of the mRNA is read co-
don by codon in a 5'?3' direction of the mRNA, starting with an AUG codon.
AUG codes for methionine, and therefore newly synthesized proteins always begin
with this amino acid at their amino terminus. Protein biosynthesis is catalyzed by
ribosomes. Both eukaryotic and prokaryotic ribosomes consist of a large and a
small subunit, and both subunits are composed of several proteins and rRNAs. In
eukaryotic cells the small ribosomal subunit first associates with an initiation
tRNA (Met-tRNA
i
) and binds the mRNA at its 5' cap. Once attached, the complex
scans along the mRNA until reaching the start AUG codon. In most cases this is
the first AUG codon in the 5'?3' direction. This position indicates the translation
start and determines the reading frame. Finally, during initiation the large riboso-
mal subunit is added to the complex and the ribosome becomes ready for protein
synthesis. Each ribosome has three binding sites: one for the mRNA and two for
tRNAs. In the beginning the first tRNA binding site, also called P site, contains
49
2.4 Expression of Genes
Fig. 2.12 During translation the genetic information of the mRNA
is converted into the corresponding polypeptide. More details are
given in the text.
