Molina-Par?s C., Lythe G. (editors) Mathematical Models and Immune Cell Biology
Подождите немного. Документ загружается.


5 The Cyton Model 113
First and Subsequent Divisions and Progressor Fraction
Although the PDFs for death and division may, in principle, vary for each division,
we find that for most cases it is sufficient to distinguish between undivided cells,
and those in subsequent generations. Detailed analysis of a CpG-stimulated B cell
approach in vitro shows that the PDFs for death and division times vary only slightly
with generation after initial division [8].
We introduced the concept of the progressor fraction and discussed its division
dependence. Here we apply a functional form to this division dependence, thus re-
ducing the number of free parameters that are necessary to specify the progressor
fraction in the model. The form used in [4], is based on the observation that the
ultimate division number achieved by a cohort of cells is approximately normally
distributed over the generations [5]. As is the case with death and division times, the
undivided cells behave in a way that is different from cells in subsequent genera-
tions, and for this reason we let the progressor fraction for undivided cells remain a
free parameter unconstrained by the functional form used for the subsequent gener-
ations. This implies that the fraction of cells in generation i that are division capable
is given by
i
D
8
ˆ
ˆ
ˆ
ˆ
ˆ
<
ˆ
ˆ
ˆ
ˆ
ˆ
:
0
if i D 0,
Z
1
iC1
N .n;
dest
;
dest
/dn
Z
1
i
N .n;
dest
;
dest
/dn
if i>0,
(5.5)
where
dest
;
dest
are the mean and standard deviation of the Gaussian PDF that
describes division destiny:
N .n;;/ D
1
p
2
2
exp
.n /
2
2
2
:
Comparison with Experiment
This gives a model with 11 free parameters, listed in Table 5.1, which are obtained
empirically by fitting to experimental data in the form of CFSE timecourses. The
values of these parameters will be dependent on experimental conditions such as the
strength and type of stimulus, as well as the presence of different cytokines and their
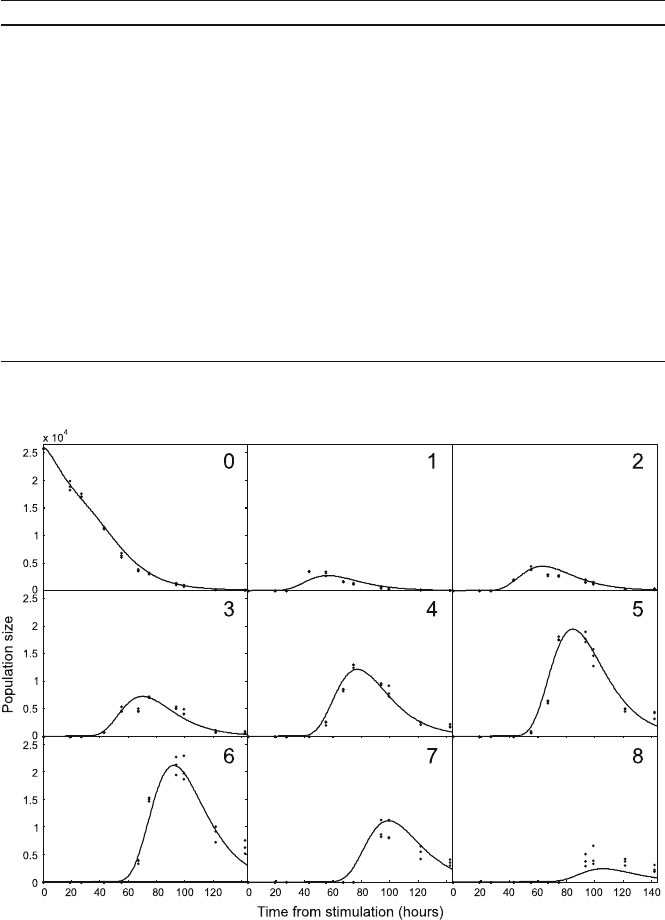
concentrations. In Fig. 5.4 we show a comparison between the best-fit model pre-
diction for the population of cells in each division compared with experimental time
course. The data was obtained from CFSE analysis of a system of LPS stimulated
B cells. The best-fit values for the free parameters were obtained by minimising the
sum of the squared residuals between predicted and measured values.
114 C. Wellard et al.
Table 5.1 A list of free parameters used in the cyton model of lymphocyte proliferation
Parameter Description Parameter Description
0
B
Mean time to first division
C
D
Mean time to die for cells
in subsequent divisions
0
B
Standard deviation of the time
to first division
C
D
Standard deviation of the time
to die in subsequent division
0
D
Mean time to die for undivided
cells
0
Progressor fraction for undivided
cells
0
D
Standard deviation of the time
to die for undivided cells
dist
Mean generation number for the
PDF characterising the division
destiny
C
B
Mean time to subsequent division
dist
Standard deviation of the generation
number for the PDF
characterising the division
destiny
C
B
Standard deviation of the time
to subsequent division
Fig. 5.4 A fit to time course data obtained from CFSE analysis of LPS stimulated B cells. Here
we show the best-fit prediction for the populations of each generation as a function of time (line),
compared to the measured values (points)
5 The Cyton Model 115
Stochastic Variation in Population Number
Although the cyton model is based on the idea that the fundamental cellular
processes of death and division are inherently stochastic, we have only presented
a solution for the average behaviour of the system. The solution of these equa-
tions gives an estimate for the number of cells that would be expected over many
replicates of an identical experiment, but does not give any information about fluctu-
ations in this result due to the stochastic nature of the underlying processes. In large
systems the variation between replicates due to these fluctuations will usually be
small, generally much lower than the variation due to experimental considerations,
however in small systems these stochastic fluctuations can be observed. In [8], the
authors present multiple time course experiments obtained from the time-lapse mi-
croscopy of CpG-stimulated B cells. In each case the clone produced from a single
progenitor was followed throughout the duration of the response, and the number of
cells in each generation recorded as the response progressed. In this experiment sig-
nificant variation was observed in the population sizes of the different clones. While
the model as formulated in (5.4) is sufficient to reproduce the average behaviour of
these clones, it is not sufficient to encapsulate the stochastic variation. To do this
requires a new approach.
Branching Process Models
The theory of branching process [9] has been used extensively in mathematical
biology to simulate cell population dynamics, from discrete time models [10]to
more complex continuous time models [11–13]. Here we use a generalisation of the
Bellman–Harris model to obtain a set of recursion relations for the probability gen-
erating function (PGF) for the number of cells in each generation as a function of
time. This PGF can be used to calculate the mean population dynamics, as well as
the expected stochastic fluctuations around this mean.
We begin by defining the random variable Z
i
.t/ to denote the number of live
cells in generation i at time t. The PGF for the number of cells in each generation,
given a single cell in generation i at time t D 0 is defined as
F
i
.s;t/ D E
"
Y
i
0
s
Z
i
0
.t/
i
0
ˇ
ˇ
ˇ
Z
i
.0/ D 1; Z
j
.0/ D 0 8j ¤ i
#
; (5.6)
where EŒx denotes the expectation value of the random variable x. Following the
prescription for a generalised Bellman–Harris model [12,13] gives a recursion rela-
tion for this PGF
F
i
.s;t/ D .1Q
B
i
.t//.1Q
D
i
.t//s
i
CQ
D
i
.t/C
Z
t
0
F
iC1
.s;t/
2
q
B
i
./d: (5.7)

116 C. Wellard et al.
Here the q
B
i
.t/; q
D
i
.t/ are the effective probability distributions defined in (5.1),
and Q
B
i
.t/; Q
D
i
.t/ are the corresponding cumulative functions. The first term in
the above equation takes into account the case in which the cell has neither died
nor divided, the contribution for this case is s
i
. The second term takes into account
the case in which the cell has died, here the contribution is one. Finally the third
term takes into account the cases in which the cell divided at a time t ,andthe
contribution here is equal to the product of two PGFs for cells in the subsequent
generation,born at t .
Mean Population Dynamics
The mean dynamics of the model can be obtained using
j
i
.t/ D
@F
i
.s;t/
@s
j
ˇ
ˇ
ˇ
sD0
,where
j
i
.t/ gives the expected number of cells of generation j at time t given a single
progenitor of generation i at t D 0. When applied to (5.7)thisgives
j
i
.t/ D .1 Q
B
i
.t//.1 Q
D
i
.t//ı
i;j
C 2
Z
t
0
j
iC1
.t /q
B
i
./d: (5.8)
This recursion relation can be evaluated by noting that
j
i
.t/ D0; 8j<i, and by
assuming some limit on the number of divisions that a cell has undergone. This limit
may be imposed either by a progressor fraction that effectively vanishes beyond a
certain generation, or by assuming a minimum division time for cells.
Stochastic Fluctuations
The advantage of the branching process approach is that, in principle, it is possible
to calculate the entire probability distribution for the populations size at any point
in time. In practice this is quite difficult, and usually unnecessary. A much easier
prospect, and one that offers great utility, is to calculate the variance in population
size. This variance will give a measure of the uncertainty in the population size
resulting from the stochastic nature of the underlying processes. To calculate the
variance, we begin with the second derivative of the PGF
j;k
i
.t/ D
@
2
F
i
.s;t/
@s
k
@s
j
ˇ
ˇ
ˇ
sD0
.
When applied to (5.7)thisgives
j;k
i
.t/ D 2
Z
t
0
j;k
iC1
.t / C
j
iC1
.t /
k
iC1
.t /
q
B
i
./d: (5.9)
From this we can use the relations
j;k
i
.t/ D
(
j;k
i
.t/
j
i
.t/
k
i
.t/ j ¤ k
j;j
i
.t/ C
j
i
.t/
j
i
.t/
2
j D k
(5.10)
5 The Cyton Model 117
to calculate the covariances of the population of cells in generations j; k at time t
resulting from an initial cell of generation i.
Differentiation
Branching process models have been used to model differentiation in neuronal
cells [11]. Here we extend our model to include possible differentiation processes
for lymphocytes. The cyton model can be generalised to treat general differentiation
processes in the same way as it treats death and division. To motivate this generali-
sation we note that various differentiation events in activated B cells, such as isotype
switching and commitment to antibody-secreting cells, have been shown to be divi-
sion linked [14,15]. Similarly we assume that the death and division processes are
dependent on the differentiation state, or type of the cell, and that the probability of
differentiating to a particular state is dependent on the current state of the cell.
Thus we hypothesise the existence of a molecular mechanism responsible for
each possible differentiation process, that acts independently of all other process,
and acts in a stochastic manner. In analogy to death and division these processes
can be characterised by a PDF for the differentiation time, which is dependent on
the state of the cell, as well as its generation number. We define the PDF for time-
to-differentiation from type j to type k for a cell in division i as p
k
i;j
.t/. In analogy
with progressor fraction, we define a division dependent differentiation fraction
k
i;j
which gives the fraction of cells of type j that are capable of differentiation to type
k in division i. In this case the effective distributions, which give the probability
density at time t,aregivenby
q
B
i;j
.t/ D
i;j
p
B
i;j
.t/
1 P
D
i;j
.t/
Y
k
1
k
i;j
P
k
i;j
.t/
;
q
D
i;j
.t/ D p
D
i;j
.t/
1
i;j
P
B
i;j
.t/
Y
k
1
k
i;j
P
k
i;j
.t/
;
q
k
i;j
.t/ D
k
i;j
p
k
i;j
.t/.1 P
D
i;j
.t//
1
i;j
P
B
i;j
.t/
Y
k
0
¤k;j
1
k
0
i;j
P
k
0
i;j
.t/
:
Here P.t/ denotes the cumulative distribution of the relevant PDF.
Using the same methodology as previously we can derive a recursion relation for
the PGF
F
i;j
.s;t/ D
1 Q
B
i;j
.t/
1 Q
D
i;j
.t/
Y
k¤j
1 Q
k
i;j
.t/
s
i;j
C Q
D
i;j
.t/
C
Z
t
0
F
iC1;j
.s;t /
2
q
B
i;j
./d C
X
k¤j
Z
t
0
F
i;k
.s;t /q
k
i;j
./d:
(5.11)
118 C. Wellard et al.
As previously, we can use this relation to derive an expression for the expected
population of cells of type j
0
in generation i
0
given an initial cell of type j in
generation i
i
0
;j
0
i;j
.t/ D .1 Q
B
i;j
.t//.1 Q
D
i;j
.t//
Y
k¤j
.1 Q
k
i;j
.t//ı
i;i
0
ı
j;j
0
C2
Z
t
0
i
0
;j
0
iC1;j
.t /q
B
i;j
./d C
X
k¤j
Z
t
0
i
0
;j
0
i;k
.; t /q
k
i;j
./d:
Similarly we can use
i
0
;j
0
Ii
00
;j
00
i;j
.t/ D 2
Z
t
0
i
0
;j
0
iC1;j
.t /
i
00
;j
00
iC1;j
.t / C
i
0
;j
0
Ii
00
;j
00
iC1;j
.t /
q
B
i;j
./d
C
X
k¤j
Z
t
0
i
0
;j
0
Ii
00
;j
00
i;k
.t /q
k
i;j
./d;
to calculate values for the covariances.
Simulation
The methods mentioned so far require numerical solution of a set of deterministic
integral equations for the mean number of cells or the variance. Higher order mo-
ments can also be calculated however each comes at further computational expense
to the previous. An alternative approach is to simulate the cells directly. Although
this approach is generally slower to calculate the mean value, there is no further cost
in calculating the variance and higher moments. Further, it is more flexible and can
be quickly adapted to different sorts of models. Also it can be readily extended to
provide readouts of common experimentally observed quantities.
Modelling Cell Properties
Each cell being simulated carries with it a set of numbers which represent its physi-
cal state. In some cases these numbers are decided when the cell is created, in others
they vary over (simulated) time. The following is a non-exhaustive list of what might
be stored:
– When the cell was created. This allows age related properties of the cell to be
calculated.
5 The Cyton Model 119
– A unique ID. This way relationships between cells can be kept so that correlations
and clonal effects can be explored.
– Cell fate. In the case of the Cyton model, whether a cell dies or divides and when
this happens can be decided at the time a cell is created.
– Cell type. Whether the cell has committed to becoming antibody secreting cell
and if isotype switching has occurred.
– Stage of cell cycle and cell size. By storing where a cell is in the cell cycle
the amount of DNA can be calculated. This allows comparison to be made with
flow cytometry techniques that can measure DNA content such as DAPI and
Hoechst stains which bind to the DNA in a stoichiometric manner. Estimating
cell size allows connection to be made with forward scatter measurements from
flow cytometry.
– Fluorophore content. If the stage of the cell cycle and DNA content are known
then it is also possible to make connection with Bromodeoxyuridine (BrdU)
experiments. BrdU is a molecule that can replace thymidine during DNA repli-
cation. In BrdU experiments, cells’ exposure to BrdU is limited to certain times.
Its incorporation can be measured using immunohistochemistry techniques. Nor-
mally it is used to provide confirmation that cells are dividing during a particular
time (when BrdU is present) but potentially it can be used to provide more infor-
mation about the parameters for a model such as Cyton [16–18].
– Affinity for antigen. This enables the models to be implemented which link a
cell’s ability to proliferate to its affinity, thereby addressing repertoire issues.
Implementation
In practice the simulation of cells is done using a discrete event simulator. This
requires the maintenance of a list of cells and their associated states and a time-
ordered list of events that can occur to them. Simulation is done asynchronously.
That is, instead of stepping through time, the programme processes a sequence of
queued events. In this way, time resolution can be made arbitrarily small (within
machine precision) without incurring computational expense. Examples of events
that need to be queued are
– Cell division and death
– Changing of some cell property such as a cell type and phase
– Making measurement corresponding to those made at experimental time points
Assuming that there are no interactions between cells then the computational cost
of this method scales no worse than O.N/ times the cost of insertion into the event
list where N is the number of cells.
120 C. Wellard et al.
References
1. Lyons A, Parish C (1994) Determination of lymphocyte division by flow cytometry. J Immunol
Methods 171:131–137
2. Hasbold J, Gett A, Rush J, Deenick E, Avery D, Jun J, Hodgkin P (1999) Quantitative analy-
sis of lymphocyte differentiation and proliferation in vitro using carboxyfluorescein diacetate
succinimidyl ester. Immunol Cell Biol 77:516–522
3. Gett A, Hodgkin P (2000) A cellular calculus for signal integration by T cells. Nat Immunol
1:239–244
4. Hawkins E, Turner M, Dowling M, van Gend C, Hodgkin P (2007) A model of immune regu-
lation as a consequence of randomized lymphocyte division and death times. Proc Natl Acad
Sci USA 104:5032–5037
5. Turner M, Hawkins E, Hodgkin P (2008) Quantitative regulation of B cell division destiny by
signal strength. J Immunol 181:374–382
6. Boer RD, Ganusov V, Milutinovic D, Hodgkin P, Perelson A (2006) Estimating lymphocyte
division and death rates from CFSE data. Bull Math Biol 68:1011–1031
7. Leon K, Faro J, Carneiro J (2004) A general mathematical framework to model generation
structure in a population of asynchronously dividing cells. J Theor Biol 229:455–476
8. Hawkins E, Markham J, McGuinness L, Hodgkin P (2009) A single-cell pedigree analysis of
alternative stochastic lymphocyte fates. Proc Natl Acad Sci USA 106:13457
9. Kimmel M, Axelrod DE (2001) Branching processes in biology. Springer, New York
10. Yates A, Chan C, Strid J, Moon S, Callard R, George A, Stark J (2007) Reconstriction of cell
population dynamics using CFSE. BMC Bioinformatics 8:196
11. Yakovlev A, Mayer-Proschel M, Noble M (1998) A stochastic model of brain cell differentia-
tion in tissue culture. J Math Biol 37:49
12. Hyrien O, Mayer-Pr¨oschel M, Noble M, Yakovlev A (2005) A stochastic model to analyze
clonal data on multi-type cell populations. Biometrics 61:199–207
13. Subramanian V, Duffy K, Turner M, Hodgkin P (2008) Determining the expected variability
of immune responses using the cyton model. J Math Biol 56:861–892
14. Hodgkin P, Lee J, Lyons A (1996) B cell differentiation and isotype switching is related to
division cycle number. J Exp Med 184:277–281
15. Hasbold J, Corcoran L, Tarlinton D, Tangye S, Hodgkin P (2004) Evidence from the gen-
eration of immunoglobulin G-secreting cells that stochastic mechanisms regulate lymphocyte
differentiation. Nat Immunol 5:55–63
16. Yanagisawa M, Dolbeare F, Todoroki T, Gray J (1985) Cell cycle analysis using numerical
simulation of bivariate DNA/bromodeoxyuridine distributions. Cytometry 6:550
17. Cain S, Chau P (1997) A transition probability cell cycle model simulation of bivariate
DNA/bromodeoxyuridine distributions. Cytometry 3:239
18. Hodgkin PD, Hawkins ED, Hasbold J, Gett AV, Deenick EK, Todd HF, Hommel M (2006)
Monitoring T cell proliferation. In: Nagorsen D, Marincola FM (eds) Analyzing T Cell
Responses: how to analyze cellular immune responses against tumor associated antigens.
Springer, chapter 6, pp 123–142

Chapter 6
Modelling Intravital Two-Photon Data
of Lymphocyte Migration and Interaction
Marc Thilo Figge and Michael Meyer-Hermann
Abstract Multi-photon microscopy is a powerful tool for imaging lymphocyte
migration and interaction in intact biological organs. These experiments gener-
ate quantitative data on the cell motility, shape dynamics, and contact duration of
cellular interactions. In this chapter we review mathematical models that have suc-
cessfully contributed to the interpretation of these data with regard to lymphocyte
migration and interaction. The examples involve different modelling approaches and
range from T cell priming in secondary lymphoid tissue to B cell affinity maturation
in germinal centres.
Introduction
Adaptive immunity implies the diligent communication of specific information that
is facilitated by lymphocyte migration in lymphoid tissue and their interaction with
other components of the immune system. For example, T cells scan antigen present-
ing cells and may become activated if the T cell receptors recognize specific foreign
peptides embedded in major histocompatibility molecules. T cell priming by den-
dritic cells in lymph nodes occurs in subsequent phases that clearly differ in the T
cell behaviour with regard to migration and interaction [1]. Differences in the mi-
gration behaviour of the distinct types of T cells in diverse lymphoid tissues remain
to be explained [2].
The salient feature of B cells is that the B cell receptors undergo affinity matu-
ration in order to optimize the immune response against pathogenic antigens. This
process takes place in follicular structures of lymphoid organs, referred to as ger-
minal centres [3, 4], and involves intercellular interactions in the selection process
for high-affinity B cells. Germinal centres have a peculiar morphology consisting of
M.T. Figge (
)
Applied Systems Biology, Friedrich Schiller University Jena, Leibniz Institute
for Natural Product Research and Infection Biology – Hans Kn¨oll Institute,
Beutenbergstrasse 11a, 07745 Jena, Germany
e-mail: thilo.figge@hki-jena.de
C. Molina-Par´ıs and G. Lythe (eds.), Mathematical Models and Immune Cell Biology,
DOI 10.1007/978-1-4419-7725-0
6,
c
Springer Science+Business Media, LLC 2011
121
122 M.T. Figge and M. Meyer-Hermann
two distinct zones, termed the dark and the light zone. The dark zone is the region
of B cell proliferation, while in the light zone B cells undergo selection. How B cell
migration between the zones is realized is still a matter of debate today [5–8].
For the last few years, lymphocyte migration and interaction have been exten-
sively explored by non-invasive intravital microscopy [9]. The main achievement
of this technique is that lymphocytes can be followed in real time and within their
natural cellular environment. The generated data include records of cell tracks, i.e.
the time-dependent position of single cells, as well as cellular contact times of in-
teractions. The working principle of imaging by multi-photon microscopy is briefly
summarized, together with an overview of dynamical cell properties that can be ex-
plored by this experimental technique. The rapidly growing body of experimental
data is calling for mathematical methods that are suitable for analyzing these data
in order to extract meaningful conclusions initiating new insights and experiments.
The art of mathematical modelling is to choose an adequate modelling approach.
This process starts by specifying the questions to be answered and continues by
identifying all variables and connections between them that are relevant for answer-
ing these questions. Each modelling approach has its assets and drawbacks. As a
rule of thumb, computationally cheap methods often only give a very rough repre-
sentation of reality and, therefore, they often only provide answers to questions of a
very general kind. Questions with regard to specific system properties typically re-
quire detailed calculations involving sophisticated computational methods. In many
cases, however, a detailed modelling approach also demands knowledge about pro-
cesses and parameters that are not yet accessible experimentally. Estimating the
quantitative impact of these processes ultimately leaves the questioner with the task
to decide whether or not the obtained answers are reasonable. Ideally, different mod-
elling approaches can be combined where answers to questions obtained at a certain
level of resolution serve as input for modelling at a level of higher resolution.
We consider three modelling approaches of lymphocyte migration and interac-
tion that differ in the level of resolution with respect to the functional properties of
the cells. The first is a statistical modelling approach for the analysis of cell tracks as
observed in two-photon experiments. Single cells are viewed as independent point
particles and the only functional aspect that is taken into account is their ability to
migrate. Next, we consider cellular systems in which different types of cells migrate
and interact with each other. This is captured by an agent-based modelling approach,
where each cell represents a discrete agent that interacts with other cells and is mon-
itored during its whole lifetime. Finally, the agent-based modelling approach is used
with a sufficiently high spatial resolution to go beyond the point particle represen-
tation of cells in order to study the dynamics of cellular shape deformations under
migration and interaction.
