Hogg S. Essential microbiology
Подождите немного. Документ загружается.

JWBK011-11 JWBK011-Hogg August 12, 2005 19:21 Char Count= 0
268

JWBK011-11 JWBK011-Hogg August 12, 2005 19:21 Char Count= 0
11
Microbial Genetics
A gene is a sequence
of DNA that usually en-
codes a polypeptide.
When any living organism reproduces, it passes on
genetic information to its offspring. This information
takes the form of genes, linear sequences of DNA that
can be thought of as the basic units of heredity. The total
complement of an organism’s genetic material is called
its genome.
How do we know genes
are made of DNA?
The concept of the gene as an inherited physical entity determining some aspect of
an organism’s phenotype dates back to the earliest days of genetics. The question of
what genes are actually made of was a major concern of molecular biologists (not that
they would have described themselves as such!) in the first half of the 20th century.
Since it was recognised by this time that genes must be located on chromosomes, and
that chromosomes (in eucaryotes) comprised largely protein and DNA, the reasonable
assumption was made that genes must be made up of one of these substances. In the
early years, protein was regarded as the more likely candidate, since, from what was
known of molecular structure at the time, it offered far more scope for the variation
which would be essential to account for the thousands of genes that any organism must
possess. The road to proving that DNA is in fact the ‘stuff of life’ was a long and hard
one, which can be read about elsewhere; we shall mention below just some of the key
experiments which provided crucial evidence.
In 1928 the Englishman Fred Griffith carried out a seminal series of experiments
which not only demonstrated for the first time the phenomenon of genetic transfer
in bacteria (a subject we shall consider in more detail later in this chapter), but also
acted as the first step towards proving that DNA was the genetic material. As we shall
see, Griffith showed that it was possible for heritable characteristics to be transferred
from one type of bacterium to another, but the cellular component responsible for this
phenomenon was not known at this time.
Attempts were made throughout the 1930s to isolate and identify the transform-
ing principle, as it became known, and in 1944 Avery, MacLeod and McCarty pub-
lished a paper, which for the first time, proposed DNA as the genetic material. Avery
and his colleagues demonstrated that when DNA was rendered inactive by enzy-
matic treatment, transforming ability was lost from a cell extract, but if proteins,
269
JWBK011-11 JWBK011-Hogg August 12, 2005 19:21 Char Count= 0
270 MICROBIAL GENETICS
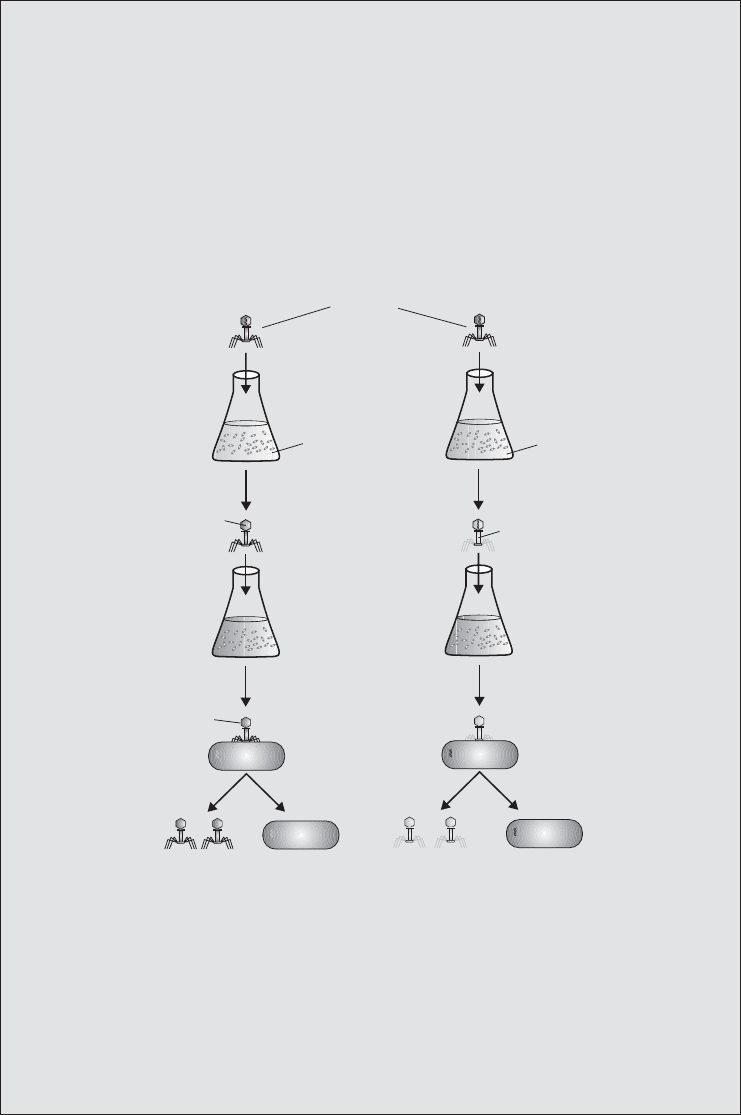
Box 11.1 Hershey and Chase: the Waring blender experiment
In 1952 Alfred Hershey and Martha Chase provided further experimental evidence
that DNA was the genetic material. In their experiment, the bacteriophage T2 was
grown with E. coli cells in a medium with radiolabelled ingredients, so that their pro-
teins contained
35
S, and their DNA
32
P. The phages were harvested, and mixed with
a fresh culture of E. coli. They were left long enough for the phage particles to infect
the bacteria, (but not long enough to produce new phage particles and lyse the
cells). The culture was then subjected to mechanical agitation in a Waring blender,
which, it was hoped, would remove the ‘shell’of the phages from the outside of the
bacteria, but leave the injected genetic material inside.
T2 phage
32
P
35
S
DNA
labelled
Protein
labelled
Infect bacteria in
labelled media
Isolate phage
Infect bacteria in
unlabelled media
Separate phage ghosts
from bacterial cells
Ghost
Ghosts
unlabelled
Bacteria labelled
with
32
P
Ghosts labelled
with
35
S
Bacteria
unlabelled
From Reece, RJ: Analysis of Genes and Genomes, John Wiley & Sons, 2003. Repro-
duced by permission of the publishers
The bacteria were sedimented by centrifugation, leaving the much lighter phage
‘shells’ in the supernatant. When solid and liquid phases were analysed for
32
P and
35
S, it was found that nearly all the
32
P was associated with the bacterial cells, while
the great majority of the
35
S remained in the supernatant. The conclusion drawn
from these results is that it was the
32
P-labelled DNA which had been injected into
the bacteria, and which was therefore the genetic material.
JWBK011-11 JWBK011-Hogg August 12, 2005 19:21 Char Count= 0
DNA REPLICATION 271
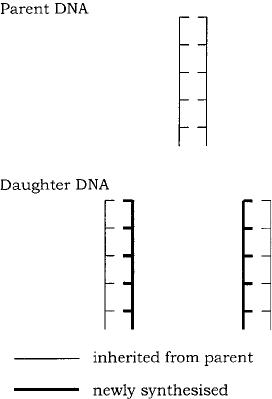
Figure 11.1 DNA replication is semiconservative. Following replication, each new DNA
molecule comprises one strand from the parent DNA and one newly synthesised strand
carbohydrates or any other cellular component was similarly inactivated, the ability
was retained. In spite of this apparently convincing proof, the pro-protein lobby was
not easily persuaded. It was to be several more years before the experimental results of
Alfred Hershey and Martha Chase (Box 11.1) coupled with Watson and Crick’s model
for DNA structure (Figure 2.23) finally cemented the universal acceptance of DNA’s
central role in genetics.
DNA replication
The elucidation of the structure of DNA by Watson and Crick in 1953 stands as one
of the great scientific breakthroughs of the 20th century. An important implication of
their model lay in the complementary nature of the two strands, as they commented
themselves in their famous paper in Nature:
It has not escaped our notice that the specific pairing we have postulated immediately suggests a
possible copying mechanism for the genetic material.
The mechanism they proposed became known as the semiconservative replication of
DNA, so called because each daughter molecule comprises one parental strand and one
newly synthesised strand (Figure 11.1). The parental double helix of DNA unwinds and
each strand acts as a template for the production of a new complementary strand, with
new nucleotides being added according to the rules of base-pairing. In 1957, 4 years
after the publication of Watson and Crick’s model, Matthew Meselson and Franklin
Stahl provided experimental proof of the semiconservative nature of DNA replication
(Box 11.2). The exact way in which replication takes place in procaryotic and eucaryotic
cells differs in some respects, but we shall take as our model replication in E. coli, since
it has been studied so extensively.
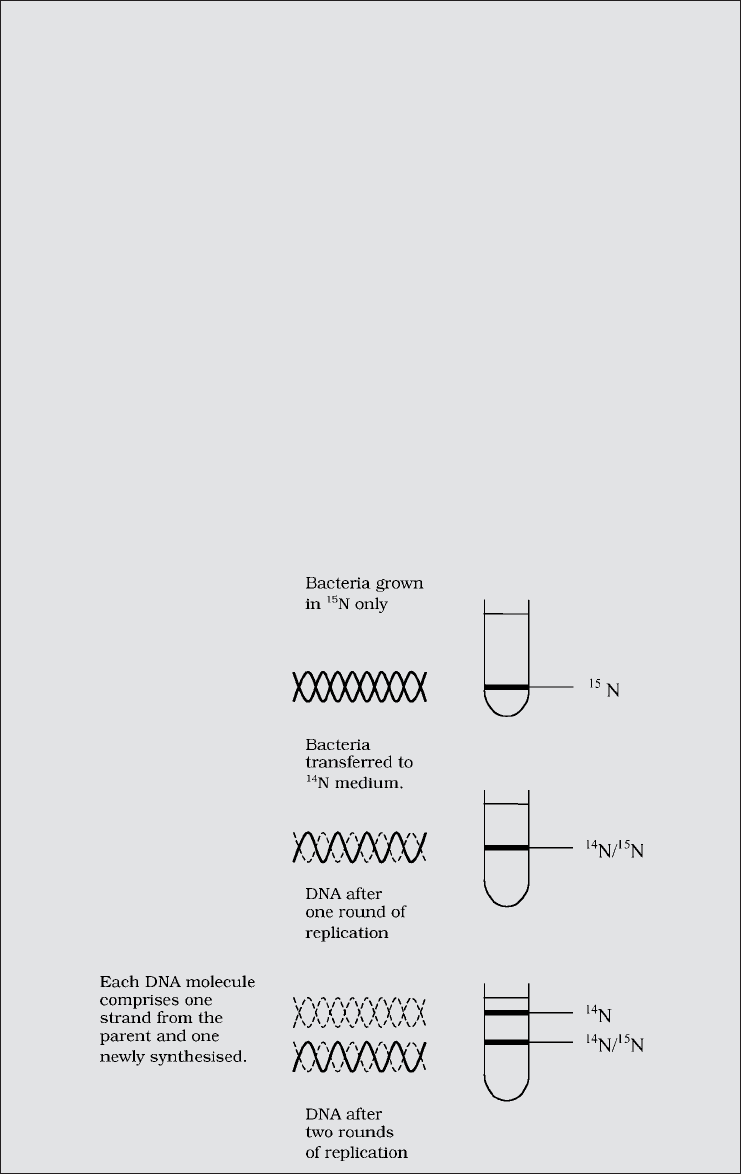
JWBK011-11 JWBK011-Hogg August 12, 2005 19:21 Char Count= 0
Box 11.2 Experimental proof of semiconservative replication
The semiconservative model of DNA replication, as suggested by Watson and Crick,
was not the only one in circulation during the 1950s. Meselson and Stahl provided
experimental evidence which not only supported the semiconservative model, but
also showed the other models to be unworkable. Their elegantly designed exper-
iment used newly developed techniques to differentiate between parental DNA
and newly synthesised material.
First, they grew E. coli in a medium with ammonium salts containing the heavy
isotope
15
N as the only source of nitrogen. This was done for several generations of
growth, so that all the cells contained nitrogen exclusively in the heavy form.
The bacteria were then transferred to a medium containing nitrogen in the nor-
mal,
14
N form. After a single round of replication, DNA was isolated from the culture
and subjected to density gradient centrifugation. This is able to differentiate be-
tween DNA containing the two forms of nitrogen, as
15
N has a greater buoyant
density and therefore settles at a lower position in the tube.
Meselson and Stahl found just a single band of DNA after centrifugation, with a
density intermediate between that of
14
N and
15
N, indicating a hybrid molecule, as
predicted by the semiconservative model. After a second round of E. coli replication
in a medium containing
14
N, two bands of DNA were produced, one hybrid and one
containing exclusively
14
N, exactly as predicted by the semiconservative model, but
inconsistent with other hypotheses.
272

JWBK011-11 JWBK011-Hogg August 12, 2005 19:21 Char Count= 0
DNA REPLICATION 273
DNA replication in procaryotes
You may recall from Chapter 4 that bacteria multiply by a process of binary fission;
before this occurs, each cell must duplicate its genetic information so that each daughter
cell has a copy.
DNA replication involves the action of a number of specialised enzymes:
r
helicases
r
DNA topoisomerases
r
DNA polymerase I
r
DNA polymerase III
r
DNA primase
r
DNA ligase.
DNA replication takes
place at a replication
fork, a Y-shaped struc-
ture formed by the sep-
arating strands. The fork
moves along the DNA as
replication proceeds.
Replication begins at a specific sequence called an ori-
gin of replication. The two strands of DNA are caused to
separate by helicases (Figure 11.2), while single-stranded
DNA binding proteins (SSB) prevents them rejoining.
Opening out part of the double helix causes increased
tension (supercoiling) elsewhere in the molecule, which
is relieved by the enzyme DNA topoiosomerase (some-
times known as DNA gyrase). As the ‘zipper’ moves
along, and more single stranded DNA is exposed, DNA
polymerase III adds new nucleotides to form a comple-
mentary second strand, according to the rules of base-
pairing. DNA polymerases are not capable of initiating
the synthesis of an entirely new strand, but can only
extend an existing one. This is because they require a
free 3
-OH group onto which to attach new nucleotides.
Thus, DNA polymerase III can only work in the 5
–3
direction. A form of RNA polymerase called DNA pri-
mase synthesises a short single strand of RNA, which
can be used as a primer by the DNA polymerase III.
(Figure 11.2).
A primer is a short
sequence of single-
stranded DNA or RNA
required by DNA poly-
merase as a starting
point for chain exten-
sion.
When replication occurs, complementary nucleotides are added to one of the strands
(the leading strand) in a continuous fashion (Figure 11.2). The other strand (the lagging
strand), however, runs in the opposite polarity; so how is a complementary sequence
synthesised here? The answer is that DNA polymerase III allows a little unwinding to
take place and then, starting at the fork, works back over from a new primer, in the 5
–3
direction. Thus, the second strand is synthesised discontinuously, in short bursts, about
1000–2000 nucleotides at a time. These short stretches of DNA are called Okazaki
fragments, after their discoverers.
On the lagging strand, a new RNA primer is needed at the start of every Okazaki
fragment. These short sequences of RNA are later removed by DNA polymerase I, which
JWBK011-11 JWBK011-Hogg August 12, 2005 19:21 Char Count= 0
274 MICROBIAL GENETICS
single-stranded
binding protein
5′
5′
3′
3′
5′
3′
5′
5′
3′
3′ 5′
helicase
helicase
RNA primer
leading strand
DNA polymerase III
Okazaki
fragment
lagging strand
DNA
primase
DNA
polymerase
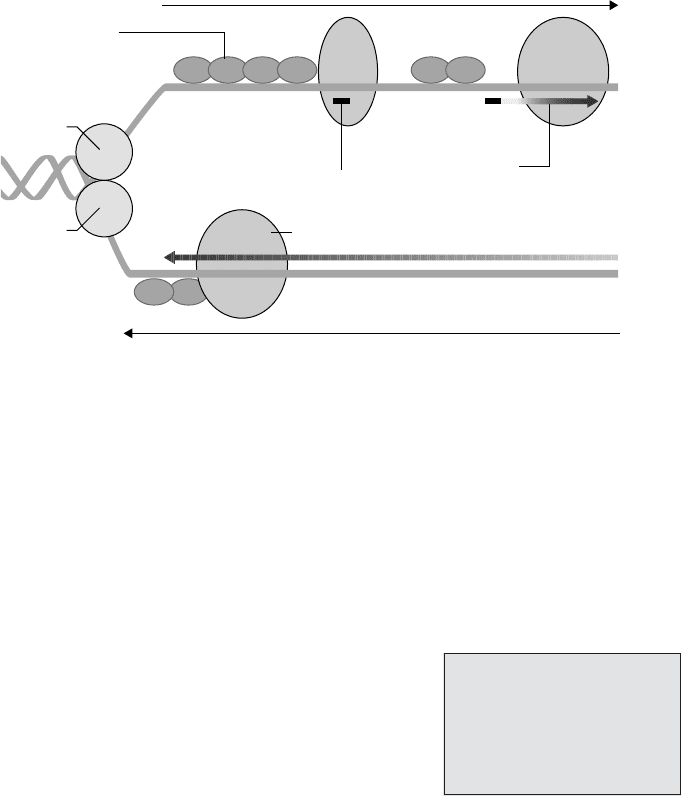
Figure 11.2 DNA replication takes place at a replication fork. Strands of DNA are sep-
arated and unwound by helicase and DNA gyrase, and prevented from rejoining by the
attachment of single-strand binding proteins. Starting from an RNA primer, DNA poly-
merase III adds the complementary nucleotides to form a second strand. On the leading
strand, a single primer is set down (not shown), and replication proceeds uninterrupted in
the same direction as the fork. Because DNA polymerase III only works in the 5
→ 3
di-
rection, on the lagging strand a new primer must be added periodically as the strands open
up. Replication here is thus discontinuous, as a series of Okazaki fragments (see the text).
From Bolsover, SR, Hyams, JS, Jones, S, Shepherd, EA & White, HA: From Genes to Cells,
John Wiley & Sons, 1997. Reproduced by permission of the publishers
DNA ligase repairs
breaks in DNA by re-
establishing phosphodi-
ester bonds in the sugar-
phosphate backbone.
then replaces them with DNA nucleotides. Finally, the
fragments are joined together by the action of DNA
ligase. Replication is bidirectional (Figure 11.3), with
two forks moving in opposite directions; when they
meet, the whole chromosome is copied and replication
is complete
*
.
What happens when replication goes wrong?
It is clearly important that synthesis of a complementary second strand of DNA should
occur with complete accuracy, but occasionally, a non-complementary nucleotide is
inserted; this may happen as frequently as once in every 10 000 nucleotides. Cells,
however, are able to have a second attempt to incorporate the correct base because of
the proofreading activity of the enzymes DNA polymerase I and III. These are able to
*
This rather long-winded description hardly does justice to a process that incorporates new nucleotides at a
rate of around 1000 per second!

JWBK011-11 JWBK011-Hogg August 12, 2005 19:21 Char Count= 0
WHAT EXACTLY DO GENES DO? 275
Direction of replication fork
Replication bubble
Figure 11.3 DNA replication is bidirectional. Two replication forks form simultaneously,
moving away from each other and developing a replication bubble
cut out the ‘wrong’ nucleotide and replace it with the correct one. As a result of this
monitoring, mistakes are very rare; they are thought to occur at a frequency of around
one in every billion (10
9
) nucleotides copied. Mistakes that do slip through the net result
in mutations, which are discussed later in this chapter.
DNA replication in eucaryotes
In eucaryotic organisms, the DNA is linear, not circular, so the process of replication
differs in some respects. Genome sizes are generally much bigger in eucaryotes, and
replication much slower, so numerous replication forks are active simultaneously on
each chromosome. Replication takes place bidirectionally, creating numerous replica-
tion bubbles (Figure 11.4) which merge with one another until the whole chromosome
has been covered.
What exactly do genes do?
At the start of the 20th century Archibald Garrod had proposed that inherited disorders
such as alkaptonuria may be due to a defect in certain key metabolic enzymes, thus
offering for the first time an explanation of how genetic information is expressed. His
ideas were not really developed however, until the work of George Beadle and Edward
Tatum in the 1940s, whose experiments with the bread mould Neurospora led to the
formulation of the one gene, one enzyme hypothesis. Although now acknowledged to be
Figure 11.4 DNA replication in eucaryotes. Many replication bubbles develop simultane-
ously; they extend towards each other and eventually merge. The arrows denote the direction
of the replication forks

JWBK011-11 JWBK011-Hogg August 12, 2005 19:21 Char Count= 0
276 MICROBIAL GENETICS
Box 11.3 One gene, one enzyme: not quite true
Beadle and Tatum proposed that each gene was responsible for the production of
a specific enzyme. However all proteins, not just enzymes, are encoded by DNA,
and furthermore, some have a quaternary structure (see Chapter 2) with different
polypeptide subunits being encoded by different genes. The hypothesis was there-
fore modified to one gene, one polypeptide. Later still, it emerged that even this
is not always the case, as some genes do not encode proteins at all, but forms of
RNA.
somewhat over-generalised (see Box 11.3), this model proved useful in the years when
the molecular basis of gene action was being elucidated.
Having established that genes are made of DNA, and having a model for the structure
of DNA that explained how it was able to copy itself, the way was open in the 1950s
for scientists to work out the mechanism by which the information encoded in a DNA
sequence was converted into a specific protein.
How does a gene direct the synthesis of a protein?
You may recall from Chapter 2 that both DNA and proteins are polymers whose ‘build-
ing blocks’ (nucleotides and amino acids respectively) can be put together in an almost
infinite number of sequences. The sequence of amino acids making up the primary
structure of a protein is determined by the sequence of nucleotides in the particular
gene responsible for its production. It does this not directly, but through an intermedi-
ary molecule, now known to be a form of RNA called messenger RNA (mRNA). It is this
intermediary that carries out the crucial task of passing the information encoded in the
DNA sequence to the site of protein synthesis. This unidirectional flow of information
can be summarised:
DNA mRNA
DNA replication
Protein
and is often referred to as the Central Dogma of biology, because of its applicability to
all forms of life. Proposed by Crick in the late 1950s, this is still accepted as being a
true model of the basic events in protein synthesis. Sometimes the message encoded in
DNA is transcribed into either ribosomal RNA (rRNA) or transfer RNA (tRNA); these
types of RNA are not translated into proteins but represent end-products in themselves.
(In the last chapter, we saw that retroviruses have proved an exception to one part of
the Central Dogma, since they possess enzymes capable of forming DNA from an RNA
template.)
The conversion of information encoded as DNA into the synthesis of a polypeptide
chain occurs in two distinct phases (Figure 11.5); first the ‘message’ encoded in the
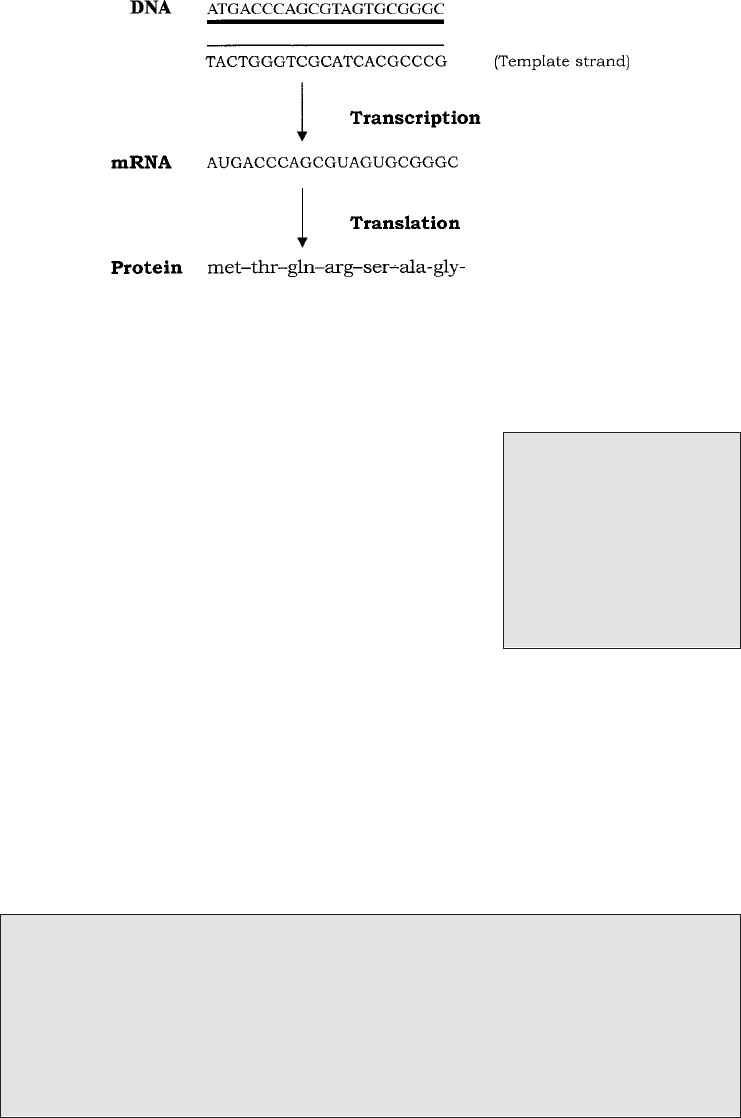
JWBK011-11 JWBK011-Hogg August 12, 2005 19:21 Char Count= 0
WHAT EXACTLY DO GENES DO? 277
Figure 11.5 Transcription and translation. The sequence on one strand of DNA is tran-
scribed as a molecule of mRNA (with uracil replacing thymine). The triplet code on the
mRNA is then translated at ribosomes as a series of amino acids
Messenger RNA (mRNA)
is formed from a DNA
template (transcription)
and carries its encoded
message to the ribo-
some, where it directs
synthesis of a polypep-
tide (translation).
DNA sequence of a gene is converted to mRNA by tran-
scription, then this directs the assembly of a specific se-
quence of amino acids during translation. We shall dis-
cuss how this happens shortly, but first we need to con-
sider the question: how does the sequence of nucleotides
in a gene serve as an instruction in the synthesis of pro-
teins?
The genetic code
Messenger RNA carries information (copied from a DNA template) in the form of a
genetic code that directs the synthesis of a particular protein. The nature of this code
was worked out in the early 1960s by Marshall Nirenberg, Har Gobind Khorana and
others (Box 11.4). The message encoded in a gene takes the form of a series of triplets
(codons). Of the 64 possible three-letter combinations of A, C, G and U, 61 correspond
to specific amino acids, while the remaining three act as ‘stop’ messages, indicating that
reading of the message should cease at that point (Figure 11.6). It is essential that the
reading of the message starts at the correct place, otherwise the reading frame (groups of
Box 11.4 The genetic code is (almost) universal
The genetic code was first worked out using E. coli, but soon found to apply to other
organisms too. It seemed reasonable to assume that the code was universal, i.e.
applicable to all life forms. It has been shown, however, that certain genes such
as those found in the mitochondria of some eucaryotes employ a slight variation of
the code. Mitochondria have their own transcription enzymes, ribosomes and tRNAs
and so are able to use a modified system.
