Hogg S. Essential microbiology
Подождите немного. Документ загружается.


P1: FCG/SPH P2: FCG/SPH QC: FCG/SPH T1: SPH
JWBK011-14 JWBK011-Hogg August 12, 2005 16:32 Char Count= 0
358 ANTIMICROBIAL AGENTS
r
if administered orally, it must not be inactivated by the acid environment of the
stomach, and must be capable of being absorbed by the small intestine.
Antibiotics should not
produce a hypersensitiv-
ity (allergic) reaction in
the host. This is caused by
an extreme response by
the host immune system,
and is not the same as
toxicity.
r
an antibiotic should not have any significant effect on
the resident microflora of the host.
r
it should not be easy for the target pathogen to estab-
lish resistance against an antibiotic.
r
side-effects such as allergic reactions should be
minimal.
r
it should be sufficiently stable to have a good shelf
life, without special storage considerations.
How do antibiotics work?
All antibiotics have the common property of interfering in some way with a normal,
critical function of the target bacterial cell. The most commonly used antibiotics exert
their effect by one of the following methods:
1 Inhibition of cell wall synthesis (group I)
2 Disruption of cell membranes (group II)
3 Interference with protein synthesis (group III)
4 Interference with nucleic acid synthesis (group IV)
Table 14.2 lists examples of each group. Those antibi-
otics belonging to groups I and III are better able to dis-
criminate between procaryotic and eucaryotic cells, and
consequently show more selective toxicity and a higher
therapeutic index.
Group I: Inhibitors of cell wall synthesis
The therapeutic index
provides a measure of
the selective toxicity
of a chemotherapeu-
tic agent. It is the ra-
tio between the con-
centration at which the
substance causes harm
to its host (toxic dose)
and that at which it
is required to be clin-
ically effective (thera-
peutic dose). It is there-
fore desirable for an
antibiotic to have a high
therapeutic index.
The main group which work in this way are the β-lactam
antibiotics, so-called because they contain a β-lactam
ring in their structure. Included among this group are the
penicillins and the cephalosporins. See also Box 14.4.
You may recall from our discussion of bacterial cell
wall structure in Chapter 3 that an important factor in
the strengthening of the peptidoglycan component of
the bacterial cell walls is the cross-linking of chains by
transpeptidation. It is this process which is acted on by
the β-lactams; they bind irreversibly to the transpep-
tidase enzyme, forming covalent bonds with a serine

P1: FCG/SPH P2: FCG/SPH QC: FCG/SPH T1: SPH
JWBK011-14 JWBK011-Hogg August 12, 2005 16:32 Char Count= 0
ANTIBIOTICS 359
Table 14.2 Some commonly used antibiotic classes
Inhibitors of cell wall synthesis Penicillins, cephalosporins
Disrupters of cell membranes Polymixins, polyenes
Inhibitors of protein synthesis Streptomycin, tetracyclines
Inhibitors of nucleic acid synthesis Rifamycins
residue within the enzyme’s active site. The cell wall continues to form, but becomes
progressively weaker as more new, unlinked, peptidoglycan is set down. Since bacteria
are generally to be found in a hypotonic environment, as the wall weakens, water enters
the cell, leading to swelling and then lysis.
Penicillins The first β-lactam antibiotic to be discovered was benzylpenicillin, or
penicillin-G, whose action is restricted to Gram-positive bacteria, because it is unable
to penetrate the Gram-negative cell wall. It is effective against Gram-positive bacteria
when administered intramuscularly, but cannot be taken by mouth because it is broken
down in the acid conditions of the stomach. Another naturally occurring penicillin,
penicillin-V, represented an advance inasmuch as it is less acid-labile and can therefore
Semisynthetic penicillins
are based on the core
structure of the naturally
occurring molecule, with
the addition of chem-
ically synthesized side
chains.
be taken orally. All the penicillins are based on a core
structure or nucleus called 6-amino-penicillanic acid
(Figure 14.2); extensive research has led to the develop-
ment of many variants of this, the so-called semisynthetic
penicillins. These have attached to their nucleus novel
side chains not encountered in nature, and have over-
come some of the problems inherent in naturally occur-
ring penicillins such as instability and narrow specificity
(Figure 14.3).
Ampicillin is a semi-synthetic penicillin that has a broader specificity (Box 14.5) than
Penicillin G; it is appreciably more effective against Gram-negative bacteria such as
Salmonella and E. coli, its hydrophobic nature making it better able to penetrate their
outer membrane. It has the additional benefit of being acid-stable and can therefore be
taken orally.
Another drawback to natural penicillins is that they are susceptible to naturally
occurring bacterial β-lactamases (also called penicillinases), which breaks a bond in the
β-lactam core of the penicillin molecule (Figure 14.4). Sometimes, β-lactam antibiotics
Box 14.4 β-lactam antibiotics have a second mode of action
The β-lactams also act by preventing the natural regulation of enzymes called au-
tolysins. These enzymes function by breaking down peptidoglycan in a controlled
fashion, causing breaks to allow for the addition of new peptidoglycan as the cell
grows, and are normally regulated by naturally occurring inhibitors. The β-lactams
neutralise the activity of these inhibitors, leading to further breakdown of the cell
wall.
P1: FCG/SPH P2: FCG/SPH QC: FCG/SPH T1: SPH
JWBK011-14 JWBK011-Hogg August 12, 2005 16:32 Char Count= 0
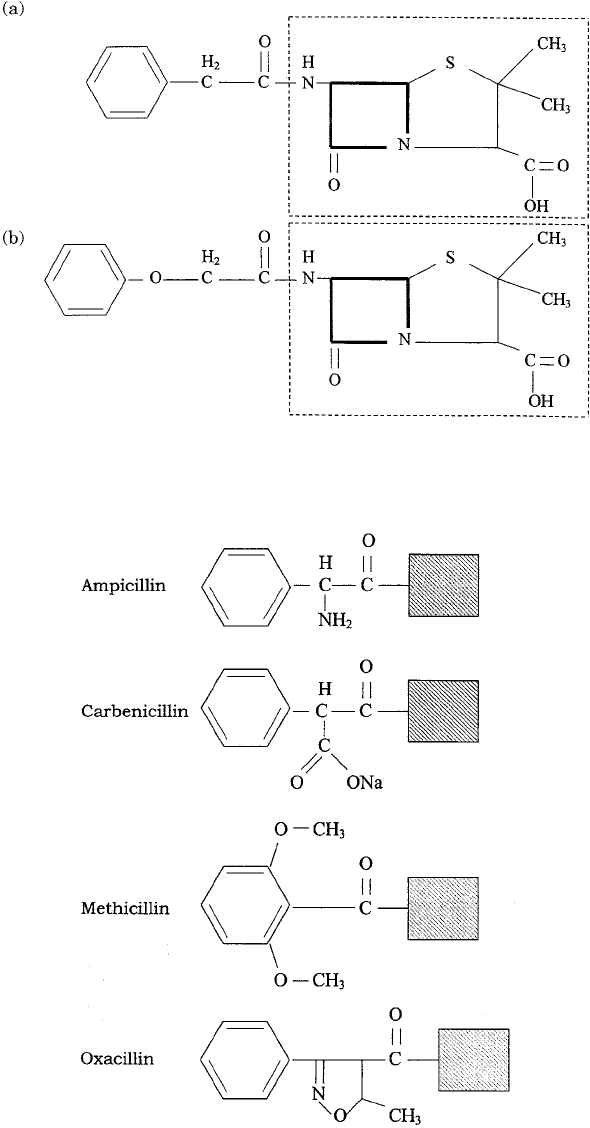
Figure 14.2 Naturally occurring penicillins. (a) Penicillin G (benzylpenicillin); (b) penicillin
V. The dotted outline covers the 6-aminopenicillanic acid nucleus present in all variants of
penicillin. The heavy outline denotes the β-lactam ring
Figure 14.3 Some important semisynthetic penicillins. The shaded square represents the
6-aminopenicillanic acid nucleus common to all penicillins (see Figure 14.2)
360

P1: FCG/SPH P2: FCG/SPH QC: FCG/SPH T1: SPH
JWBK011-14 JWBK011-Hogg August 12, 2005 16:32 Char Count= 0
ANTIBIOTICS 361
Box 14.5 Broad spectrum or narrow spectrum?
Certain antibiotics, due to the mechanism of their action, are only effective against
a few different pathogens, while others can be used successfully against many
different kinds. They are said to have, respectively, a narrow spectrum and a broad
spectrum of activity.
On the face of it, all things being equal, you would expect your doctor to choose
the antibiotic with the broadest possible spectrum of activity, but this isn’t always
the wisest option. When the cause of an infection isn’t known, it makes sense to
hedge one’s bets and prescribe a broad-spectrum antibiotic (‘whatever it is, this
should sort it!’), but this policy is not without its dangers. The drug is likely to kill off
many of the host’s own resident microflora, which can lead to a superinfection,
and the development of antibiotic-resistant strains is also made more likely. If the
identity of the pathogen is suspected, an appropriate narrow-spectrum drug is to be
preferred.
are taken in combination with a β-lactamase inhibitor such as clavulanic acid. This
binds to the β-lactamase with a high affinity, preventing it from acting on the antibiotic.
Some semisynthetic penicillins such as methicillin and oxacillin are resistant to attack by
the β-lactamases that can render certain bacteria resistant to their naturally occurring
forms.
An anaphylactic shock is
an extreme form of hy-
persensitivity reaction.
Penicillin is not an appropriate treatment for the es-
timated 1–5 per cent of adults who show an allergic re-
action to it; in extreme cases, death from anaphylactic
shock can result.
Cephalosporins The cephalosporins, like the penicillins, have a structure based on a
β-lactam ring (Figure 14.5). They also exert their effect on transpeptidases, but gen-
erally have a broader specificity and are more resistant to the action of β-lactamases.
Ceftriaxone, for example, is now used in the treatment of gonorrhoeal infections, caused
by penicillin-resistant strains of Neisseria gonorrhoeae. In addition, patients who are
allergic to penicillin are often treated with cephalosporins. Cephalosporins were first
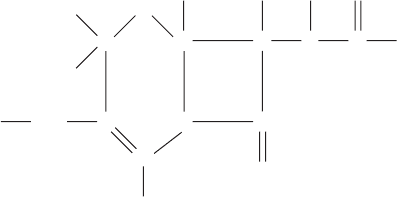
Figure 14.4 Action of β-lactamase on penicillin. A number of bacteria, especially staphylo-
cocci, possess the enzyme β-lactamase (penicillinase), which inactivates penicillin by cleavage
of the β-lactam ring at the point marked by the arrow
P1: FCG/SPH P2: FCG/SPH QC: FCG/SPH T1: SPH
JWBK011-14 JWBK011-Hogg August 12, 2005 16:32 Char Count= 0
362 ANTIMICROBIAL AGENTS
CCCNCR
1
H HO
H
CN
C
O
S
C
COOH
H
H
CH
2
R
2
Figure 14.5 Cepahalosporins are based on a nucleus of 7-amino-cephalosoranic acid,
which, like the penicillins, features a β-lactam ring (shown as a square). Note that each
molecule has two variable side chains
isolated in the late 1940s from a marine fungus called Cephalosporium acremonium,
and came into general use in the 1960s. So-called second, third and fourth genera-
tion cephalosporins have been developed to widen the spectrum of activity to include
many Gram-negative organisms, and to keep one step ahead of pathogens developing
resistance to earlier versions.
Both penicillins and cephalosporins are also used prophylactically, that is, in the
prevention of infections, prior to surgery in particularly vulnerable patients.
Other antibiotics that affect the cell wall Carbapenems are β-lactam antibiotics pro-
duced naturally by a species of Streptomyces. A semisynthetic form, imipenem, is active
against a wide range of Gram-positive and -negative bacteria, and is used when resis-
tance to other β-lactams has developed.
Bacitracin and vancomycin are two other antibiotics that exert their effect on the cell
wall, but by a different mechanism. Bacitracin is derived from species of Bacillus and
acts on bactoprenol pyrophosphate, the lipid carrier molecule responsible for transport-
ing units of peptidoglycan across the cell membrane to their site of incorporation into
the cell wall (see Chapter 3). Its use is restricted to topical (surface) application, since
its use internally can cause kidney damage. Vancomycin is a highly toxic antibiotic with
a narrow spectrum of use against Gram-positive organisms such as streptococci and
staphylococci. It is particularly important in its use against infections caused by organ-
isms resistant to methicillin and the cephalosporins, such as methicillin-resistant Staphy-
lococcus aureus (MRSA) (see Resistance to Antibiotics below). It is not absorbed from
the gastrointestinal tract and is therefore most commonly administered intravenously.
Group II: Antibiotics that disrupt cell membranes
Polymixins are a class of antibiotic that act by disrupting the phospholipids of the
cytoplasmic membrane and causing leakage of cell contents. Produced naturally by a
species of Bacillus, polymixins are effective against pseudomonad infections of wounds
and burns, often in combination with bacitracin and neomycin (an inhibitor of protein
synthesis; see below). Their toxicity makes them unsuitable for internal use. Polyene
antibiotics such as amphotericin and nystatin are antifungal agents that act on the
P1: FCG/SPH P2: FCG/SPH QC: FCG/SPH T1: SPH
JWBK011-14 JWBK011-Hogg August 12, 2005 16:32 Char Count= 0
ANTIBIOTICS 363
sterol components of membranes; they are discussed more fully towards the end of this
chapter.
Group III: Inhibitors of protein synthesis
Antibiotics that act by affecting protein synthesis generally have a relatively broad spec-
trum of action. As we saw in our historical review earlier in this chapter, streptomycin
was the first antibiotic that was shown to be effective against Gram-negative organisms.
Its discovery in 1943 was particularly welcome since such organisms were unaffected
by penicillin or sulphonamides. It proved to be particularly useful in the treatment of
tuberculosis, the causative agent of which, Mycobacterium tuberculosis, is protected
against the effects of penicillin by the waxy layer of mycolic acids in its cell wall.
Streptomycin belongs to a group of antibiotics called aminoglycosides, which act
by binding to the 30S subunit of the bacterial ribosome, preventing attachment of
the 50S subunit to the initiation complex (Figure 14.6). They can thus discriminate
between procaryotic (70S) and eucaryotic (80S) ribosomes, and consequently have a
relatively high therapeutic index (although not as high as cell wall inhibitors). Other
members of this group are gentamicin, kanamycin and neomycin. Like some other
‘wonder drugs’, streptomycin has proved to have undesirable side-effects; these have
led to it being replaced in most applications by safer alternatives. In addition, bacterial
resistance to streptomycin is widespread, further diminishing its usefulness. Use of the
5’
PE A
3’
50S
subunit
30S
subunit
tRNA
mRNA
a)
b)
c)
d)
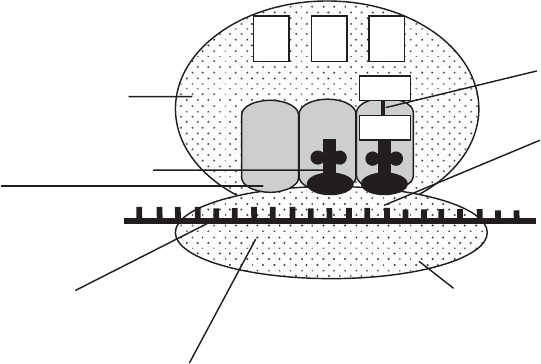
Figure 14.6 Inhibitors of protein synthesis. (a) By binding to the 30S subunit of the bac-
terial ribosome, aminoglycosides block the attachment of the 50S subunit. This prevents
completion of the initiation complex, thus protein synthesis is inhibited. (b) Tetracyclines
distort the shape of the 30S subunit, preventing the attachment of the appropriate amino-
acyl tRNA. (c) Chloramphenicol inhibits peptidyltransferase and prevents formation of new
peptide bonds. (d) Macrolides such as erythromycin bind to the 50S subunit, preventing
elongation of the growing peptide chain
P1: FCG/SPH P2: FCG/SPH QC: FCG/SPH T1: SPH
JWBK011-14 JWBK011-Hogg August 12, 2005 16:32 Char Count= 0
364 ANTIMICROBIAL AGENTS
aminoglycosides as a group has diminished since the development of later generation
cephalosporins and the tetracyclines.
Tetracyclines also work by binding to the 30S ribosomal subunit, preventing the
attachment of aminoacyl tRNA, and therefore extension of the peptide chain (Fig-
ure 14.6). They are yet another group of antibiotics produced by Streptomyces spp.
Both natural and semisynthetic tetracyclines are easily absorbed from the intestine,
allowing them to be taken orally. Coupled with their broad specificity (the broadest of
any antibiotic), this led to inappropriately widespread use in the years following their
discovery, sometimes resulting in complications caused by the destruction of the normal
resident microflora. Tetracyclines are still used for a number of applications, notably to
treat a variety of sexually transmitted diseases.
Two important antibiotics which act on the larger, 50S, subunit of the procaryotic
ribosome are erythromycin and chloramphenicol. Both combine with the subunit in
such a way as to prevent the assembly of amino acids into a chain (Figure 14.6). Chlo-
ramphenicol was the first antibiotic to be discovered with a broad spectrum of activity;
it also derives originally from Streptomyces spp., but is nowadays produced synthet-
ically. Its use has become severely restricted since it was shown to have some serious
side-effects, notably on the bone marrow, but it remains the agent of choice for the
treatment of typhoid fever.
Erythromycin is the best known of the macrolide group of antibiotics. Unlike chlo-
ramphenicol, it has a large hydrophobic molecule and is unable to gain access to most
Gram-negative bacteria, thus restricting its spectrum of activity. Erythromycin can be
taken orally and has a similar spectrum of activity to penicillin G; it is often used instead
of penicillin in the treatment of staphylococcal and streptococcal infections in children.
It is particularly appropriate for this application as it is one of the least toxic of all
commonly used antibiotics.
Group IV: Inhibitors of nucleic acid synthesis
Rifampin belongs to a group of agents called rifamycins. It acts by inhibiting the enzyme
RNA polymerase, thereby preventing the production of mRNA. Rifampin is used against
the mycobacteria that cause tuberculosis, an application for which its ability to penetrate
tissues makes it well suited. Unlike most antibiotics, rifampin interacts with other drugs,
often reducing or nullifying their effect. When used in high doses, it has the unusual side-
effect of turning secretions such as tears, sweat and saliva, as well as the skin, an orange-
red colour. As we have already seen, the quinolone group of synthetic antimicrobial
drugs act by disrupting DNA replication.
Resistance to antibiotics
The global increase in resistance to antimicrobial drugs, including the emergence of bacterial
strains that are resistant to all available antibacterial agents, has created a public health problem
of potentially crisis proportions.
American Medical Association, 1995

P1: FCG/SPH P2: FCG/SPH QC: FCG/SPH T1: SPH
JWBK011-14 JWBK011-Hogg August 12, 2005 16:32 Char Count= 0
RESISTANCE TO ANTIBIOTICS 365
Box 14.6 Where did antibiotic resistance come from?
Have genes responsible for antibiotic resistance always existed in nature, or have
they arisen since the development and widespread use of antibiotics? The answer,
almost certainly, is the former. A sample of an E. coli strain, freeze-dried in 1946, was
revived many years later, and found to have plasmid-encoded genes for resistance
to streptomycin and tetracycline, neither of which were in clinical use until some
years after the culture was preserved. It seems likely that bacteria possessed these
genes to protect against naturally occurring antibiotics, an idea supported by the
fact that R-plasmids have been found in non-pathogenic soil bacteria. Also, resis-
tance to a number of antibiotics has been demonstrated in soil and water bacteria
from sources sufficiently remote to be free from anthropogenic influence.
As we have already suggested, the impact of certain antibiotics can be greatly reduced
due to the development of resistance by target pathogens (Box 14.6). This represents the
greatest single challenge facing us in the fight against infectious diseases at the start of
the 21st century. Fleming himself foresaw that the usefulness of penicillin might become
limited if resistant forms of pathogens arose.
Nosocomial infections
are ones that are ac-
quired in hospitals or sim-
ilar locations. Some 5--10
percent of hospital pa-
tients acquire such an in-
fection during their stay.
This may prove to be
fatal, especially among
the elderly and immuno-
compromised. As well as
the human cost, such in-
fections extend the av-
erage time spent in hos-
pital and therefore add
greatly to the costs of
treatment.
Not long after penicillin was put into general use,
strains of Staphylococcus aureus were found which
did not respond to treatment, and by 1950 penicillin-
resistant S. aureus was a common cause of infections in
hospitals. A decade later, a semi-synthetic form of peni-
cillin, methicillin, was introduced; this was not affected
by the β-lactamase enzymes that inactivated Penicillin
G, and was used to treat resistant forms. Within years,
however, came the first reports of strains of S. aureus
that did not respond to methicillin. The incidence of
methicillin-resistant S. aureus (MRSA) has increased
greatly since, and it represents the major source of noso-
comial infections. In 1980, synthetic fluoroquinolones
were introduced to counter the threat of MRSA, but
within a year 80 per cent of isolated strains had devel-
oped resistance to these too. Vancomycin is regarded as a
last-resort treatment for MRSA, for a number of reasons;
it has a number of serious side-effects, its widespread use
would encourage resistance against it, and it is extremely
expensive.
A case of vancomycin-resistant Staphylococcus aureus (VRSA) emerged in Japan in
1996; a few months later it had reached the USA. This represents a serious threat; some
of these strains respond to treatment with a cocktail of antibiotics, but already people
have died from untreatable VRSA infections. In 2003, a strain of VRSA was shown
to have obtained its vancomycin resistance by cross-species transfer from a strain of
Enterococcus faecalis.
P1: FCG/SPH P2: FCG/SPH QC: FCG/SPH T1: SPH
JWBK011-14 JWBK011-Hogg August 12, 2005 16:32 Char Count= 0
366 ANTIMICROBIAL AGENTS
How does antibiotic resistance work?
We saw earlier in this chapter how antibiotics exert their effects in a variety of ways, so
it should come as no surprise that there is no single mechanism of resistance. Resistance
may be natural, that is, intrinsic to the microorganism in question, or it may be acquired.
Some bacteria are able to resist antibiotic action by denying it entry to the cell;
penicillin G for example is unable to penetrate the Gram negative cell wall. Others
can pump the antibiotic back out of the cell before it has had a chance to act, by
means of enzymes called translocases; this is fairly non-specific, leading to multiple drug
resistance. Other bacteria are naturally resistant to a particular antibiotic because they
lack the target for its action, for example, mycoplasma do not possess peptidoglycan,
the target for penicillin’s action.
To avoid the action of an antibiotic, bacteria may be able to use or develop alter-
native biochemical pathways, so that its effect is cancelled out. Many pathogens can
secrete enzymes that modify or degrade antibiotics, causing them to lose their activ-
ity; we have already seen that penicillins can be inactivated by enzymatic cleavage of
their β-lactam ring. Similarly, chloramphenicol can be acetylated, while members of the
aminoglycoside family can be acetylated, adenylated or phosphorylated, all leading to
loss of antimicrobial activity.
Mutations may occur which modify bacterial proteins in such a way that they are
not affected by antimicrobial agents. You will recall that streptomycin normally acts by
binding to part of the 30S subunit on the bacterial ribosome; the actual binding site is
a protein called S12. Mutant forms of the S12 gene can lead to a product which still
functions in protein synthesis, but loses its ability to bind to streptomycin. Similarly,
mutations in transpeptidase genes in staphylococci means they do not bind to penicillin
any more, so cross-linking of the cell wall is not inhibited.
How does resistance arise?
Occasionally, mutations occur spontaneously in bacteria, which render them resistant
to one antibiotic or another. Usually the mutation leads to a change in a receptor or
binding site such as those just described, rendering the antibiotic ineffective. The changes
are usually brought about by point mutations (see Chapter 11) occurring at very low
frequency on chromosomal DNA. Bacteria can, however, become resistant much more
rapidly by acquiring the mutant resistance-causing gene from another bacterium. This
is called transmissable antibiotic resistance; it occurs mainly as a result of bacterial
conjugation, and is the cause of most of the resistance problems we presently face.
Transmissable resistance was first reported in Japan in the late 1950s, when multi-drug
resistance in Shigella was shown to have been acquired by conjugation with resistant
E. coli in a patient’s large intestine. E. coli is known to transfer R (resistance) plasmids
to several other gut bacteria including Klebsiella, Salmonella and Enterobacter, as well
as Shigella. Whereas chromosomal mutations usually result in a modification to the
drug’s binding site, genes carried on plasmids code for enzymes which inactivate it, (e.g.
β-lactamases) or lead to its exclusion from the cell (translocases).
There is a strong link between the use of a particular antibiotic in a locality and the
incidence of resistant bacterial strains. This is because of selective pressure favouring
P1: FCG/SPH P2: FCG/SPH QC: FCG/SPH T1: SPH
JWBK011-14 JWBK011-Hogg August 12, 2005 16:32 Char Count= 0
ANTIBIOTIC SUSCEPTIBILITY TESTING 367
the resistant forms of a bacterium. Fortunately this can, at least in part, be reversed,
as several studies have shown, where a more restricted use of certain antibiotics over
several years was followed by a reduction in the incidence of resistant bacterial forms.
Antibiotic susceptibility testing
In order to determine the most appropriate antimicrobial agent to use against an infec-
tion, it is necessary to determine the susceptibility of the pathogen. There are several
ways of doing this, but here we describe the two most commonly employed techniques.
The tube dilution assay determines the minimum inhibitory concentration (MIC) of
the antibiotic, that is, the lowest concentration at which it prevents growth of a given
organism. A series of tubes containing increasingly dilute preparations of the antibiotic
is introduced into a broth with a standard number of test organisms and incubated. The
lowest concentration in the series to show no microbial growth is the MIC (Figure 14.7).
9.9ml
growth
medium
Antimicrobial Agent
0.1ml 0.1ml 0.1ml0.1ml0.1ml0.1ml0.1ml0.1ml
MIC
No growth Growth
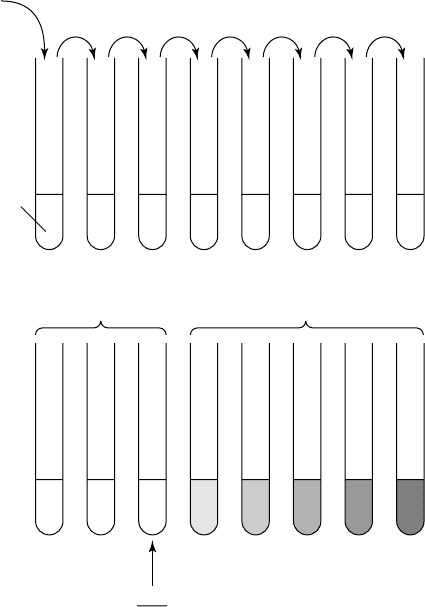
Figure 14.7 Minimal inhibitory concentration (MIC). The test organism is incubated with
serially diluted antibiotic. The lowest concentration capable of preventing microbial growth
is the MIC
