Scheffler Immo E. Mitochondria
Подождите немного. Документ загружается.

THE ELECTRON TRANSPORT CHAIN 211
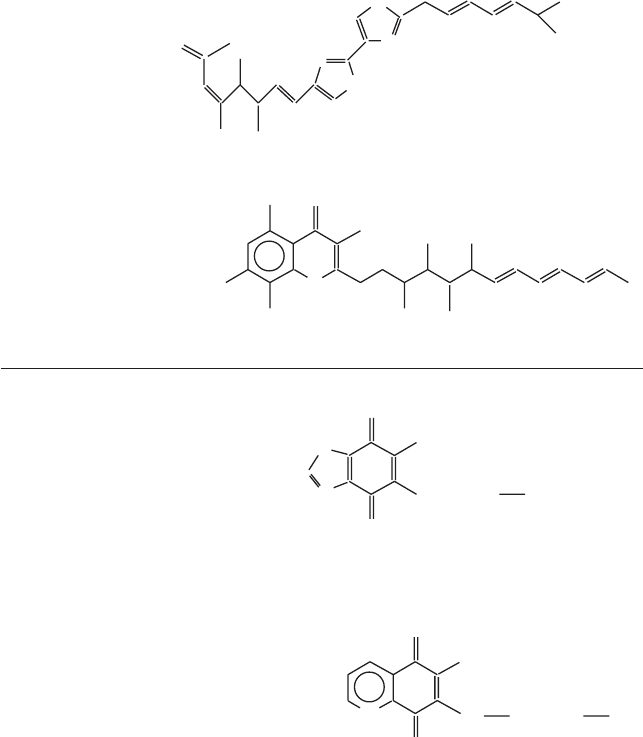
Figure 5.17 Different subclasses of inhibitors frequently used in the study of complex
III. The fargets for the different drugs in groups A, B, and C are discussed in the text
(see also Figure 5.14 )
O
NH
2
OCH
3
OCH
3
S
N
S
N
Myxothiazol
O
O
OH
H
3
CO
H
3
CO
CH
3 CH
3
OCH
3
H
3
CO
Stigmatellin A
undecylhydroxydioxobenzothiazole
UHDBT
O
O
OH
S
N
(CH
2
)
1
CH
3
heptadecylmercaptohydroxyquinoline
quinone
HMHQQ
O
O
N
OH
S (CH
2
)
1
CH
3
A
B
mark a major milestone in the history of electron transport and respiration.
As always, these papers settled some older arguments and answered a number
of questions quite defi nitively, but a wealth of new questions were raised to
understand at the atomic level how electron transport is intimately coupled
to proton pumping. Only high - resolution computer images with the added
capability of rotating the image and zooming in and out of the structure can
give a true sense of this achievement and its powerful infl uence on future
thinking about these fundamental processes in bioenergetics. As one review
212 MITOCHONDRIAL ELECTRON TRANSFER
summarized the situation at the time: “ The monster is subdued, but far from
tamed ” (159) . In the meantime, more structures have been solved, and existing
structures have been refi ned at higher resolution. The interested reader can
fi nd a wealth of information and links at a website dedicated to cytochrome
oxidase ( http://www - bioc.rice.edu/ ∼ graham/CcO.html ).
The functionally homologous but simpler bacterial structure also became
available in 1995 (160) , by itself a major achievement. A remarkable conserva-
tion of structural features helps to emphasize those aspects most basic and
essential for function. Certain differences will emphasize the requirement for
regulatory mechanisms in higher organisms that may not exist or may be dif-
ferent in bacteria.
A discussion must start with the composition of the complex. The mamma-
lian complex contains 13 subunits, the yeast complex is composed of 9 subunits,
and one may expect to see further variations in the total number of subunits
in different organisms. In all organisms the three largest subunits (I, II, III)
are encoded by the mitochondrial genome and synthesized in the matrix. For
the remaining subunits the nomenclature can be confusing to the uninitiated:
In the mammalian complex we have subunits IV, Va, Vb, VIa, VIb, VIc, VIIa,
VIIb, VIIc, and VIII; in yeast the subunits are IV, V, VI, VII, VIIa, and VIII.
Antimycin A
HO
HO
HO N
OH
CH
3
CH
3
CH
3
CH
3
O
O
OH
OH
(CH
2
)
8
CH
3
O
N
Funiculosin
Heptylhydroxyquinoline-N-oxide
CH
3
CH
3
NH
CHO
OH
C
O
NH
O
CH
3
O
O
(CH
2
)
5
CH
3
O
CH(CH
3
)
2
C
O
CH
2
C
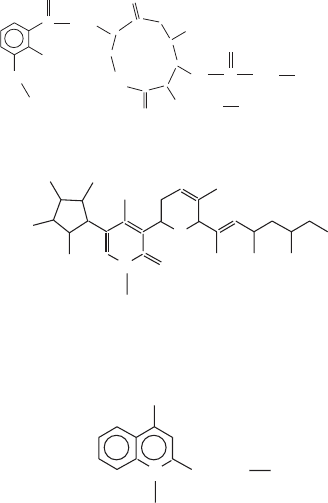
Figure 5.17 (Continued)
THE ELECTRON TRANSPORT CHAIN 213
And while a biochemist may refer to subunit VIIa in yeast, a geneticist might
refer to it as the COX9 gene product.
A large number of biochemical and elegant spectroscopic analyses had
established some time ago that the complex contained two hemes (aa
3
) and
two copper centers. In fact, the cytochromes a and a
3
were among the fi rst
described by D. Keilin in 1925. The X - ray structure has probably been most
defi nitive in defi ning two additional metal centers containing Mg and Zn.
Studies on the simpler bacterial cytochrome c oxidases and comparisons of
homologous peptides have given clues about the association of these impor-
tant metal centers with specifi c peptides which are now totally confi rmed for
yeast and mammals. Subunit I binds the heme a and heme a
3
prosthetic groups
and also forms the Cu
B
redox center. Subunit II binds the Cu
A
center. Subunits
I – III form the core of the enzyme. While subunits I and II can carry out the
redox reactions and proton pumping, they become irreversibly inactivated
during one cycle, and subunit III has been proposed to maintain the structural
integrity of the complex (see reference 156 for a recent review and listing of
many references). The other subunits may perform regulatory functions or
play a role in insulation, stabilization, or assembly. Such a regulatory function
is indicated by the existence of tissue - specifi c isoforms for the subunits VIa,
VIIa, and VIII in mammals (161, 162) .
In by now standard fashion, the yeast nuclear genes have been disrupted
one at a time, and in the absence of either subunit IV, VI, VII, or VIIa there
is no cytochrome c oxidase activity and cytochromes aa
3
are missing. A knock-
out of the COX8 gene (subunit VIII) causes only a 20% reduction of activity
at normal aa
3
levels. There are two isoforms of subunit V, either one is required
for activity, but complexes with either Va or Vb differ slightly in their kinetic
and spectroscopic properties. The yeast complex may have additional subunits,
making it more similar to the mammalian complex, but it has to be clarifi ed
whether they are bona fi de subunits, scaffolding proteins required only for
assembly, or simply contaminants.
For many years, laboratories had attempted to defi ne the topology of these
subunits within the inner membrane by reactions with membrane imperme-
able reagents, by cross - linking and co - precipitation, and by an exploration of
the location of epitopes with a large variety of monoclonal antibodies (163,
164) . In light of the solution of the crystal structure, these studies need not be
reviewed here, but the development of the technology and reagents will con-
tinue to be useful tools in the examination of the enzyme, particularly in
comparisons between the normal enzyme and enzymes derived from individu-
als with mitochondrial mutations in the genes for the three largest subunits
(164) .
There are 28 transmembrane helices contributed by the various subunits.
Viewed from the top of the membrane, they form an irregular cluster surround-
ing the metal centers, in effect solubilizing the metal sites within the lipid
bilayer and also “ insulating ” the electron path from the surrounding lipid
layers. Many helices at the outside of the cluster probably have no function
214 MITOCHONDRIAL ELECTRON TRANSFER
other than to serve in the assembly and stabilization of the complex. Notably,
most of the transmembrane helices are not parallel to each other, and they are
not vertical to the plane of the membrane. Three subunits (Va, Vb, and VIb)
are not within the membrane, and they are associated as peripheral membrane
proteins with extramembrane domains of the membrane subunits.
In considering the structure in detail, it is profi table to decompose the
problem into separate structure – function relationships. The central issue is the
path of electrons from the substrate ferrocytchrome c to oxygen. Another
problem of fundamental importance is to understand the proton pumping
activity associated with electron transport. Finally, regulatory functions, stabi-
lization of the complex, and the assembly mechanism may be addressed from
a consideration of the high - resolution structure. The emergence of the struc-
ture in two major stages refl ects this order of priority.
The representation of the metal sites within the overall contours of the
complex and in relationship to the membrane was the fi rst major achievement
(157) , followed one year later by the entire structure (158) (Figures 5.18
and 5.19 ).
The two metal ions Mg
2+
and Zn
2+
are not redox centers. The Zn
2+
ion is
quite some distance away from the “ action, ” and its major function appears to
be in the stabilization of the three - dimensional structure of the complex. The
Mg
2+
ion, on the other hand, is between Cu
A
and heme a
3
, and, although not
directly involved, it may have more than a simple stabilizing role, either in
electron transport (and its direction) or in the coupling with proton transfer.
While the spatial arrangements and precise distances between the other
metal centers were fi nally determined by the crystal structure, numerous inge-
nious spectroscopic studies had contributed to a path for the electrons which
in a simplifi ed form has the following appearance:
ferrocyt c Cu Cu heme a heme a Cu O
AA 3B 2
→−→ → −→
In this scheme the Cu
A
center (with two copper ions) is located near the
surface of the complex facing the intermembrane space and hence it is able
to be brought in close contact with the reduced cytochrome c. The two heme
groups are buried within the complex as electron carriers with a redox poten-
tial determined by the surrounding protein. The unique feature is the bimetal-
lic heme iron – copper reaction center, which is also the site to which oxygen
approaches for its activation and conversion to water. To understand more
detail, one must understand the nature and redox behavior of the Cu
A
– Cu
A
center; and, most importantly, the mechanism of the transfer of electrons to
molecular oxygen from the heme a
3
– Cu
B
combination.
The Cu
A
– Cu
A
center has the [2Cu – 2S
γ
] cyclic structure in which the two
copper ions are coordinated with two cysteines, two histidines, a methionine,
and a peptide carbonyl of a glutamate of subunit II. These ligands, except for
the carbonyl group, had been proposed previously on the basis of mutagenesis
experiments that were fully confi rmed by the X - ray structures. The presence
THE ELECTRON TRANSPORT CHAIN 215
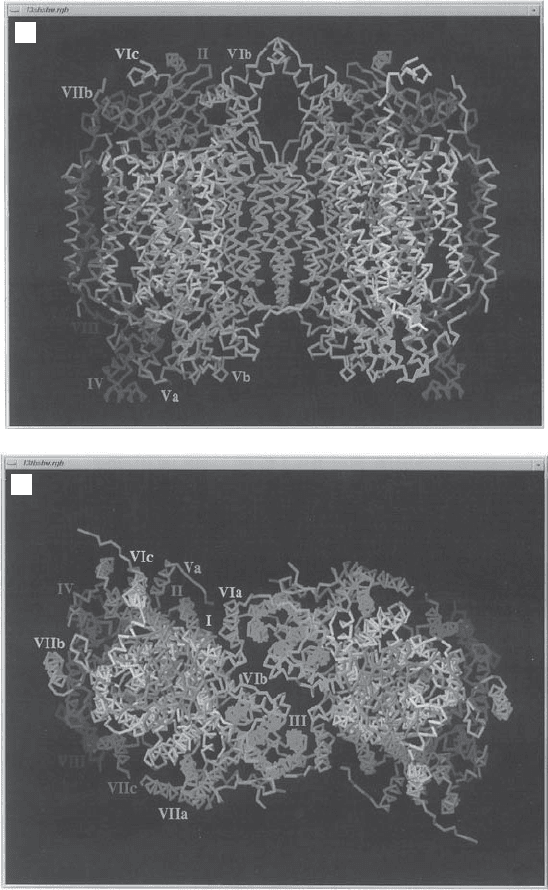
Figure 5.18 X - ray structure of complex IV. (A) Side view showing the transmembrane
segments of the various subunits; matrix side at the bottom. (B) View from the cytosolic
side. (The photographs were kindly provided by Dr. S. Yoshikawa of the Himeji Insti-
tute of Technology, Hyogo, Japan. From reference 157 with permission.) See color
plates.
B
A
216 MITOCHONDRIAL ELECTRON TRANSFER
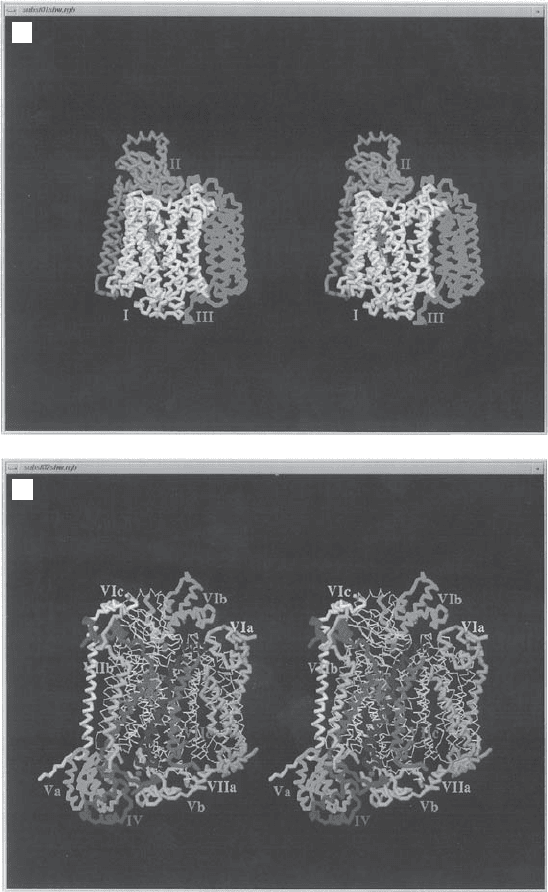
Figure 5.19 X - ray structure of complex IV. Stereo images of the C
α
- backbone tracings.
(A) Subunits I, II, and III. (B) Subunits IV, Va, Vb, VIa, Vib, Vic, VIIa, VIIb, and VIII.
(The photographs were kindly provided by Dr. S. Yoshikawa of the Himeji Institute of
Technology, Hyogo, Japan. From reference 157 with permission.) See color plates.
B
A
THE ELECTRON TRANSPORT CHAIN 217
of two copper ions per site A had long been uncertain and controversial. An
insightful account of the history of the Cu
A
site in cytochrome oxidase has been
written by H. Beinert (165) . It is highly instructive in tracing the development
of ideas about Cu in complex IV, in emphasizing the development of methodol-
ogy and techniques (purifi cation, spectroscopy, molecular genetics). Many
general concepts in protein structure with special emphasis on metal – protein
(ligand) interactions ultimately had to be applied to the understanding of a
complex with seven metal ions, three of them Cu. It now appears that there are
major similarities to the [2Fe – 2S] centers, where one - electron transfers can be
facilitated by delocalizing the charge between the two metal ions.
The most unusual and interesting structure is the binuclear heme a
3
– Cu
B
center, and major interest in this center arises from the nature of the reaction
occurring there. It is usually presented as a simple addition of electrons and
protons to oxygen:
442e
−
++ →OH HO
2
+
2
However, details of this process are more complicated (156, 166, 167) . With
20% oxygen in the atmosphere, the organic material on the earth ’ s surface
would spontaneously burst into fl ame if the activation of molecular oxygen
were simpler and energetically more favorable. The low normal reactivity of
O
2
arises from the triplet electronic ground state of oxygen, with two unpaired
electrons, which makes both single - and two - electron transfers from singlet -
state donors kinetically slow. Thus, the thermodynamically highly favorable
reduction of oxygen to water can be kept in check, except at high temperatures
or at reaction centers such as the one under discussion. The latest scheme of
oxygen activation proposed by Branden and colleagues is shown in Figure
5.20 . The cycle starts with R (Fe(II)/Cu(I)). Oxygen binds to the heme iron
(A), but the proximity of the copper ion allows two electrons from within the
center to be transferred to oxygen, converting it to the peroxy ion (O
2
2 −
) and
resulting in the oxidation of the metal ions to P
M
(Fe(IV)/Cu(II), including a
tyrosine radical at the catalytic site. The fi rst of the four electrons from the
upstream heme a enters together with a proton to form the intermediate F. In
the F → OH transition another electron and a proton are added and the
oxygen molecule is split. The proton redistribution leaves Fe(III) and Cu(II),
each coordinated with a hydroxyl ion. The hydroxyl ions are released one at
a time as water in two more steps (O
H
→ E and E → R) with the addition of
one proton and one electron in each to return to the starting position Fe(II)/
Cu(I). Many of these steps have been resolved by optical measurements at
low temperatures, or by time - resolved Raman spectroscopy (168) . For the
more intricate biophysical, thermodynamic, and kinetic aspects of this problem
the reader is referred to the specialized literature (e.g., references 156 and 167
and references therein). Not all aspects of the mechanism are resolved to
everyone ’ s satisfaction, especially with regard to the steps that drive the proton
pumping. What makes the pump unidirectional? Recent expert reviewers have
218 MITOCHONDRIAL ELECTRON TRANSFER
referred to this enzyme as a “ hysteric ” enzyme; that is, its properties are depen-
dent on its history, which depends (at least under experimental, in vitro condi-
tions) on the rate of delivery of the electrons from cyt c.
A most important consequence arises when this overall process is per-
turbed by the premature departure of partially reduced oxygen. Partial reduc-
tion yields the superoxide radical (O
2
·
−
), from which hydrogen peroxide (H
2
O
2
)
or the hydroxyl radical (OH
·
) can be derived. These species are often referred
to collectively as the “ reactive oxygen species ” (ROS), and their production
is strongly implicated in oxygen toxicity and mutagenesis of mitochondrial
DNA (see Chapter 7 ). A more elaborate discussion of the ROS and their
reactions will be presented in Section 5.7 . In the present context it is important
to clarify whether reactive oxygen species produced in mitochondria arise
from incomplete reduction of oxygen in complex IV. It has been estimated
that a signifi cant fraction (a few percent??) of oxygen entering mitochondria
is reduced only partially, and this fraction may increase under abnormal condi-
tions or as a result of electron transport complexes impaired by mutations.
Normal individuals suffer from various affl ictions when exposed to greater
than 21% oxygen, and prolonged exposure can lead to lung damage. Prema-
ture babies exposed to higher - than - normal oxygen concentrations in incuba-
tors can develop retrolental fi broblasia (ocular damage) unless carefully
monitored and given α - tocopherol as a free radical scavenger. It is generally
accepted that mitochondrial ROS production does not occur at the site
designed for reduction of oxygen in complex IV, but rather from the reaction
of the highly soluble and diffusible oxygen with reduced upstream sites of the
Figure 5.20 Proposed sequence of reactions for transferring four electrons from
cytochrome c to an oxygen molecule to form two molecules of water. (Adapted from
reference 155 .)
O
2
2+
Fe
Cu
1+
YOH
2+
Fe
Cu
1+
YOH
O O
-
4+
Fe
Cu
2+
YO
.
OH
O
2-
-
-
4+
Fe
Cu
2+
YO
OH
O
2-
4+
Fe
Cu
2+
YO
O
2-
-
H O
2
R
E
O
F
P
P
A
M
R
H
3+
Fe
Cu
2+
YOH
OH
OH
-
-
3+
Fe
Cu
1+
YOH
OH
-
H O
2
H O
2
1H
+
1e
-
1H
+
,
1e
-
1H
+
,
1e
-
1H
+
,
1e
-
1H
+
,
1e
-
THE ELECTRON TRANSPORT CHAIN 219
ETC. In other words, a leakage of electrons can occur from high potential sites,
especially in damaged or abnormal mitochondria .
The complete solution of the structure of complex IV in mammals and of
the homologous simpler complex in bacteria has greatly stimulated the discus-
sion of the mechanism of proton pumping coupled to electron transport (156,
158, 160, 167) , but here even a structure at 1.8 - Å resolution cannot give an
absolutely defi nitive answer. Previous approaches employing mutagenesis had
suggested the existence of two proton channels: one for protons that become
associated with oxygen to form water, and another for the protons pumped
across the membrane. From theoretical considerations, one would expect the
proton channels to consist of a network of hydrogen - bonded side chains whose
pK
a
’ s would be controlled by conformational changes associated with redox
reactions. Acidic groups on either side of the membrane must be alternately
accessible, and their p K must vary depending on the oxidation state of the
enzyme. Protons are shuttled through the protein complex along a string of
residues that have been referred to as a “ proton wire ” (169) . The relevant
residues were identifi ed primarily with the help of mutagenesis experiments
on the bacterial enzyme(s). Several such networks were found, and included
in these localized structures were cavities likely to contain water molecules
that can participate in proton conduction. Since the original X - ray structures
were reported in 1995, additional structures have now been examined of the
enzyme in the fully reduced, fully oxidized, azide - bound, and carbon monox-
ide - bound states (170) . These studies greatly refi ne the conformational changes
associated with the redox reaction and contribute to the formulation of more
specifi c proton pathways and models (see Figure 5.21 ). In fact, the power of
modern computers is such that the behavior of the entire enzyme can nowa-
days be simulated by molecular dynamics, with various restraints imposed
depending on the cost of the calculations (167) . The proton pathways from the
matrix to the a
3
– Cu
B
center appear to be resolved, but the subsequent path to
the IMS remains to be further elucidated. The D pathway (Figure 5.21 ) appears
to be involved in the pumping of all four protons. Although the pathways may
be identifi ed, there is still the challenge to understand how the rate and direc-
tion of movement are controlled by the electron fl ow. Finally, one must also
identify a water channel for the escape of the product, H
2
O, and possibly a
pathway through which the substrate O
2
can approach the heme a
3
– Cu
B
reac-
tion site. Potential pathways have been suggested from a consideration of the
crystal structure, but none of them are wide enough in the static crystal struc-
ture to accommodate these molecules. Thus, rapid, reversible conformational
changes have been postulated to operate in the opening and closing of such
channels. Expert, comprehensive, and up - to - date reviews (156, 167, 171 – 173)
should be consulted for further details.
Finally it should be noted that most of the more sophisticated biophysical
studies have been performed with the prokaryotic complex from Rhodobacter
sphaeroidis . In the eukaryotic enzymes the basic mechanisms of electron
transport, oxygen reduction and proton pumping are likely to be similar, but
superimposed on these will be regulatory mechanisms from ligand binding and
220 MITOCHONDRIAL ELECTRON TRANSFER
protein phosphorylation (see Section 5.6 ). Cytochrome oxidase may have the
capacity to regulate its own H
+
/ e
–
ratio — that is, the effi ciency with which
proton pumping is coupled to electron fl ow; this may act as a protective mecha-
nism to prevent the membrane potential to become too high (167) . The specifi c
role of the “ supernumerary ” subunits remains a challenge for the future.
In closing this section, one should not dwell on the outstanding issues and
unsolved problems. Instead it should be recognized that since the discovery of
the Atmungsferment by O. Warburg almost seven decades ago, this enzyme has
been one of the most important objects of study in the area of bioenergetics. As
such, it has attracted the attention of some of the greatest minds in the history
of biochemistry. The whole structure at high resolution (1.8 Å ) represents a
crowning achievement. Words fail to describe it fully, but most institutions and
many individual laboratories can now obtain the structure via the internet, or
on computer disks for display on a workstation. When combined with three
dimensional vision achieved with special goggles, the inspection and explora-
tion of this structure becomes an awsome spectacle even for the nonspecialist.
5.2.8 The Assembly of the Electron Transport Chain Complexes
While the compositions, structures, and functions of the electron transport
complexes are now understood in impressive detail, a remaining challenge is
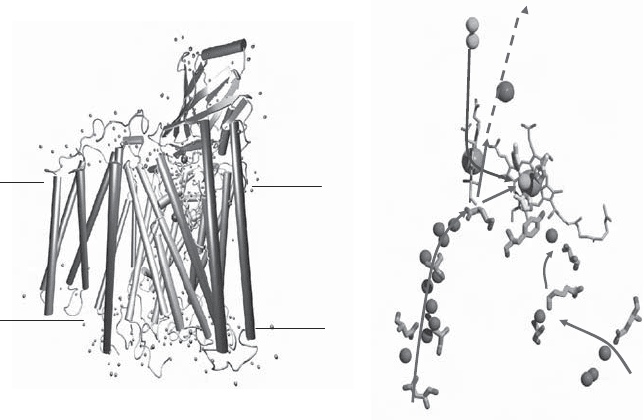
Figure 5.21 (A) Schematic view of complex IV with the four essential subunits (155) .
(B) The proposed D and K pathways for the entry of protons to form water and to be
pumped across the membrane. The path from the region of the hemes to the outside
is still poorly defi ned. (From reference 155 with permission.) See color plates.
A
SU II
SU I
SU III
SU IV
P side
N
side
Cu
A
Cu
B
heme a
3
heme a
Y288
T359
S299
K362
E101
II
E286
Mg
2+
D pathway
K pathway
D132
N139
N121
S201
e
−
H
+
H
+
B
