Vaccari D.A., Strom P.F., Alleman J.E. Environmental Biology for Engineers and Scientists
Подождите немного. Документ загружается.

Answer 10 mm/unit 9.3 units ¼93-mm-longciliate. 10 mm/unit 161 units ¼
1610 mm ¼1.61-mm field diameter.
11.2.2 Electron Microscopes
Electron microscopes utilize electron beams instead of light and thus are able to achieve
greater resolution (Figure 11.8). The two major types are the transmission electron micro-
scope (TEM) and the scanning electron microscope (SEM).
The TEM is analogous to a light microscope, with a beam of electrons being trans-
mitted through the sample rather than light. Instead of glass lenses, the TEM uses electro-
magnets to control and focus the beam. The result is the p otential for much higher
magnification, with levels of resolution down to a few tenths of a nanometer (nearly
1000 times better than what can be achieved with a light microscope).
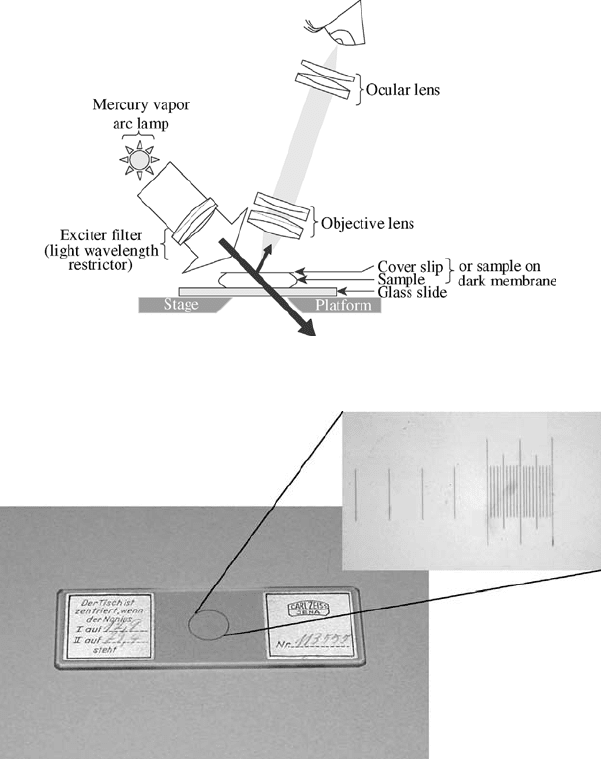
Figure 11.6 Fluorescence microscopy: a fluorescent image is viewed against a darkened or black
background.
Figure 11.7 Stage micrometer used to calibrate dimensions under microscope.
298
QUANTIFYING MICROORGANISMS AND THEIR ACTIVITY
The disadvantages of TEM microscopy include the much highe r cost and complexity,
and the need for sophisticated preparation of samples, such as cutting thin sections of the
cells to be observed. Of course, these techniques kill the cells. A main use of TEM is
looking at internal cell structures.
SEM offers somewhat less magnification, with resolutions down to around 10 nm. It
is based on the detection of backscattered rather than directly transmitted electrons. Typi-
cally, the specimen’s surface is treated with an electron reflective coating, such as gold. As
the applied electron beam is scanned back and forth across this sample , the scattered elec-
trons provide a three-dimensional image of the coated surface. However, care must be
taken that the coating procedure itself does not produce artifacts (false observations).
11.3 SAMPLING, STORAGE, AND PREPARATION
11.3.1 Sampling
Sampling is a critical, although often insufficiently considered, part of most analyses. The
best that any analytical procedure can hope to do is to determine the analyte in the sample
accurately. Thus, the qual ity of the information received about the system of interest can
only be good as the samples on which it is based.
Sampling of microorganisms can provide some additional concerns because of their
special nature. Extra care must be taken to avoid contamination, since even a small num-
ber of inadvertently added organisms could grow quickly and produce a large error. On
the other hand, toxic agents present in the sample could continue to kill organisms, lead-
ing to underestimates of their numbers. The chlorine residual left after disinfection of
water or wastewater is a common example of this. Also, in many systems microorganisms
will adhere to surfaces. This might make special sampling techniques necessary.
11.3.2 Storage
Many types of microbial analysis must be done almost immediately. If not, the microbes
may continue to grow or die, changing their total numbers and also the relative ratio of
various species. Refrigeration will help in many cases, but storage times are still typically
limited to only a few hours. This means, among other things, that composite samples
Light
SEM
ST or SP
TEM
1 Å 10 Å
100 µm 10 µm 1 µm
100 nm
0.1 µm
10 nm 1 nm
0.1 nm
100 Å
Electron Microscopes
Eukaryote
cells
Prokaryote
cells
Viruses
Proteins
Amino
acids
Atoms
Figure 11.8 Resolution with various microscopes. Electron microscopes include scanning (SEM),
transmission (TEM), and scanning tunneling (ST) or scanning probe (SP).
SAMPLING, STORAGE, AND PREPARATION 299
(samples made by mixin g several individual, or grab, samples) taken over a 24-hour
period to try to determin e average conditions are not acceptable for many microbial
tests. On the other hand, some methods (e.g., some DNA analyses) allow rapid freezing
or other preservation techniques for long- term storage.
11.3.3 Preparation
For some samples, the numbers of microorganisms may be too low for the desired ana-
lysis, and a concentration technique will first be employed. This might involve centrifu-
ging a large sample to collect the organisms in a smaller, more concentrated volume.
Alternatively, filtration might be used to collect the organisms from a large volume on
the filter surface. In another type of filtration, planktonic (floating) eukaryotes may be
concentrated during collection by using a plankton net that is dragged through the
water. For viruses (and a few cellular organisms), adsorption in a column of packed
solid material, followed by elution into a small volume, may be used.
In other samples, conce ntrations of microorganisms may be too high for counting,
resulting in the need for dilution. This might be with filtered water from the same source,
or more commonly with a standard phosphate buffer or saline solution. In most case s,
distilled or other high-purity water cannot be used, as it would cause an osmotic shock
to the organisms (and possibly their death). A common approach is to make serial
dilutions. Often, this consists of making a 10-fold dilution, then using that to make
another 10-fold dilution, and repeating until the desired concentration is reached.
If microorganisms are adhering to a surface, it may be necessary to try to remove them.
If they are adhering to each other , or to small particles of soil or other material, a dispersion tech-
nique such as blending is sometimes used in an attempt to separate them. Dispersing adhered or
aggregated microorganisms remains an important problem in the analysis of many samples.
Removing extraneous or interfering materials from a sample also may be useful. Some
chemicals can be removed by adding a reagent to react with them (e.g., sodium thiosul-
fate, Na
2
S
2
O
3
, can be added to eliminate residual chlorine). Soluble interfering substances
might be separated from the microbes by filtration or centrifugation. Removing particu-
lates can be more difficult, since microorganisms may adhere to them.
11.4 DETERMINING MICROBIAL BIOMASS
11.4.1 Measurem ents of Total Mass
Prokaryotic cells are usually incredibly minute, with a typical individual mass on the
order of only 10
12
g [1 picogram (pg) ]. Since an average adult human typically has a
mass of 50 to 100 kg, we are more than 10
16
times larger. This number is so huge that
it does not even have a commonly known name (a trillion is ‘‘only’’ 10
12
). To put things in
perspective, this is about the same ratio as the few seconds you spend reading this sen-
tence compared to the entire 4.6 billion year existence of our planet.
Thus, the mass of these cells cannot normally be measured individually. However, in a
number of cases it may be important to know the collective biomass of microorganisms
present in a system. In laboratory cultures grown on soluble media, this can be measured
by filtration followed by washin g (to remove salts), drying, and direct weighing of the
material collected, giving a dry weight. This is also done for suspensions in systems
such as activated sludge, even though most of the material included as ‘‘biomass’’ is
300 QUANTIFYING MICROORGANISMS AND THEIR ACTIVITY
not actually living cells. (It is believed that more than 90% is commonly dead cells or
other nonliving organic particulate material.) Similarly, biofilms and growths on the sur-
face of a laboratory agar plate can be scraped off, suspended in buffer, and measured in
this way, although for biofilms, again, much of the ‘‘biomass’’ will be film polymers and
other nonliving materials. Biomass in the air (or other gases) might be estimated in a simi-
lar way after collection by filtration, although in most cases the nonliving material would
again be a major interference.
One correction that is sometimes used for these methods based on suspended solids
(SS) is to consider only the volatile suspended solids (VSS) fraction. The SS sample is
ashed (or burned off by heating to 500
C) to drive and burn off the volatile materials
(which are usually mainly organic), and the change in mass determined. This can be used
to correct for the presence of inorganic materials, such as salts and metal oxides, but does
not differentiate between living and nonliving organic matter. Also, 15% of a microor-
ganism’s dry mass typically will be ash (Section 11.1.1).
In some cases, centrifugation can be substituted for filtration. The compacted solids,
referred to as the pellet, may be dried and weighed, usually after washing (resuspending
in liquid and centrifuging again). In an application used at some sewage treatment plants,
operators centrifuge a sample of the sludge and compare the pellet volume with a calibra-
tion chart to provide a rough estimate of the mass. This test can be rapidly done (10 min-
utes) for process monitoring using only an inexpensive small centrifuge, compared to the
more time-consuming and expensive steps of filtration, drying (requiring an oven), and
precise weighing (requirin g an analytical balance).
These types of measurements are not practical for many other types of samples, such as
soils, solid wastes, and most sludges. In those case s, as well as often for those described
above, counts are performed instead of mass measurements. If desired, mass might then
be estimated from the number of microorganisms (Section 11.5.3). However, counting has
its own limitations in such samples (see below).
One other approach that has been used with varying success in soils is referred to as a
fumigation technique. The microorganisms in a sample are killed by exposure to a toxic
gas such as chloroform. The gas is then removed and the sample is inoculated with a
small number of newly added bacteria. Since the killed microorganisms then serve as food
for new growth, the amount of carbon dioxide that is produced by the microbial activity can
be related (using proper controls and calibration) to the amount of biomass originally present.
11.4.2 Measurem ents of Cell Constituents
Rather than try to measure the total mass of organisms present, sometimes it is more prac-
tical or desirable to measure the mass of a particular cell const ituent. For example, it may
be possible to treat a sample to extract the microbial protein or DNA that is present and to
quantify it by weight or by a chemical means. The amount of such fundamental cell con-
stituents present may then be correlated to the biomass present (see Table 11.4, for exam-
ple). In some cases these measures may be even more useful then total biomass estimates,
since the amounts per cell vary less than for some other constituents, such as exocellular
polymer or food storage products.
Another cell constituent of interest for estimating biomass is ATP. It is present in all
organisms, and since it disappears quickly upon death of a cell, it can serve as a useful
indicator of viabl e (living) biomass. However, the amount of ATP present in a cell can
vary considerably with its physiological state.
DETERMINING MICROBIAL BIOMASS 301
Planktonic algae concentrations can often be estimated by measurement of chloro-
phyll a. This method also can be used for cyanobacteria, and in fact includes them in
the estimates obtained. The sample is prepared by grinding the cells and extracting the
pigment with acetone. The chlorophyll is then quantified by measuring absorbance at
664 nm with a spectrophotometer, or fluorescence with a fluorometer, or by high-perfor-
mance liquid chromatography (HPLC). Corrections may be necessary due to the presence
of other pigments.
Some methods have been developed to extract and identify cell constituents that are
characteristic of a particular group of microorganisms, including cell wall constituents,
fatty acids, and DNA or RNA seque nces that are specific for particular taxa. So far
these are used primarily to help identify what types of microorganisms are present, but
quantitative methods are also being developed that can be used to estimate biomass.
11.5 COUNTS OF MICROORGANISM NUMBERS
Counts of microorganisms may be made by direct microscopic techniques or through
indirect methods such as culturing.
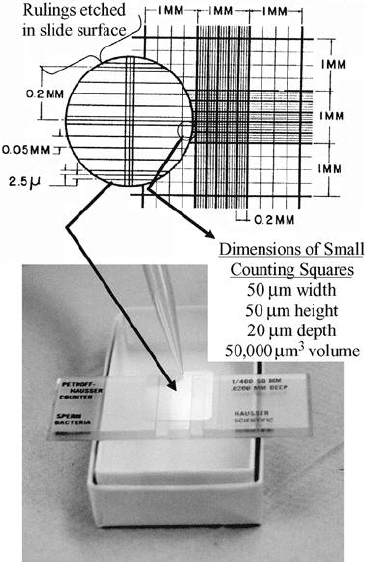
Figure 11.9 Petroff–Hauser counting cell.
302
QUANTIFYING MICROORGANISMS AND THEIR ACTIVITY
11.5.1 Direct Counts
In some samples, microorganisms can simply be counted under the microscope. Counting
chambers—specialized microscope slides with wells holding a fixed volume—are avail-
able for this purpose. Some, such as the Petroff–Hauser cell (Figure 11.9), also have a grid
marked on them. An example of a chamber with a deeper well used for larger organisms
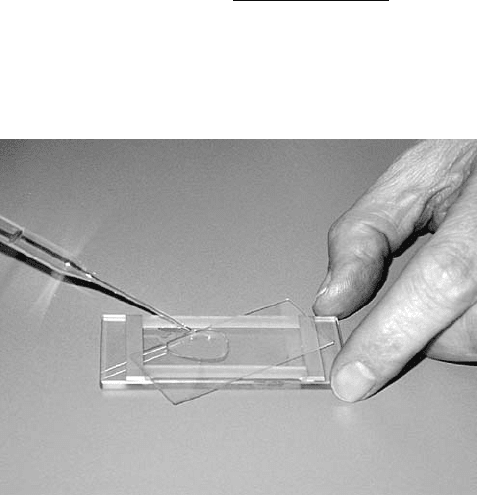
or associations such as filaments or floc is the Sedgwick–Rafter cell (Figure 11.10); it has
an area of 1000 mm
2
and a depth of 1 mm, thus holding a volume of 1.0 mL. Fresh live
samples are used when possible, especially for protozoa and metazoa, so that activity such
as motility or feeding can be observed.
Alternatively, a fixed volume (usually 1 drop, or about 0.05 mL ¼50 mL) can be added
to a regular slide, and a coverslip (usually square, 22 to 25 mm on a side) placed over it
(spreading the sample under the coverslip). The number of cells seen in one field (the
visible area seen through the eyepiece, usually about 1.6 mm in diameter at 100 and
0.16 mm at 1000) under the micro scope is then multiplied by the ratio of the area of
the coverslip to the field area to get the concentration (count per volume).
Example 11.2 An average of 3.2 ciliated protozoans were seen per field when 10 fields
were counted at 100. If the sample size was 0.05 mL, the microscope field at that mag-
nification is 1.6 mm in diameter, and a 22 22 mm coverslip was used, what is the con-
centration of ciliates present?
Answer Area of field A
f
¼ pr
2
¼ pð1:6=2mmÞ
2
¼ 2:01 mm
2
=field
Area under coverslip A
c
¼ 22 mm 22 mm ¼ 484 mm
2
=coverslip
Conc: of ciliates ¼ 3:2= field
484 mm
2
=coverslip
2:01 mm
2
=field
0:05 mL=coverslip
¼ 15;000=mL
Figure 11.10 Filling a Sedgewick–Rafter cell.
COUNTS OF MICROORGANISM NUMBERS 303
For observation of highly motile protozoa, it is sometimes desirable to slow down their
activity. This can be done by adding a compound such as methylcellulose to increase
viscosity, or by using an inhibitory compound such as nickel sulfate.
Although small metazoa such as rotifers and nematodes can often be enumerated along
with the protozoa, larger forms may require specialized tec hniques. Some, in fact, may
actively avoid being drawn into a small sampling or subsampling device, such as a pipette.
Others, as well as larger associations of proka ryotes such as large floc, may simply not fit
through the opening of fine tip pipettes.
Because of their silica shells, a special technique can be used for diatoms. After wash-
ing of the sample with distilled water, organic matter is destroyed by heat or an acid-
oxidation step, leaving the shells for counting.
Note that using appro ximations from the above example, 5000 cells/mL (500 mm
2
/
2mm
2
divided by 0.05 mL) would be needed in a sample for there to be one cell on aver-
age per microscope field at 100. For prokaryotic cells, which are usually too small to
count at low magnification, concentrations of above 10
6
mL
1
(2 per field at 1000) are
usually necessary for direct counting. If there are too many cells to count (>10
8
mL
1
),
the sample might be diluted first. If the sample contains too few microorganisms, filtration
might then be used to concentrate them on a filter surface for viewing.
However, even with dilution or concentration, this simple counting approach is limited
to samples with little extraneous material that would prevent clear viewing and that con-
tain individual, dispersed cells rather than cells in flocs or biofilms. To allow direct count-
ing in samples such as activated sludge (flocculated), biofilms, and soils, a variety of
stains and probes have been developed, including many that are fluorescent. These
allow differentiation between inert particles and cells, between living and dead cells, or
even among specific strains of organisms.
One such approach uses a diacetate ester of fluorescein, which is able to pass into the
cytoplasm of cells. Once inside, the ester is rapidly hydrolyzed by nonspecific est erase
enzymes. The free fluorescein (which, as its name suggests, is fluorescent ) that is released
is trapped in the cell, thereby making it readily visible with a fluorescence microscope.
The acridine orange direct count (AODC) uses another fluorescent dye that passes
into the cytoplasm and then binds to nucleic acids, g iving an orange or green color.
This does not require the activity of enzymes within the cel l and thus may give a higher,
total count, including both viable and nonviable (dead) cells.
One problem with the AODC method is that clay particles also may appear orangish,
potentially interfering with cell counts. This has contributed to the increased popularity of
a blue fluorescent dye, 4
0
,6-diamido-2-phenylindole (DAPI), which binds more specifi-
cally with DNA.
If antibodies to the surface of a specific organism can be produced, the fluorescent
antibody technique can be used. In this method the antibody is conj ugated with (chemi-
cally attached to) a fluorescent dye. When this preparation is added to the sample, it
attaches to the cell so that the surface of the target organisms fluoresce.
Genetic probes can also be used to stain specific groups of organisms for counting.
The fluorescent in situ hybridization (FISH) technique (Section 10.4.4) is one popular
example.
Another alternative is the use of metabolic stains. These typically utilize a dye that is
reduced by (accepts electrons from) a cell’s metabolic activities to form a colored product.
A number of tetrazolium dyes such as 2-(p-iodophenyl)-3-(p-nitrophenyl)-5-phenyl tet-
razolium chloride (INT ), for example, are capable of passing through the cell membrane
and then accepting electrons by way of dehydrogenase enzymes. The result is conversion
304 QUANTIFYING MICROORGANISMS AND THEIR ACTIVITY
to dark red to deep-purple formazan crystals that are trapped within the cell. Thus, a
count of respiring cells can be made. The formazan can also be eluted with isopropyl alco-
hol and an indirect measure of cell concentration (number or mass) made from a spectro-
photometric measurement of the intensity of the color released.
Filamentous bacteria (including act inomycetes, many cyanobacteria, and some gram
negatives, such as Sphaerotilus and Beggiatoa), fungi, and algae can pose a problem in
quantification in that a single ‘‘count’’ might be composed of only one cell a few micro-
meters long or might contain many cells extending to a length of 1 mm or more. In some
cases the number of cells in the filament may be counted or the filament length measured,
perhaps for conversion to a biomass estimate (similar to the calculations of Section
11.5.3). Similarly, counts of colonial protozoa and algae may require some enumeration
of individuals or a measure of the size of the colony. For bacterial associations such
as floc, individual counts may be impossible, but the size of the association may be
measured.
Image analysis is a tool that can be combined with several of the techniques described
above. A digital image of the microscope field is captured and can then be analyzed using
computer software to pick out and quantify specific size (including filament length or floc
size), shape, and/or color (including from stains and fluorescence) objects. This informa-
tion can then be provided as total or differential counts, and can also be used to estimate
cell volume (and hence mass; Section 11.5.3).
11.5.2 Indirect Methods
Several commonly used methods of estimating the number of microorganisms in a sample
depend on culturing them in or on a medium that supports their growth. These methods
require the use of aseptic technique, in which sterile materials (initially containing no
organisms) are used along with a variety of measures to prevent contamination by
other organisms (e.g., from the air, the analyst’s hands, or the lab bench surface). They
also involve a period of incubation (time for growth under designated conditions) of from
one to several days for most common tests.
In one type of approach (use d for most plate count and membrane filtration methods;
see below), the organisms are grown on a solid surface until each initial cell produces a
visible colony consisting of many millions of daughter cells. The number of colonie s is
then counted and gives an estimate of the number of colony-forming units (CFUs) origin-
ally present. It is usually assumed that each colony arose from a single cell (or spore), but
in some cases it is likely that a small cluster of cells or a filament was present initially and
gave rise to the colony. This is especially problematic for fungi and actinomycetes, whose
counts can increase dramatically with sporulation even though little change in the actual
biomass present has occurred.
Another approach (used in most probable number techniques and some plate count and
membrane filtration methods) is to grow the cells in or on a medium until they produce
some detectable change that indicates their presence. This might be a change in pH,
increased turbidity, the disappearance of a substrate, or the appearance of a product.
Often, color indicators are used to make the activity readily apparent.
One advantage of culturing techniques is that they often can be performed in a way that
minimizes interferences from extraneous materials. They may also be used to count cells
at lower densities than direct methods (without a concentration step). However, culturing
requires a medium on which the organisms of interest can grow. Since no single
medium is suitable for all prokaryotes, a true total count cannot be obtained. In fact, total
COUNTS OF MICROORGANISM NUMBERS 305
culturable bacterial counts in many samples are one to two orders of magnitude lower
than total direct counts.
On the other hand, media can be made intentionally selective, so that only certain
organisms will grow. For example, a selective substrate can be used, or a chemical that
is inhibi tory to nontarget organisms can be incorporated. Similar ly, selective tempera-
tures, salinities, or pHs can be used. Additionally, media can be differential, so that
among the organisms that do grow, the ones of interest can be told apart from others.
Often, this is done by a color reaction.
Some common approaches are described briefly below. By varying the medium used
and the incubation conditions (e.g., temperature, aero bic vs. anaerobic), a wide variety of
microorganisms can be enumerated.
Plate Counts Probably the most common methods of estimating bacterial numbers are
plate counts. These use agar, or occasionally, some other solidifying agent, to convert
liquid medium (broth) into a solid. Agar is particularly useful because it does not melt
in water until heated to boiling, but it does not resolidify until it is cooled below 45 to
48
C. However, because some bacteria can digest agar (an organic polymer produced
by certain red algae), it cannot be used in some special applications.
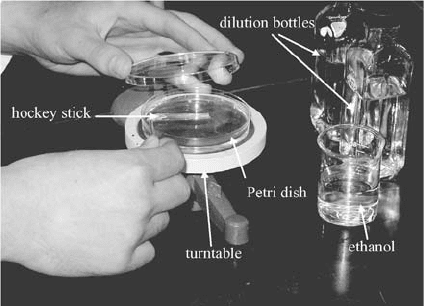
In the spread plate technique (Figure 11.11) , an agar medium (often, 10 to 15 mL) is
added to a small (commonly 100 mm in diameter by 15 mm high) sterile glass or plastic
petri dish and allowed to solidify. A small (0.1 mL) amount of sample or dilution is then
placed on top and spread over the surface of the medium using a bent glass rod called a
hockey stick. Visible colonies that grow on the surface of the medium after incubation are
counted.
Example 11.3 If an average of 38 colonies is counted for plates containing 15 mL of
agar medium on which 0.1 mL of a 10
4
dilution of a sample was spread-plated, what
was the concentration of CFUs in the original sample?
Answer
CFUs=mL ¼ 38 colonies=plate 0:1mL 10
4
¼ 3:8 10
6
CFU=mL
Note that the amount of agar is irrelevant.
Figure 11.11 Spread plating.
306
QUANTIFYING MICROORGANISMS AND THEIR ACTIVITY

For pour plates, a 1-mL sample or dilution is put on the bottom o f a petri dish. Then
10 to 20 mL of the desired molten agar medium is added, and the dish is covered and
swirled to mix the cont ents. The medium solidifies with the microorganisms distributed
throughout it. After incubation, the visible colonie s are counted.
Aside from the other limitations of methods involving culturing, incubation time is also
an important factor affecting the accuracy of plate counts. If insufficient time is allowed,
of course, some colonies will not yet have grown to visible size. On the other hand, if too
much time is allowed, some colonies may grow so large that they cover up others, again
leading to underestimation. Alternatively, secondary colonies may form (e.g., from motile
bacteria swimming away to start new colonies), giving overestimations.
Plate counts are typically suitable for 20 to a few hundred CFUs per plate. Lower
counts have poor precision, and at higher numbers they crowd each other and interfere
with growth. If cell density is too high, the plate is recorded as TNTC (too numerous
to count), and it is recognized that a greater dilution factor should have been used.
Since it is often difficult to know ahead of time how many organisms are present, usually
a range of dilutions are plated; some may end up with no colonies, others TNTC, but
hopefully those in the middle will have countable numbers.
Membrane Filtration Methods Microorganisms can be collected on the surface of a
membrane by filtration (Figure 11.12). If the membrane filter is then incubated on the sur-
face of a growth medium in a small (usually, 50 mm in diameter) sterile petri dish, the
colonies that form give an estimate of the original numbers present. The medium can
either be a prepoured agar (similar to a spread plate) or a broth contained in an absorbent pad.
Bunsen
burner
to vacuum
low power
microscope
for counting
filter funnel
forceps in
ethanol
membrane filter
dilution
bottle
Bunsen
burner
to vacuum
low power
microscope
for counting
filter funnel
forceps in
ethanol
membrane filter
dilution
bottle
Figure 11.12 Membrane filtration.
COUNTS OF MICROORGANISM NUMBERS 307
