Klipp E., Herwig R., Kowald A., Wierling C., Lehrach H. Systems Biology in Practice: Concepts, Implementation and Application
Подождите немного. Документ загружается.


hybrids by complementary base pairing. Instead, it is based on the specific interac-
tion between antibody and antigen.
4.1.4.1 Southern Blotting
The technique of Southern blotting was invented in 1975 and is used to analyze
complex DNA mixtures (Southern 1975). Normally, the target DNA is digested by re-
striction enzymes and separated by gel electrophoresis. If the DNA is of genomic ori-
gin, the number of resulting fragments will be so large that no individual bands are
visible. If we are interested in a certain gene and know its sequence, small DNA frag-
ments can be synthesized, which are complementary to the gene. However, before
the actual hybridization step, the digested DNA has to be transferred from the gel
onto the surface of a nitrocellulose or nylon membrane so that the DNA molecules
are accessible for the hybridization. This is achieved with the help of a blotting appa-
ratus, as shown in Fig. 4.5. Originally the transfer was achieved by placing the nitro-
cellulose filter between the gel and a stack of blotting paper. Capillary forces lead to a
flow of water and DNA fragments from the gel onto the blotting paper. In this way,
the DNA gets trapped by a nitrocellulose or nylon filter. Nowadays, blotting ma-
chines that transfer the DNA by applying a voltage across the gel and membrane are
used. Once the DNA is blotted, the membrane is placed into a plastic bag and incu-
bated for several hours with a solution containing the labeled DNA probe. In the
case of a radioactive label, the membrane is placed against an X-ray film. The radio-
active DNA fragments expose the film and form black bands that indicate the loca-
tion of the target DNA.
With this technique, not only can the presence of the gene of interest be tested,
but also modifications of the gene structure (in the case of a mutation) can be stu-
died. By performing several Southern blots with DNA probes that correspond to dif-
ferent regions of the gene, modifications such as deletions and insertions can be de-
tected. Point mutations, however, cannot be identified with Southern blotting.
120
4 Experimental Techniques in a Nutshell
Fig. 4.5 Elementary steps required for South-
ern blotting. Following gel electrophoresis, the
DNA fragments are treated with an alkaline solu-
tion to make them single-stranded. The nitrocel-
lulose or nylon membrane is sandwiched be-
tween the gel and a stack of blotting paper and
the DNA is transferred onto the membrane
through capillary forces. Finally, the membrane
is incubated with the labeled DNA probe (here,
radioactive labeling) and the bands are then vi-
sualized by X-ray film exposure. (Courtesy of Dr.
P. Weingarten, Protagen AG.)

4.1.4.2 Northern Blotting
Northern blotting is very similar to Southern blotting. The only difference is that
mRNA, not DNA, is used for blotting. Although the experimental technique is very
similar, Northern blotting can be used to answer different questions than Southern
blotting. Even though mRNA is only an intermediate product on the way from the
gene to the protein, it is normally a reasonable assumption that the amount of
mRNA is correlated to the amount of the corresponding protein in the cell. Northern
hybridization is therefore used not only to verify the existence of a specific mRNA
but also to estimate the amount of the corresponding protein via the amount of
mRNA. Since the expression profile of genes varies among tissues, Northern blot-
ting gives different results for different organs, in contrast to Southern blotting,
which is based on genomic DNA.
Surprisingly, the technique did not get its name from the researcher who invented
it; rather, it was named as a humorous allusion to the double meaning of the name
of E. M. Southern, who gave his name to the Southern blot.
4.1.4.3 Western Blotting
So far we have seen techniques for blotting different types of nucleic acids. A similar
type of technique exists for proteins, the Western blot. As with the Northern blot, its
name indicates only that it is a special type of blotting technique. Depending on the
problem at hand, 1D or 2D protein gels can be used for blotting. It is more difficult
to obtain specific probes for proteins than for nucleic acids. Apart from special cases,
antibodies that are directed against the desired protein are used. If no commercial
antibodies are available for the protein of interest, they have to be produced, e.g., by
immunizing rabbits.
Once the protein is transferred to the nitrocellulose membrane, it is incubated
with the primary antibody. The primary antibody recognizes the protein and forms
an antibody-protein complex with the protein of interest. In a further step, the mem-
brane is incubated with the so-called secondary antibody, which is an antibody
against the primary antibody. If the primary antibody was obtained by immunizing a
rabbit, the secondary antibody could be a goat-anti-rabbit antibody. This is an anti-
body from a goat that recognizes all rabbit antibodies. The secondary antibody is che-
mically linked to an enzyme, such as horseradish peroxidase, that catalyzes a chemi-
luminescence reaction, and exposure of an X-ray film finally produces bands, indi-
cating the location of the protein-antibody complex. The intensity of the band is pro-
portional to the amount of protein. Unlike Northern blotting, Western blotting gives
a direct estimate of the protein content. The secondary antibody serves as signal am-
plification step. This is the reason that the enzyme is not linked directly to the pri-
mary antibody.
4.1.4.4 In situ Hybridization
The described blotting and hybridization techniques are applied to mixtures of nu-
cleic acids or proteins that have been extracted from cells or tissues. During this pro-
cess, all information about the spatial location is lost. In situ hybridization solves this
problem by applying DNA probes directly to cells or tissue slices.
121
4.1 Elementary Techniques

One common application is the location of specific genes on chromosomes. For
this purpose metaphase chromosomes, which have been exposed to a high pH to se-
parate the double strands, are incubated with labeled DNA probes. This makes it
possible to directly see where and how many copies of the gene are located on the
chromosome. If the label is a fluorescent dye, the technique is called FISH (fluores-
cent in situ hybridization). Not only chromosomes but also slices of whole tissues
and organisms can be hybridized to DNA probes. This can be used to study the spa-
tial and temporal expression pattern of genes by using probes specific to certain
mRNAs. This method is often used to study gene expression patterns during embry-
ogenesis. Finally, immunostaining uses antibodies to localize proteins within cells.
Knowledge about the subcellular localization often helps one to better understand
the functioning or lack of functioning of the studied proteins.
4.1.5
Further Protein Separation Techniques
4.1.5.1 Centrifugation
One of the oldest techniques for the separation of cell components is centrifugation.
This technique fractionates molecules (and larger objects) according to a combina-
tion of size and shape. However, in general it is true that the larger the object, the
faster it moves to the bottom. A typical low-speed centrifugation (around 1000-fold
gravitational acceleration, g) collects cell fragments and nuclei in the pellet, at med-
ium speeds (50,000 g) cell organelles and ribosomes are collected, and at ultrahigh
speeds (up to 500,000 g) even typical enzymes end up in the pellet. Of course, such a
separation is not quantitative, but repeating the centrifugation after re-dissolving the
pellet improves the purity. The sedimentation rate for macromolecules is measured
in Svedberg units, S, after Theodor Svedberg, who invented ultracentrifugation in
1925. S depends on the mass (m) and density of the particle (r
par
), as well as on the
density (r
sol
) and friction ( f ) of the medium. These properties control the velocity
(v) of the particle (o
2
= angular velocity, r = distance from center of rotation).
S
v
o
2
r
m 1 r
sol
=r
par
f
(4-2)
The ribosomal subunits, for instance, got their name from their sedimentation
coefficient (40S subunit and 60S subunit). Because the friction is controlled not only
by the size of the particle but also by its shape, S values are not additive. The com-
plete ribosome (40S plus 60S) sediments at 80S and not at 100 S.
From Eq. (2) it also follows that the sedimentation rate is zero if the densities of
the particle and the surrounding medium are identical. This is the basis for the equi-
librium centrifugation method, in which the medium forms a stable density gradi-
ent (caused by the gravitational forces) such that the density of the studied particles
lies within the density range of the gradient. If the centrifugation is run long en-
ough, the particles move to the position where the densities of the medium and the
particle are identical and form stable bands there. Thus, equilibrium centrifugation
122
4 Experimental Techniques in a Nutshell
separates the molecules by density, independent of their size. The necessary density
gradients are typically formed with saccharose or cesium chloride (CsCl).
4.1.5.2 Column Chromatography
Other classical separation techniques include the different forms of column chroma-
tography (Fig. 4.6). A column (often made of glass, a few centimeters wide and a few
dozen centimeters tall) is filled with a solid carrier material and the protein mixture
is placed on top of it. Then a buffer is slowly washed through the column and takes
the protein mixture along with it. However, different proteins are held back to a dif-
ferent degree by the column material and arrive at different times at the bottom of
the column. The eluate can be fractionated and tested for the presence of the desired
protein. The ratio of desired protein to total protein is a measure of the purity, which
can additionally be checked on normal SDS gels. Different column materials are
available and the success of a separation often depends critically on the choice of ma-
terial.
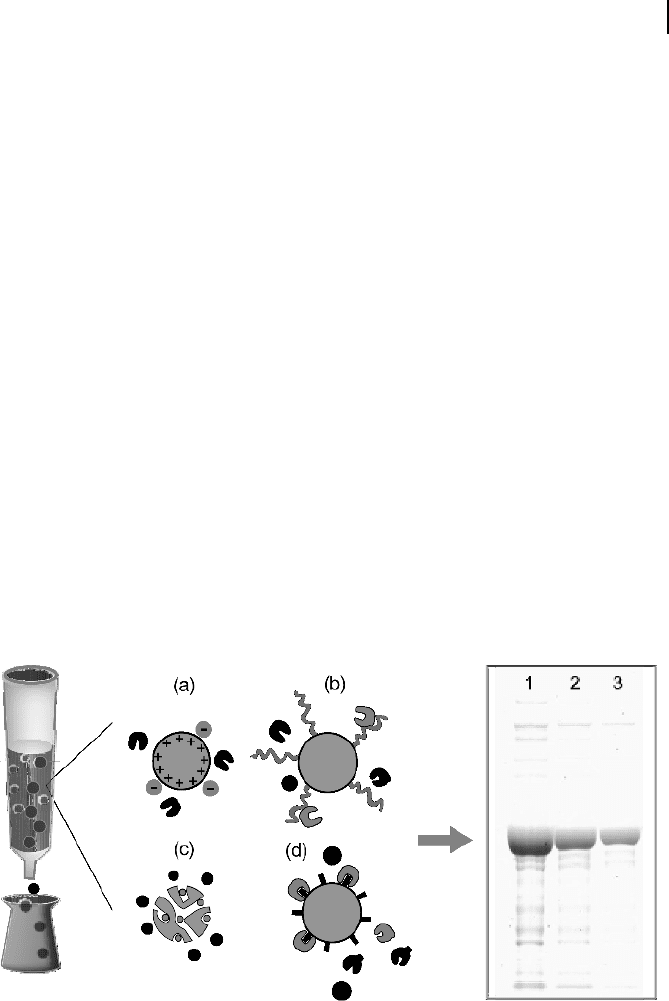
The material for ion exchange chromatography (Fig. 4.6a) contains negatively
(carboxymethyl or phosphor cellulose) or positively (diethylaminoethyl cellulose)
charged beads that can be used to separate hydrophilic proteins according to charge.
The binding between the proteins and the beads is also controlled by the salt concen-
tration and pH of the elution buffer. Some proteins possess hydrophobic surfaces that
can be used to separate proteins by hydrophobic interaction chromatography (Fig.
4.6b). For this purpose short aliphatic side chains are attached to the surface of the
column material. Gel filtration chromatography (Fig. 4.6c) is also often used to sepa-
rate proteins according to size. The beads contain a range of pores and channels that
123
4.1 Elementary Techniques
Fig. 4.6 In column chromatography a protein
mixture is placed on top of the column material
and then eluted with buffer. Different types of
material are available to separate the proteins
according to (a) charge, (b) hydrophobicity,
(c) size, or (d) affinity to a specific target mole-
cule. Often, different chromatographic steps
have to be used successively to purify the desired
protein to homogeneity (lanes 1–3 of the gel).
(Courtesy of Dr. P. Weingarten, Protagen AG.)

allow small molecules to enter, which increases their retention times. This allows
not only their separation by size but also estimation of the absolute size of a protein
or protein complex. A more recent development is affinity chromatography (Fig.
4.6d), which makes use of highly specific interactions between a protein and the col-
umn material. This can be achieved, for instance, by chemically linking antibodies
to the column material. For affinity chromatography the column is loaded with the
protein mixture in the first step. The proteins of interest bind, while the other pro-
teins pass through the column. In the second step, the elution process is started by
using a high-salt or high-pH buffer that frees the bound protein from the column.
A major improvement regarding speed and separating power was achieved
through the development of high performance liquid chromatography (HPLC). The
columns are much smaller and the carrier material is packed more densely and
homogenously. To achieve reasonable buffer flow rates, very high pressures (up to
several hundred atmospheres) are needed. Therefore, special pumps are used and
the column housing is made of steel.
The enrichment factor of a single chromatographic step is normally between 10-
and 20-fold. However, since many proteins represent only a tiny fraction of the total
protein content of a cell, different chromatographic columns often have to be used
consecutively. A notable exception is affinity chromatography, which can achieve en-
richments up to 10
4
in a single step. In combination with modern recombinant
DNA techniques, affinity chromatography has many applications. Recombinant pro-
teins can be designed to contain special short sequences of amino acids that do not
compromise the functionality of the protein but can serve as molecular tags. A short
stretch of histidine residues is called a His-tag and is specifically recognized by a
nickel surface or special His antibodies.
4.2
Advanced Techniques
4.2.1
PCR
The polymerase chain reaction (PCR) was invented in the mid-1980s and allows the
billion-fold amplification of specific DNA fragments (typically up to 10 kbp) directly
from genomic DNA (Saiki et al. 1985). A pair consisting of short oligonucleotides
(15–25 bp), the primers, is synthesized chemically such that they are complementary
to an area upstream and downstream of the DNA of interest. DNA is made single-
stranded by heating, and during the cooling phase primers are added to the mixture,
which then hybridize to the single-stranded DNA (Fig. 4.7). In the next step a DNA
polymerase extends the primers, doubling the copy number of the desired DNA frag-
ment. This concludes one PCR cycle. Each additional cycle (denaturation, annealing,
and amplification) doubles the existing amount of DNA that is located between the
primer pair. Thus, 30 cycles correspond to a 2
30
= ~10
9
-fold amplification step (30–40
cycles are typically used). A special heat-stable DNA polymerase is used that remains
124
4 Experimental Techniques in a Nutshell
active during the heating step and thus obviates the need to add fresh enzyme after
each cycle. Today, the different steps of the PCR reaction don’t have to be performed
manually. Small, automated PCR machines, also called thermal cyclers, can perform
dozens of PCR reactions in parallel, and the typical 30 cycles are finished in roughly
half an hour.
In the last few years, sequence information for many complete genomes has be-
come available, which allows one to use PCR to clone genes directly from genomic
DNA without the use of DNA libraries. PCR has revolutionized modern molecular
genetics and has many applications. For instance, by combining reverse transcrip-
tase (which makes a DNA copy from RNA) with PCR, it is also possible to clone
mRNA with this technique. The extreme sensitivity of PCR also makes it the method
of choice for forensic studies. Highly variable tandem repeats are amplified and used
to determine whether genetic material that comes from hair follicle cells, saliva, or
blood stains belongs to a certain suspect. This is possible because different indivi-
duals have tandem repeats of different length, which results in amplified DNA frag-
ments of different length. By looking at a large number of different tandem repeats,
the chances of a false positive result can be made arbitrarily small. This principle is
also the basis of paternity tests.
Another type of application is the use of multiple, or degenerate, primers. This
can be useful when a consensus sequence is known and the aim is to amplify genes
that contain this conserved sequence. Degenerate primers are also used if a stretch
125
4.2 Advanced Techniques
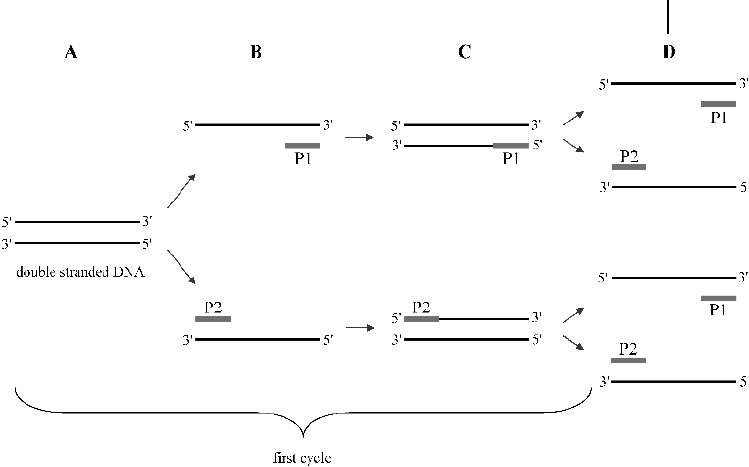
Fig. 4.7 The polymerase chain reaction. (a)
After double-stranded DNA is heated to obtain
single strands, (b) short DNA primers (P1 and
P2) are hybridized to a region that is upstream
or downstream, respectively, of the DNA of inter-
est. (c) A DNA polymerase is then used to ex-
tend the primers. (d) Then the cycle is repeated
by melting the DNA and annealing new primers.
For clarity the primers that were used in the first
cycle are not drawn in bold in the second cycle.

of the amino acid sequence of a new protein is known (e.g., through protein sequen-
cing) and the corresponding gene has to be found. Because of the degeneracy of the
genetic code, several nucleotide sequences can code for a given amino acid sequence.
The use of degenerate PCR can solve this problem.
An important, more recent, development is the technique of real-time PCR, which
is especially suited to quantify the amount of template that was initially present.
Classical PCR is normally unable to give quantitative results because of saturation
problems during the later cycles. Real-time PCR circumvents these problems by
using fluorescent dyes that either intercalate in double-stranded DNA or are bound
to sequence-specific oligonucleotides (TaqMan probe). The increase of fluorescence
with time is used, in real time, as an indicator of product generation during each
PCR cycle.
4.2.2
DNA and Protein Chips
4.2.2.1 DNA Chips
DNA chips, also called DNA microarrays, are a recently developed method for the
high-throughput analysis of gene expression (DeRisi et al. 1997). Instead of looking
at the expression of a single gene, microarrays allow one to monitor the expression
of several thousand genes in a single experiment, resulting in a global picture of the
cellular activity. This idea is also the basis of the systems biological modeling ap-
proach, and chip data can therefore be a very important source of data.
A microarray experiment starts with the construction of the chip (Fig. 4.8) from a
DNA library (see Section 4.1.2). The inserts of individual clones are amplified by
PCR (a single primer pair that is specific for the vector used to construct the library
can be used) and spotted in a regular pattern on a glass slide or nylon membrane.
These steps are normally automated and performed by robots. Then, total mRNA is
extracted from two samples that we would like to compare (e.g., yeast cells before
and after osmotic shock). Using reverse transcriptase, the mRNA is transcribed into
126
4 Experimental Techniques in a Nutshell
Fig. 4.8 DNA chips are used to study the ex-
pression level of thousands of genes in parallel.
Individual genes from a DNA library are ampli-
fied by PCR and spotted onto glass slides. cDNA
is prepared from the cells or tissues that are to
be compared and is labeled with red and green
fluorescent dyes. The color reveals, after hybridi-
zation, the relative mRNA amounts in the two
sources. Finally, the expression data are clus-
tered.

cDNA and labeled with a fluorescent dye. It is important that the dyes emit light at
different wavelengths; red and green dyes are commonly used. The cDNAs are now
incubated with the chip where they hybridize to the spot that contains the comple-
mentary DNA fragment. After washing, the ratio of the fluorescence intensities for
red and green are measured and displayed as false color pictures (Fig. 4.8, middle).
Red or green spots indicate a large excess of mRNA from one or the other sample,
while yellow spots show that the amount of this specific mRNA was roughly equal
in both samples. Very low amounts of both mRNA samples result in dark spots.
These ratios can, of course, also be quantified numerically and used for further cal-
culations, such as the generation of a clustergram. For this analysis, a complete link-
age cluster of the genes that were spotted on the chip is generated and the mRNA ra-
tio (represented as color) is displayed in this order (Fig. 4.8, right). This helps to test
whether groups of related genes (maybe all involved in fatty acid synthesis) also
show a similar expression pattern.
Oligonucleotide chips, a variant of DNA chips, are based on an alternative experi-
mental design. Instead of spotting cDNAs, short oligonucleotides (25–50mer) are
used. Approximately a dozen different and specific oligonucleotides are used per
gene. In this case only one probe of mRNA is hybridized per chip, and the ratio of
fluorescence intensity of different chips is used to estimate the relative abundance of
each mRNA. Chips from the companies Affymetrix or Agilent are most commonly
used for this approach.
During the past several years, this technique has been used to study such diverse
problems as the effects of caloric restriction and aging in mice (Lee et al. 1999), in-
fluence of environmental changes on yeast (Causton et al. 2001), and the conse-
quences of serum withdrawal on human fibroblasts (Eisen et al. 1998).
4.2.2.2 Protein Chips
Despite the large success of DNA chips, it is clear that the function of the genes is
realized through the proteins and not by the mRNAs. Therefore, efforts are under
way to construct chips that consist of spotted proteins instead of DNA. In this case
the starting point is an expression library for obtaining large quantities of the recom-
binant proteins. The proteins are spotted and fixed on a glass slide and can then be
incubated with interaction partners. These could be (1) other proteins to study pro-
tein complexes, (2) antibodies to quantify the spotted proteins or to identify the re-
cognized antigens, (3) DNA to find DNA-binding proteins, or (4) drugs to identify
compounds of pharmaceutical interest (Cahill and Nordhoff 2003).
However, the generation of protein chips poses more problems than DNA chips
because proteins are not as uniform as DNA. One challenge is to express sufficient
amounts of recombinant proteins in a high-throughput approach. Another problem
is that the optimal conditions (temperature, ionic strength, etc.) for interaction with
the reaction partner are not identical for different proteins. But the field is making
rapid progress, and protein chips will be used more often in the near future.
The main advantage of DNA and protein chips is the huge amount of data that
can be gathered in a single experiment. However, there are also points that need
careful consideration. The quality of an expression profile analysis based on array
127
4.2 Advanced Techniques

data is highly dependent on the number of repeated sample measurements and
on the array preparation, hybridization, and signal quantification procedure (Wier-
ling et al. 2002). The many samples on the chip also pose a problem regarding
multiple testing. If care is not taken, a large number of false positives is to be ex-
pected.
4.2.3
Yeast Two-hybrid System
The yeast two-hybrid (Y2H) system is a technique for the high-throughput detec-
tion of protein-protein interactions. It rests on the fact that some transcription fac-
tors (TF), such as the yeast Gal4 gene, have a modular design. The DNA-binding
domain is separated from the activating domain. To test whether two proteins (we
will call them bait and prey) interact, their genes are fused to the DNA-binding or
DNA-activating domain, respectively, of the TF (Fig. 4.9). The bait binds to the
DNA via its TF fragment. If bait and prey do interact, the two domains of the tran-
scription factor come close enough together to stimulate the expression of a repor-
ter gene. If bait and prey do not interact, the reporter gene is silent. This technique
can be used to find all interacting partners of the bait protein. For this purpose
yeast cells are transformed with an expression library containing prey proteins
fused to the activating part of the TF. Although the detection occurs in yeast, the
bait and prey proteins can come from any organism. This single-bait–multiple-prey
system can even be extended to a multiple-bait–multiple-prey approach, which
makes it possible to obtain the complete protein interactome of yeast (Uetz et al.
2000). However, as with most high-throughput techniques, the two-hybrid system
is prone to false positives, as indicated by the fact that a second study using the
same technique derived a quite dissimilar yeast interactome (Ito et al. 2001). To cor-
roborate the interactions obtained with Y2H, affinity chromatography or co-immu-
noprecipitation can be used.
128
4 Experimental Techniques in a Nutshell
Fig. 4.9 The yeast two-hybrid system identifies protein-protein in-
teractions. The genes of the bait and prey proteins are fused to parts
of a yeast transcription factor. If bait and prey interact, the different
parts of the transcription factor come close enough together to acti-
vate the expression of a reporter gene.

4.2.4
Mass Spectrometry
The identification of individual proteins of a cell is an essential part for studying bio-
logical processes. The first step towards identification often involves a separation of
the protein broth. 2D gels (see Section 4.1.3) or the different forms of column chro-
matography (see Section 4.1.5.2) are frequently the method of choice. The separated
proteins can then be identified by cleaving them enzymatically or chemically into
specific peptides and determining the exact size of these fragments. The resulting
size distribution, the fingerprint of the protein, is compared to the theoretical finger-
prints that are calculated for all proteins in the sequence databanks. This is possible
because it is known at which amino acid sequence proteases like trypsin cut a pro-
tein.
Mass spectrometry (MS) has been used for the past 50 years to measure the
masses of small molecules with high accuracy. However, its application to large bio-
molecules has been limited by the fragility and low volatility of these materials. But
these problems have been solved with the development of the matrix-assisted laser
desorption/ionization time-of-flight (MALDI-TOF) technique. The polypeptide is
mixed with a solvent and an excess of the matrix material and is placed inside the
mass spectrometer, where the solvent evaporates. The probe is then targeted with a
laser beam that transfers most of its energy to the matrix material, which heats up
and is vaporized. During this process the intact polymer is charged and carried into
the vapor phase. An electrical field accelerates the molecules, which then traverse
down an evacuated tube, (smaller ions arrive in a shorter time at the detector than
massive ions), and the time of flight is recorded. The masses of the different ions
can then be calculated, taking into account that the TOF is proportional to the square
root of the mass divided by the charge of the ion (TOF
m=z
p
).
The accuracy of MALDI-TOF is extremely high. The error is less than 1 part in
10
5
. That means the error in determining the mass of a 100-kDa protein is less than
1 Dalton! Because of this very high accuracy, MS can also be used for sequencing of
peptide fragments. In this case two mass spectrometers have to be used in succes-
sion. The first one separates the peptides (according to their time of flight) and feeds
them individually into the second MS, where they are further fragmented at their
peptide bonds. The output of the second spectrometer is therefore mass peaks that
are separated by the mass of a single amino acid, which can be used to construct the
sequence of the peptide. With the use of degenerate primers (see Section 4.2.1), the
gene for the protein can then be cloned. This is useful if the fingerprint of the pro-
tein is not found in the database and the protein cannot be identified this way.
Finally, MS-based peptide sequencing also yields information about posttransla-
tional modifications. If an amino acid is chemically modified (i. e., phosphorylated
or glycosylated), this results in a characteristic change of the mass and thus it can be
identified.
129
4.2 Advanced Techniques
