Klipp E., Herwig R., Kowald A., Wierling C., Lehrach H. Systems Biology in Practice: Concepts, Implementation and Application
Подождите немного. Документ загружается.


8.3.1.3 Bayesian Networks
A Bayesian network (see also Section 3.5.2.3) is based on the representation of the
regulatory network as a directed acyclic graph G = [V, E]. Again, the vertices i BV re-
present genes and edges denote regulatory interactions. Variables x
i
belonging to the
vertices i denote a property relevant to the regulation, e.g., the expression level of a
gene or the amount of active protein. A conditional probability distribution
p(x
i
|L(x
i
)) is defined for each x
i
, where L(x
i
) are the parent variables belonging to
the direct regulators of i. The directed graph G and the conditional distributions to-
gether specify a joint probability distribution p(x) that determines the Bayesian net-
work. The joint probability distribution can be decomposed into
px
Q
i
px
i
Lx
i
: (8-6)
The directed graph expresses dependencies of probabilities: the expression level of
a gene represented by a child vertex depends on the expression levels of genes be-
longing to the parent vertices. Hence, it also implies conditional independencies
i(x
i
; y|z), meaning that x
i
is independent of the set of variables y given the set of
variables z . Two graphs or Bayesian networks are equivalent if they imply the same
set of independencies. In this case they can be considered as the same undirected
graph, but with varying direction of edges. Equivalent graphs cannot be distin-
guished by observation of the variables x (Friedman et al. 2000).
Example 8-2
For the network given in Fig. 8.2c the conditional independence relations are
i(x
a
;x
b
) and ix
d
;x
a
; x
b
x
c
j
. The joint probability distribution of the network is
px
a
; x
b
; x
c
; x
d
px
a
px
b
px
c
x
a
; x
b
j
px
d
x
c
j
.
Bayesian networks have been used to deduce gene regulatory networks from gene
expression data. The aim is to find the network or equivalence class of networks that
best explains the measured data. A problem is the determination of initial probability
distributions.
8.3.1.4 Boolean Networks
In the Boolean network approach (see also Section 3.5.2.2 and Section 10.3.3 for
Boolean rules), the expression level of each gene is assigned to a binary variable: a
gene is considered to be either on (1) or off (0), i.e., it is transcribed or not. The
states of the genes are updated simultaneously in discrete time steps. The new state
can depend on the previous state of the same gene or other genes. These dependen-
cies cause the Boolean network. The following termini are used: the N genes are the
N nodes of the network, the k interactions regulating the expression of a certain
gene are the k inputs of that node, and the binary expression value of each gene is its
output. Since every node can be in one of two different states, a network of N genes
can assume 2
N
different states. An N-dimensional vector of variables can describe
270
8 Modeling of Gene Expression

the state at time t. The value of each variable at time t+ 1 depends on the values of its
inputs. It can be computed by means of the Boolean rules (see Section 10.3.3). For a
node with k inputs, the number of possible Boolean rules is 2
2
k
. Although a Boolean
network is a very simplified representation of the gene regulatory network, it enables
a first computation of gene expression dynamics.
Example 8-3
For the network presented in Fig. 8.2d the following Boolean rules apply:
at 1f
a
atat Rule 1 for k 1
bt 1f
b
ct; dtnot ctand dt Rule 2 for k 2
ct 1f
c
at; btatand bt Rule 2 for k 2
dt 1f
d
ctnot ct Rule 0 for k 1
The temporal behavior is determined by the sequence of states (a, b, c, d) given an
initial state (compare also Section 10.3.3).
From Tab. 8.1 it is easy to see that this network has two different types of stationary
behavior. If the initial state of a is 0, then the system evolves towards the steady
state 0101, meaning that genes a and c are off,while genes b and d are on. If the in-
itial state of a is 1, then the system evolves towards a cyclic behavior including the
following sequence of states: 1000 ? 1001 ? 1101 ? 1111 ? 1010 ? 1000.
The sequence of states given by the Boolean transitions represents the trajectory
of the system. Since the number of states in the state space is finite, the number of
possible transitions is also finite. Therefore, each trajectory will lead either to a
steady state or to a state cycle. These states are called attractors. Transient states are
those states that do not belong to an attractor. All states that lead to the same attrac-
tor constitute the basin of attraction.
Boolean networks have been used to explore general and global properties of large
gene expression networks. Considering random networks (the number k of inputs
per gene and the corresponding Boolean rules are chosen by chance), Kauffman
(1991, 1993) has shown that the systems exhibit highly ordered dynamics for small k
271
8.3 Modeling Specific Processes in Eukaryotic Gene Expression
Tab. 8.1 Successive states in the Boolean network.
0000 ? 0001 1000 ? 1001
0001 ? 0101 1001 ? 1101
0010 ? 0000 1010 ? 1000
0011 ? 0000 1011 ? 1000
0100 ? 0001 1100 ? 1011
0101 ? 0101 1101 ? 1111
0110 ? 0000 1110 ? 1010
0111 ? 0000 1111 ? 1010

and certain choices of rules. The expectation value of the median number of attrac-
tors is about
N
p
, and the length of attractors is restricted to a value proportional to
N
p
. Kauffman suggested the interpretation of the number of possible attractors as
the number of possible cell types arising from the same gene type.
Example 8-4: Reverse engineering
Reverse engineering of Boolean networks aims to derive the Boolean interaction
rules from time-dependent gene expression data. In Section 9.6 we discuss the
REVEAL algorithm (Liang et al. 1999) that addresses this problem. We applied
this algorithm to the model described in Fig. 8.2d. Expression data were gener-
ated using the ODE model of Example 8-1. The model constructed by REVEAL
entails most of the rules with two steps (k = 1 and k = 2). Binarization of the time
course concentration data yielded 54 different state transitions, out of which 21
corresponded to correct transitions. Pre-processing the state transitions after fre-
quency of occurrence yielded a confidence value for each transition. If only the
transitions with the highest confidence are considered, we end up with seven
transitions, out of which six were correct:
The rules derived from this table are
at 1f
a
atat
bt 1
~
f
b
ct; dtctand dtor not ctand dt
ct 1
f
c
at
; bt
at
and bt
dt 1f
d
ctnot ct
The second rule,
~
f
b
, is partly wrong, whereas the others hold. The network was re-
constructed within two steps of the algorithm.
The example illustrates several practical problems of reverse engineering Boolean
networks:
1. Binarization is difficult and strongly influences the result. It is not quite clear
from expression data what the level for binarization should be. Furthermore,
the level should be determined gene-wise in order to distinguish low abundant
from high abundant genes.
272
8 Modeling of Gene Expression
State transition t t + 1 Confidence Validity
1 0000 0001 1 True
2 1101 1111 0.5 True
3 1111 1110 0.625 False
4 1110 1010 0.5 True
5 1010 1000 0.3 True
6 1000 1001 1 True
7 1001 1101 0.5 True

2. States are incomplete. In practice, most of the state transitions are missing
after binarization. In our example, only seven out of 16 state transitions occur.
3. Availability of many time points is crucial. In order to filter correct states from
false states, we have to screen many state transitions to get a stable result. In
our example, we used 100 time points to reconstruct the network.
4. Time points should not be too close to each other. The selection of time points
determines the granularity of the set of state transitions. It is a tradeoff be-
tween the detection of as many state transitions as possible and avoiding false-
positive transitions. If two time points are too close together, the transition
shows no changes since the binarization is a very rough threshold that does
not detect small concentration changes. This will lead to many false-positive
self-loops in the corresponding graph.
8.3.1.5 Gene Expression Modeling with Stochastic Equations
For gene expression processes, it can be argued that the assumptions of continuous
and deterministic concentration changes employed in the description with ODEs are
not valid. Each gene is present in only one, two, or a few copies. The number of tran-
scription factor molecules is usually small (about a dozen or a few hundred), and
even the abundance of mRNA molecules is often at the detection limit. Therefore, it
is not certain whether the actors of the considered processes are actually present, and
the character of the events becomes probabilistic. Furthermore, the involved pro-
cesses can hardly be considered as continuous. For example, in transcription it takes
a certain amount of time from the initiation until termination. The discrete and prob-
abilistic character of processes is taken into account in stochastic modeling of gene
regulation. In this approach the state variables are discrete molecule numbers x. The
quantity p(x, t) expresses the probability that there are x molecules of type x at time t.
Consideration of the probability for all possible values of x = 0, 1, 2, … yields the prob-
ability distribution. Time evolution is given by the relation
px; t Dtfpx; t
; (8-7)
where the function f comprises all processes increasing or decreasing x occurring
during the period Dt. Note that only simple reaction steps (formation of a molecule,
complex formation, degradation) but no composed processes (like in Michaelis-Men-
ten kinetics) can be regarded. The period Dt must be chosen so small that only one
reaction step may occur in Dt. Equation (8-7) gives the probability of the system
being in a certain state (having a certain number of molecules of a species) at a cer-
tain time. The simulation yields one realization of the system behavior, while an-
other simulation run may yield another realization. Equation (8-7) is discrete in va-
lues of the state variables and in time. Taking the limit Dt ?0 leads to the master
equation that is discrete in state values but continuous in time:
i
it
px; tfpx; t
: (8-8)
273
8.3 Modeling Specific Processes in Eukaryotic Gene Expression

This equation gives the probability of the system being in a certain state at a
certain time. This equation is usually hard to solve, both analytically and numeri-
cally. The Gillespie algorithm is a convenient alternative approach that is mathe-
matically equivalent and employs repeated individual stochastic simulations (Gil-
lespie 1977).
Example 8-5
The equation governing the behavior of mRNA a in the gene regulatory network
shown in Fig. 8.2 is
pa; t Dtk
a1
pa1; tk
a2
pa; tk
a3
pa; tk
a4
pa1; t: (8-9)
The term k
a1
7p(a –1,t) expresses the probability that the number of molecules
was a – 1 at time t and that one molecule has been produced in Dt (due to tran-
scription); k
a2
7p(a, t) denotes the probability that the number a did not change
at all during Dt; and the term k
a3
7p(a, t) stands for the probability that the num-
ber of molecules was a at time t and that one molecule was degraded. Finally,
k
a4
7p(a +1,t) denotes the decrease of the molecule number from a +1toa in
period Dt due to degradation of one molecule.
The advantage of stochastic modeling is the explicit consideration of uncertainties
due to the stochastic and discrete character of processes involving low molecule
numbers. In some cases, experimentally observed behavior (switch between differ-
ent states) could be explained with fluctuations that are inherent to stochastic model-
ing but not to differential equations. A problem common to stochastic equations and
ODEs is the limited knowledge about appropriate kinetic parameters. Furthermore,
the simulation of large systems demands computational power, especially for higher
molecule numbers. Therefore, algorithms are developed to combine stochastic simu-
lation for low-abundance species with deterministic simulations for high-concentra-
tion species.
8.3.2
Time Delay in Gene Regulation
An important issue in modeling gene expression is the fact that individual processes
need a certain amount of time to be finished. For example, mature mRNA is not im-
mediately available (not even in very small amounts) shortly after initiation of tran-
scription. Different approaches have been employed to handle this problem, namely,
the explicit consideration of numerous intermediate reaction steps or the incorpora-
tion of a discrete time delay.
Studying a system where a transcription factor, TF-A, activates its own transcrip-
tion, Smolen and colleagues (1998, 1999) gave an example of a model with discrete
time delay. For the mechanism (see Fig. 8.4), they consider that TF-A forms a phos-
phorylated homodimer that activates transcription by binding to an enhancer
274
8 Modeling of Gene Expression
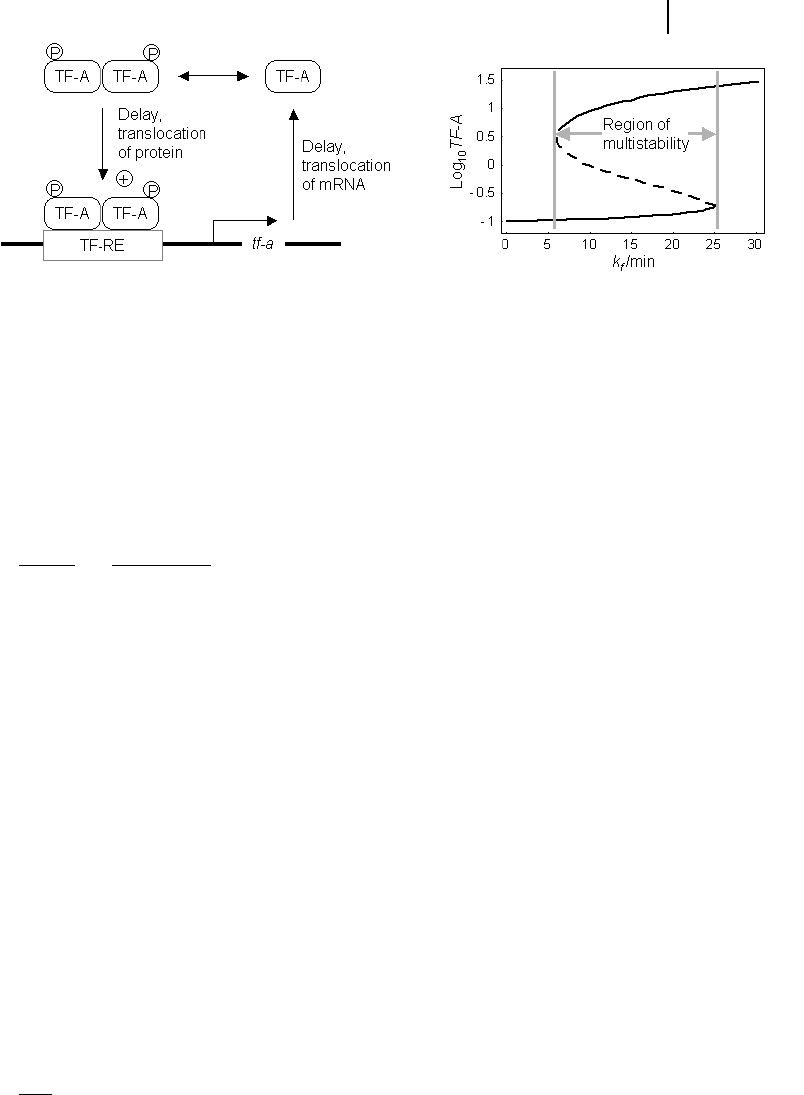
(TF-RE). Monomeric and dimeric TF-A are in rapid equilibrium. The transcription is
described with saturation kinetics dependent on the concentration of dimeric TF-A
with maximal rate k
f
and dissociation constant K
D
. Degradation of TF-A is modeled
by applying mass action kinetics with rate constant k
d
, and the basal production of
TF-A has the rate R
bas
.
Using these assumptions, the temporal behavior of TF-A is described by a single
delay differential equation
dTF-A
dt
k
f
TF-A
2
TF-A
2
K
D
()
t tk
d
TF-A R
bas
: (8-10)
The steady-state solution of Eq. (8-10) shows a bistable behavior for a range of
parameter values. There is one solution with low concentration of TF-A and a synth-
esis rate close to R
bas
and one solution with high concentration, as shown in Fig. 8.4.
Perturbations, exerted by transient changes in k
f
, can lead to a switch between both
states. The natural counterpart of these perturbations is changes in kinase or phos-
phatase activities caused by external signals.
An example for modeling time delay with several reaction steps is the model of
GATA-3 transcriptional imprinting in Th2 lymphocytes by Höfer et al. (2002). The
model describes the regulation of GATA-3 activity on the transcriptional and post-
translational level. Transcription is enhanced by two transcription factors, Stat6 and
NF-kB, and by autoactivation. Post-translational regulation involves phosphorylation,
acetylation, and interaction with inhibitory proteins.
Starting with the synthesis of the primary transcript R
1
, a series of conversions, in-
cluding splicing and nuclear export of mRNA, produces the intermediary mRNA
forms R
i
(i =2,…,m – 1) and, eventually, the functional mRNA R
m
, which is asso-
ciated to ribosomes. The translated GATA-3 polypeptide chain, G
1
, is also stepwise
modified (intermediates G
j
) to yield the active transcription factor G
n
. The respective
ODE system reads
dR
1
dt
vG
n
; tk
1
l
1
R
1
275
8.3 Modeling Specific Processes in Eukaryotic Gene Expression
Fig. 8.4 TF-A dynamics with discrete time delay.

dR
i
dt
k
i1
R
i1
k
i
l
i
R
i
i 2; :::; m
dG
1
dt
k
t
R
m
k
0
1
l
0
1
G
1
dG
j
dt
k
0
j1
G
j1
k
0
j
l
0
j
G
j
j 2; :::; m : (8-11)
The k' s and l's are first-order rate constants of conversion reactions and loss reac-
tions, respectively. The function vG
n
; t
k
B
k
S
e t
k
G
G
2
n
1 G
n
2
contains the
term e t
0; t < 0
e
t=T
; t 0
, expressing exponential decay of an external signal after
time t = 0. The intention of the model was to demonstrate the coexistence of two ex-
pression states of GATA-3 depending on autoactivation. But it can also be used to de-
monstrate time delay. Figure 8.5 shows the effect of changing the number of inter-
mediary mRNA species on the appearance of active GATA-3.
8.3.3
Modeling the Elongation of a Peptide Chain
The speed of translation as the second part of the gene expression process depends
on the velocity of the elongation process. This, in turn, is dependent on the rate of
the individual steps during growth of the nascent peptide chain and on the availabil-
ity of the different types of tRNA bound amino acids.
A very general model accounting for the dynamics of protein expression in prokar-
yotes is given by (Drew 2001)
dmRNA
dt
k
R
R mRNA k
N1
mRNA
N1
dmRNA
0
dt
k
1
a
1
mRNA
0
k
R
R mRNA
dmRNA
j
dt
k
j1
a
j1
mRNA
j
k
R
a
j
mRNA
j1
; j 1::N (8-12)
276
8 Modeling of Gene Expression
Fig. 8.5 Temporal behavior of mRNA species
R
1
and active transcription factor G
n
for differ-
ent numbers of intermediary steps in trans-
cription according to Eq. (8-11) Solid lines:
three intermediary steps (m = 3), dashed lines:
seven intermediary steps (m = 7). A higher
number of intermediary steps leads to a later
onset in the formation of transcription factor
GATA-3. Parameters: k
B
= 0.001, k
G
=5,
k
S
= 0.5, T = 15, k
i,j
= 1 (all i, j), l
i,j
= 0 (all i, j),
except of l
m,n
=1,n =3.
where mRNA denotes the concentration of the messenger RNA, mRNA
0
is the con-
centration of mRNA-ribosome complex, and mRNA
j
is the concentration of mRNA-
ribosome complex with a nascent peptide chain of length j attached. k
R
is the rate
constant for the mRNA-ribosome complex formation process and k
j
with j = 1..N are
the rate constants for the elongation steps.
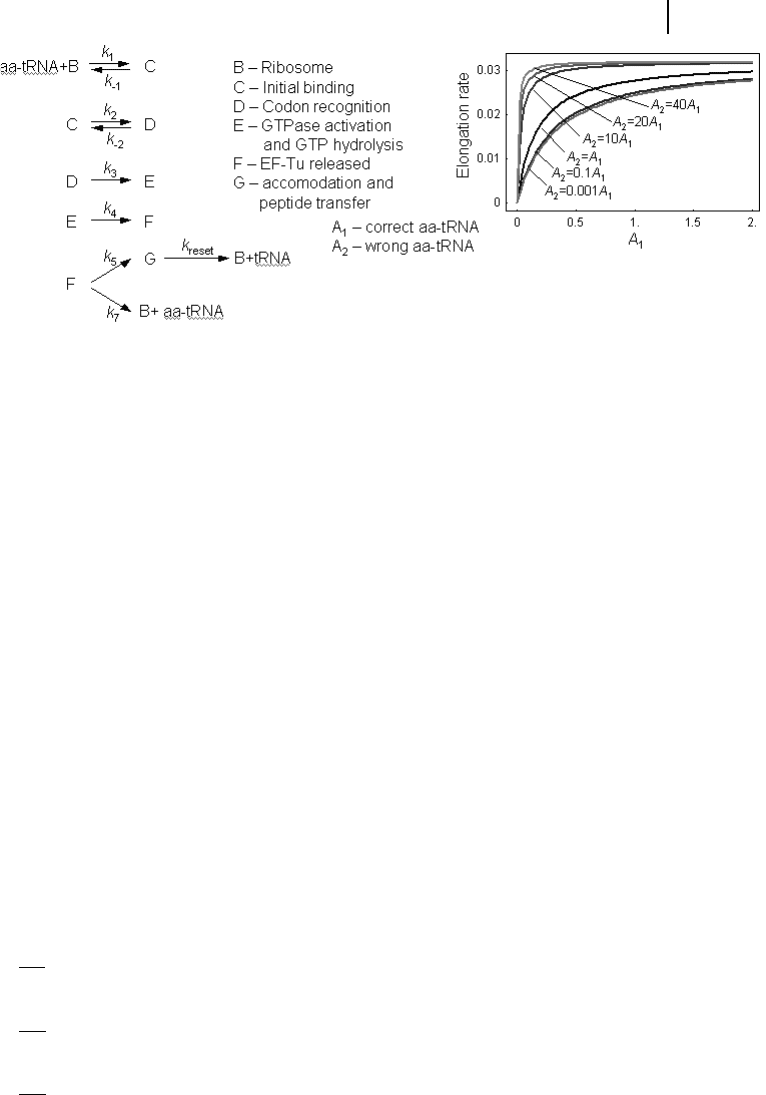
The elongation step occurring at the ribosome can also be considered in more de-
tail. In Fig. 8.6, the sequence of events during each elongation step is presented:
(1) the binding of the tRNA, which is bound to an amino acid (aa) and in complex
with the elongation factor Tu, to the ribosome; (2) recognition of the codon; (3) acti-
vation of a GTPase and GTP hydrolysis; (4) conformational change and release of
the elongation factor Tu together with a proofreading for the correct amino acid;
(5) either rejection of the wrong tRNA or accommodation of the correct tRNA; (6) for-
mation of the peptide bond and resetting of the ribosome into a form ready for the
next elongation step; (7) even the correct form of tRNA does not lead to the forma-
tion of a peptide bond in all cases. The kinetics of this mechanism depends partly on
the nature of the tRNA. The process proceeds faster if the tRNA is the correct one
that provides the amino acid to be next in the peptide sequence. The process shown
in Fig. 8.6 can be described with a set of differential equations. Denoting the correct
tRNA with A
1
and the wrong tRNA with A
2
leads to
dB
dt
k
1
A
i
B k
1
C k
r
G k
7
F i 1; 2
dC
dt
k
1
A
i
B k
1
C k
2
C k
2
D
dD
dt
k
2
C k
2
D k
3
D
277
8.3 Modeling Specific Processes in Eukaryotic Gene Expression
Fig. 8.6 Model for the elongation of a peptide chain according to
Heyd and Drew. Left: Reaction steps involved in elongation.
Right: Elongation rate for different ratios correct tRNA A
1
and wrong
tRNA A
2
.

dE
dt
k
3
D k
4
E
dF
dt
k
4
E k
5
F k
7
F
dG
dt
k
5
F k
r
G : (8-13)
This model is suited to study the temporal behavior of peptide chain elongation. It
can also be used to study the effect of tRNA supply. Figure 8.6 shows the dependence
of the elongation rate from the ratio of correct and wrong tRNA. As can be expected,
the lower the ratio of wrong tRNA, the higher the elongation rate is. If no correct
tRNA is present, then no elongation occurs. With high concentrations of correct
tRNA, the wrong tRNA is out-competed. Heyd and Drew (2003) studied a similar
system with a stochastic approach, obtaining similar results.
8.4
Modeling the Regulation of Operons in E. coli
In this section we will discus the regulation of gene expression in prokaryotes in con-
text of the operon concept. The operon concept suggests that genes are controlled by
means of operons through a single feedback regulatory mechanism known as repres-
sion. An operon consists of a set of genes preceded by a small DNA segment (the op-
erator), where repression takes place and mRNA-polymerase binds to initiate tran-
scription. An operon is repressed when an active repressor molecule binds the opera-
tor, blocking it and preventing the binding of mRNA-polymerase. The term operon
was introduced by Jacob et al. (1960) and had a deep impact on the biological sciences.
Shortly after the implementation of the operon concept, Goodwin (1965) gave the first
mathematical analysis of operon dynamics. Griffith performed a more complete ana-
lysis of simple repressible (negative feedback [Griffith 1968a]) and inducible (positive
feedback [Griffith 1968b]) gene control networks. During the last four decades, mod-
els including more and more details of the dynamic behavior of operons have been
presented along with various experimental data (Nicolis and Prigogine 1977; Santillan
and Mackey 2001; Yildirim and Mackey 2003; Mackey et al. 2004).
8.4.1
Mechanism of the Lac Operon in E. coli
E. coli prefers glucose as energy source. Under all conditions it contains the enzymes
necessary for the metabolism of glucose. If the cells are short of glucose they are
able to utilize other sugars, e.g., lactose. However, cells grown on glucose-rich med-
ium are unable to metabolize lactose immediately when they are exposed to a lac-
tose-rich medium. Only after a certain lag time they can assimilate this sugar at a
high rate. E. coli can sense the presence and concentrations of its nutrients. The con-
278
8 Modeling of Gene Expression
centration of cyclic AMP (cAMP) decreases with increasing concentration of glucose,
and a lack of glucose induces a rise in cAMP. Lactose is sensed in form of allolactose,
an isomer formed by a reaction converting the 1–4 bond of lactose into a 1–6 bond.
The adequate response of cells on the availability of nutrients is due to regulatory
processes in the cell, i.e., is due to the control of the synthesis of proteins. The chro-
mosome of E. coli consists of only one circular DNA molecule containing genes for
about 4000 proteins. The expression of many of these proteins is regulated depend-
ing on the intracellular concentration of certain metabolites. The lac operon is a tran-
scription unit that ensures that the enzymes for the lactose metabolism are ex-
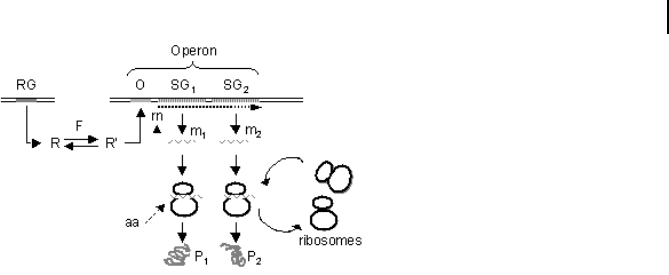
pressed only if lactose is in the medium and glucose is missing. The lac operon (see
Fig. 8.7) contains a gene for a repressor protein R. In its active form, R binds to the
operator, O, a specific DNA sequence of 21 base pairs. The operator sequence over-
laps with the RNA polymerase–binding region, the promoter, of the following struc-
tural genes. These genes code for b-galactosidase (E), permease (M), and thiogalacto-
side transacetylase.
Jacob and Monod (Jacob et al. 1960) formulated a set of general assumptions for
their model:
1. The primary product of structural genes is the messenger RNA. It is short-lived and
brings information to the ribosomes. The second transcription takes place at the ribo-
somes, polypeptides are formed, and mRNA is destroyed. Ribosomes are reused.
2. mRNA synthesis is a sequential, oriented process that can start only on specific re-
gions of the DNA, the operators. One operator may control the transcription of
several subsequent structural genes, together denoted as operon or a unit of pri-
mary transcription.
3. Besides structural genes, there are also regulatory genes that code for a repressor.
A repressor binds reversibly to a specific operator such that transcription initiation
is blocked and protein synthesis is prevented.
4. The repressor R can specifically interact with small molecules, i. e., with the effec-
tors F that change its activity.
R F $ R
0
F
0
: (8-14)
279
8.4 Modeling the Regulation of Operons in E. coli
Fig. 8.7 The operon model of Jacob
and Monod: the operon comprises the
operator O and structural genes SG
1
and SG
2
. The structural genes code for
the mRNAs m1 and m2, which in turn
are translated to the proteins P
1
and P
2
.
A regulatory gene, RG, provides the reg-
ulator R. The effector F catalyzes the
transition of the regulator from the ac-
tive to the inactive form (R and R').
R binds to the operator and prevents
binding of the RNA polymerase. If the
repressor is inactive, then transcription
of the structural genes can occur.
