Klipp E., Herwig R., Kowald A., Wierling C., Lehrach H. Systems Biology in Practice: Concepts, Implementation and Application
Подождите немного. Документ загружается.


Consider now more generally np-dimensional data vectors x
1
,…,x
n
with compo-
nent-wise mean vector
x. PCA is computed with a decomposition of the pxp-dimen-
sional empirical covariance matrix of the sample (compare Eq. (3-64) in Chapter 3):
S
1
n 1
X
n
i1
x
i
xx
i
x
T
: (9-20)
Since the matrix S is symmetric and positive semi-definite, there exist p nonnega-
tive eigenvalues l
1
::: l
p
0 (which we may assume to numerate in decreasing
order, cf. Section 3.1.2). Let r
1
,…,r
p
be the corresponding eigenvectors, such that
r
T
j
r
k
1; if j k
0; if j 6 k
Sr
j
l
j
r
j
: (9-21)
If we denote with R the pxp-dimensional matrix whose columns are composed of
the p eigenvectors from S, and if K
l
1
00
0
.
.
.
0
00l
p
0
B
@
1
C
A
denotes the pxp-dimensional
diagonal matrix whose diagonal elements are the p eigenvalues, we get the decompo-
sition
S RKR
T
: (9-22)
Geometrically, the eigenvectors of S are the main axes of dispersion of the dataset
{x
1
,…,x
n
}. The dispersion is maximally high in the first principal component, sec-
ond highest with the second principal component, etc. The dispersion in each princi-
pal component, i, equals
l
i
p
. Suppose now that an eigenvalue l
k
is close to zero.
This means that there is not much variance along that principal component at all
and that the kth coordinate of the vectors x
i
is close to zero in the transformed coor-
dinate system. Thus, this dimension does not contribute very much to the overall
dispersion of the data and can be neglected without essential loss of information. Si-
milarly, this holds for j = k + 1, …, p, since we assumed the eigenvalues to be sorted
in decreasing order. Thus, we have replaced the original p dimensions with k –1<p
dimensions that explain the relevant variance of the data sample.
An important question is how many principal components are needed to explain
a sufficient amount of the data variance. Denote each vector by its coordinates
x
i
=(x
i1
,…,x
ip
)
T
and let
x
j
be the jth coordinate of the mean vector
x. A suitable mea-
sure for the total variance of the sample is the sum of the variances of the p coordi-
nates given by
X
p
j1
1
n 1
X
n
i1
x
ij
x
j
2
Trace S
X
p
j1
l
j
: (9-23)
320
9 Analysis of Gene Expression Data

Thus, for each k < p the relative amount of variance explained by the first k princi-
pal components is given by
P
k
j1
l
k
P
p
j1
l
p
: (9-24)
In gene expression analysis, PCA is widely used to reduce the gene expression ma-
trix or its transpose matrix, the condition expression matrix, to two or three reduc-
tions. Equation (9-24) is widely used in practice to characterize the computed dimen-
sion reduction. If the amount of variance explained is high, then such a reduction
makes some sense; otherwise, the dataset is too complex to visualize with PCA.
Example 9.4: PCA examples
Figure 9.8 b shows a display after PCA for eight different bovine tissues that were
screened with a cDNA array. Six hundred cDNAs were filtered from the total set
of 20,000 cDNAs in order to represent the tissues in a suitable way. Criteria were
fold changes between tissues and reliable expression differences. PCA shows that
the brain regions (cortex, cerebellum) form a tissue cluster separated from the
others. The selected first two principal components explain approximately 82% of
the data variance, indicating that the visualization is meaningful.
9.4.3
Functional Categorization
The guilt-by-association concept so far has not been proven to hold generally. How-
ever, in many studies functional classification of specific clusters has been shown to
be successful. Since more and more functional annotations are available for genes,
e.g., through the Gene Ontology Consortium (2003), these annotations are used to
validate clustering results. Here, we can use a simple statistical method to assign
each cluster a significance P-value that judges whether it contains some untypical ag-
glomeration of functional information (compare Section 13.1).
Consider a cluster of m genes, out of which k genes belong to a certain functional
class and m–kgenes belong to different classes. Let n be the total number of genes
under analysis and K be the total number of genes annotated for that class. Is the ob-
served number untypical, or does it rather express a random distribution of the spe-
cific functional class? This problem can be translated into an urn model (cf. Example
3.15). If the m genes were randomly drawn from the total of n genes, then the prob-
ability of having exactly k out of K genes from the functional class would be given by
the hypergeometric distribution
321
9.4 Validation of Gene Expression Data

P k
K
k
n K
m k
n
m
: (9-25)
The P-value for the cluster can then be calculated as the probability of having
more than the observed number of hits of that functional group using Eq. (9-25),
i.e., p
P
jk
P j.
9.5
Classification Methods
In this section we will introduce the basic principles of classification methods for mi-
croarray data. In Section 9.5.1 we discuss the basic concepts, in Section 9.5.2 we de-
scribe the basic idea of support vector machines as one of the most important classi-
fication methods, and in Section 9.5.3 we list some alternatives. Section 9.5.4 dis-
cusses cross-validation as a method to validate the performance of classification algo-
rithms.
9.5.1
Basic Concepts
An important medical application of microarray analysis is the diagnosis of diseases
and subtypes of a disease, e.g., cancer. Normal cells can evolve into malignant cancer
cells by mutations of genes that control cell cycle, apoptosis, and other processes
(Hanahan and Weinberg 2000). Determination of the exact cancer type and stage is
essential for the correct medical treatment of the patient. The task of sample diag-
nostics cannot be undertaken by the methods discussed so far. This task defines a
complementary set of mathematical algorithms for gene expression – classification
procedures. Recall that the purpose of clustering is to partition genes (and possibly
conditions) into co-expression groups by a suitable optimization method based on
the expression matrix (Section 9.3). The purpose of classification is to assign a given
condition (e.g., a patient’s expression profile across a set of genes) to preexisting
classes of conditions (e.g., groups of patient samples from known disease stages).
The clustering methods discussed so far do not utilize any supporting tissue annota-
tion (e. g., tumor vs. normal). This information is used only to assess the perfor-
mance of the method. Such methods are often referred to as unsupervised. In con-
trast, supervised methods attempt to predict the classification of new tissues based on
their gene expression profiles after training on examples that have been classified by
an external “supervisor.”
The practical problems underlying the classification of patients to disease sub-
types are that (1) new/unknown disease classes have to be identified, (2) marker
genes that separate the disease classes have to be found, and (3) patients have to be
322
9 Analysis of Gene Expression Data

classified by assigning them to one of the classes. The first problem has already
been discussed in Section 9.3. Clustering methods can be used to group the samples
and identify subtypes of disease conditions. The identification of marker genes can
be carried out using statistical tests (compare Section 9.2) or by the selection of
groups of co-regulated genes that are significantly different across the disease sub-
types. This program has been published previously in several large-scale studies. For
example, Bittner et al. (2000) studied malignant melanoma and identified two sub-
types of cutaneous melanoma with different characteristics in cell motility and inva-
siveness. They discovered a subset of melanomas identified by mathematical cluster-
ing of gene expression in a series of samples. Many genes underlying the classifica-
tion of this subset are differentially regulated in invasive melanomas that form pri-
mitive tubular networks in vitro, a feature of some highly aggressive metastatic mela-
nomas.
Golub et al. (1999) have described a generic approach to cancer classification
based on gene expression profiling and applied this to human acute leukemias as a
test case. A class-discovery procedure automatically discovered the distinction be-
tween acute myeloid leukemia (AML) and acute lymphoblastic leukemia (ALL) with-
out previous knowledge of these classes. On the basis of 50 pre-selected “informa-
tive” genes (from a total of 6817 genes), they assigned 36 out 38 patients correctly to
the two cancer classes.
Once the set of marker genes and the disease classes have been determined, a
new patient sample can be classified according to the expression profile among the
marker genes and the comparison with the training set. We focus here on the sim-
plest classification task, i.e., the assignment to only two different classes (normal vs.
disease). Here, the general classification problem can be stated as follows. Let T be a
set of n training samples consisting of pairs x
i
; z
i
, T x
1
; z
1
; ...; x
n
; z
n
,
where x
i
is a p-dimensional vector and z
i
21; 1fgis a binary label (class label).
Each vector consists of the expression profile of the patient sample across the p mar-
ker genes and each label assigns this vector to one of the classes. Given a new query,
x 2<
p
, the classification method (classifier) has to predict the group label, z,ofx
given the training set. Thus, each classification method can be interpreted as a func-
tion F:<
p
T !1; 1fg.
9.5.2
Support Vector Machines
Support vector machines (SVMs) are the most widely used group of methods for
classification (Vapnik 1995). Several studies have been published using SVMs in re-
cent years, in particular for cancer diagnostics (e.g., Furey et al. 2000; Statnikov et al.
2004). A recent implementation can be found in Chang and Lin (2003).
The intuition of support vector machines is that of a linear decision rule. Consider
two different groups of vectors in <
p
. We want to find a hyperplane that separates
these two samples by making the least possible error. The usual problem is that there
are many such separating hyperplanes so that we have to define some kind of optimi-
zation criterion. Figure 9.9 illustrates the problem with the simple case of a two-di-
323
9.5 Classification Methods
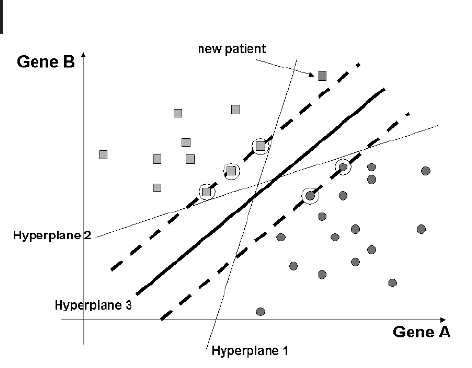
mensional space spanned by the expression levels of the patients according to two
marker genes (A and B). Hyperplanes 1 and 2 both separate the two samples perfectly.
However, given a new patient profile (red square), both methods would lead to differ-
ent classification results. The problem here is that both hyperplanes are geometrically
too close to one of the samples and thus risk misassigning a future datum.
The idea behind SVMs is to select a hyperplane that is more likely to generalize
on future data. This is achieved by finding a hyperplane that maximizes the mini-
mum distances of the closest points and thus maximizes the width of the margin be-
tween the two classes. The hyperplane is specified by the boundary training vectors
(support vectors).
Recall the classification problem from the previous section. A hyperplane can be
described by H w; b
x; wx b 0
fg
, with a vector w 2<
p
that determines the or-
ientation of the hyperplane and a scalar b that determines the offset of the hyper-
plane from the origin. Here, wx denotes the inner or dot product of the two vectors.
A hyperplane in two dimensions is given by a straight line (Fig. 9.9) and in three di-
mensions by a plane. We say that a hyperplane supports a class if all points in that
class fall on one side. Thus, we would like to find a pair w, b so that wx
i
b 1 for
the points with class label z
i
1 and wx
i
b 1 for the points with class label
z
i
1. To compute the hyperplane with the largest margin, we search two support-
ing hyperplanes for the two classes. The support planes are pushed apart until they
fall into a specific number of data vectors from each class (the support vectors
marked with a circle in Fig. 9.9). Thus, the solution depends only on these support
vectors. The distance between the supporting hyperplanes wx b 1 and
324
9 Analysis of Gene Expression Data
Fig. 9.9 Support vector machines. Two classes
of patient data are separated in the plane that is
spanned by the expression levels according to
two genes. Hyperplanes 1 and 2 yield two perfect
linear separations of the groups; however, when
classifying a new patient, they disagree in the
classification. Hyperplane 1 assigns the patient
to group 1 (circles), whereas hyperplane 2 as-
signs the patient to group 2 (squares) due to the
fact that both hyperplanes are geometrically too
close to one of the subsets. The support vector
machine classifier tries to maximize the margin
between the two groups by defining support vec-
tors (circled data points) and a hyperplane that
maximizes the minimum distances to these sup-
port vectors (hyperplane 3).

wx b 1 is equal to
2
w
kk
, where :
kk
denotes the Euclidean norm. Thus, maxi-
mizing the margin is equivalent to the following problem:
Minimize ||w||
2
subject to z
i
x
i
w b1 for i = 1, …, n. (9-26)
This problem can be represented by the Langrangian dual problem:
Minimize a values
1
2
P
i;j
z
i
z
j
a
i
a
j
x
i
x
j
P
i
a
i
subject to
P
i
z
i
a
i
0 and a
i
0 : (9-27)
Both problems lead to the same solution, i. e., a solution with the property that
w
P
i
z
i
a
i
x
i
. The classification rule found by the algorithm then reads for any new
datum
F
T
xsign
P
i
z
i
a
i
x
i
x b
: (9-28)
It should be noted that this classification rule depends only on the support vectors
since the dual problem assigns values a
i
= 0 to all other data vectors.
In practice, it might be that in the original dimension, p, no linear separation can
be performed on the training data. SVMs then map the data to a higher-dimensional
space where a linear separation is possible, using a map F : <
p
!<
m
; m > p. Since
the optimization problem involves only inner products of the vectors, an optimal se-
parating hyperplane in the projected space can be found by solving the problem for
the inner products F x
i
F x
j
. Fortunately (due to Mercer’s theorem, which is be-
yond the scope of this book), it is known that for certain mappings and any two vec-
tors the inner product in the projected dimension can be calculated using a kernel
function K:<
p
<
p
!<such that
K x
i
; x
j
F x
i
F x
j
: (9-29)
Two kernels are widely used:
. linear kernel: K x
i
; x
j
x
i
x
j
and
. polynomial kernel: K x
i
; x
j
g x
i
x
j
e
d
.
A further modification resulting from the fact that, in practice, misclassifications
will occur is the introduction of an error parameter, C, in the optimization problem
(Eq. (9-24)) and the dual problem (Eq. (9-25)). This error parameter represents the
tradeoff between the training set misclassification error and the size of the margin.
The optimization problem involving the kernel and this parameter reads
325
9.5 Classification Methods

Minimize a values
1
2
P
i;j
z
i
z
j
a
i
a
j
K x
i
x
j
P
i
a
i
subject to
P
i
z
i
a
i
0 and C a
i
0 ; (9-30)
and the classification is based on
F
T
xsign
P
i
z
i
a
i
K x
i
xb
: (9-31)
Further reading on SVMs can be found in Cristianini and Shawe-Taylor (2000).
SVMs seem to be the method of choice for classifying samples according to gene ex-
pression profiles. They have been proven to outperform other procedures, in particu-
lar in cancer diagnosis, by several independent studies (e.g., Furey et al. 2000; Statni-
kov et al. 2004).
9.5.3
Other Approaches
A very simple classification approach is the k-nearest neighbor classification method
(Duda and Hart 1973). Consider a training dataset of n pairs, (x
i
, z
i
), i = 1, …, n,ofp-
dimensional expression profiles and group labels. If a query, x, is to be classified,
then it is likely that the group label of x equals the group label of the most similar
training datum. Thus, the classification rule is given by
F
T
xz
i
0
where i
0
2 arg max fS x; x
i
; x
i
2 Tg; (9-32)
where S is a suitable similarity function between the expression profiles, for exam-
ple, the Pearson correlation coefficient. Taking into account the high error rates in
microarray experiments, it is not reasonable to base the classification on just the
nearest neighbor of the query in the training set but rather on the k-nearest neigh-
bors. Thus, the result of the classification of the query is defined as the majority vote
of these k data vectors. Further refinements are performed by weighting the group
labels of the data vectors according to their similarity to the query. K-nearest neigh-
bor methods yield surprisingly good results in many classification procedures and
can be used as a borderline test for more sophisticated algorithms.
Classification can also be combined with clustering methods using clustering-based
classification (Alon et al. 1999; Ben-Dor et al. 2000). If we consider the n training sam-
ples (tumor subtypes, cell lines etc.) as expression vectors whose coordinates are the ex-
pression levels of some genes, i.e., essentially transposing the expression matrix dis-
cussed in Section 9.3, then we can perform a clustering of the training sample in two
or more clusters. This yields groups of samples that are similar to each other based on
the selected set of genes. Clustering-based classification methods simply cluster the
query sample together with the training samples and assign the label of the highest
confidence calculated from all training labels in the same cluster to the query.
A third group of algorithms is based on boosting (Freund and Shapire 1996). The
idea of boosting is to construct a good classifier by repeated calls of weak learning
326
9 Analysis of Gene Expression Data

procedures. An example for a boosting algorithm is the AdaBoost algorithm by
Freund and Shapire.
9.5.4
Cross-validation
Methods of classification rely heavily on the use of the specific training set. It is,
however, important to get an idea of how well the learning procedure would perform
when asking for new predictions for unknown data. A method of choice to answer
this question is cross-validation. Cross-validation is a model evaluation method. The
basic idea of cross-validation is to divide the entire dataset in a training and a test
set. Some of the data is removed before training begins. Then, the data that was re-
moved can be used to test the performance of the classification method on poten-
tially “new” data. Several strategies involving this basic idea are detailed below.
9.5.4.1 The Holdout Method
In this method, the dataset is separated into two sets, called the training set and the
test set. A classifier is computed using the training set only. Then the classifier is
asked to predict the output values for the data in the test set. The errors are accumu-
lated as before to give the mean absolute test set error, which is used to evaluate the
method. It should be noted that holdout methods are very dependent on the holdout
test set and thus that evaluation can have a high variance.
9.5.4.2 k-fold Cross-validation
Here, the dataset is divided into k subsets, and the holdout method is repeated k
times. Each time, one of the k subsets is used as the test set and the other k–1 sub-
sets are put together to form a training set. Then the average error across all k trials
is computed. The advantage of this method is that it matters less how the data gets
divided. Every data point gets to be in a test set exactly once and gets to be in a train-
ing set k–1 times. The variance of the resulting estimate is reduced as k is increased.
A variant of this method is to randomly divide the data into a test and training set k
different times. The advantage of doing this is that you can independently choose
how large each test set is and how many trials you average over.
9.5.4.3 Leave-one-out Cross-validation
This method is k-fold cross-validation, with k equal to n, the number of data points
in the set. That means that n separate times, the function approximator is trained on
all the data except for one point and a prediction is made for that point. As before,
the average error is computed and used to evaluate the model. The evaluation given
by leave-one-out cross-validation error is very expensive to compute, but it has been
applied as a standard cross-validation method to classification studies.
327
9.5 Classification Methods

9.6
Reverse Engineering Genetic Networks
The methods discussed so far analyze the transcriptome level of genes in an explora-
tive way. Co-expressed genes might have similar regulatory characteristics, but it is
not possible to get information about the nature of the regulation. This task is
tackled by the analysis of genetic networks.
Genetic networks are composed of a set of molecular entities (genes, proteins,
compounds) and interactions between those entities. The purpose of these interac-
tions is to carry out some cellular functions. The dynamics of a genetic network de-
scribe functional pathways of a cell (or tissue) such as metabolism, gene regulation,
and signaling.
The task of reverse engineering of a genetic network is the reconstruction of the
interactions in a qualitative way from experimental data using algorithms that
weight the nature of the possible interactions with numerical values (cf. Section 3.5).
In contrast to the forward modeling of networks with known interactions, which
tries to determine network behavior by topological and dynamical characteristics (cf.
Chapter 8), reverse engineering is data-based and tries to estimate regulatory interac-
tions from the experimental data. Once determined, these networks can be used to
make predictions on the gene expression of the corresponding genes. In silico predic-
tions can be used to characterize vital functions of this network, e.g., predicting the
behavior of the network when knocking down a certain gene by a suitable model sys-
tem. In Section 9.6.1 we describe the simplest gene regulatory network model, the
Boolean network. Here, algorithms exist that can infer the Boolean rules from the
state transitions, which is exemplified with the REVEAL algorithm (Liang et al.
1999). In Section 9.6.2 we review alternative approaches, and in Section 9.6.3 we re-
port on recent findings that support the occurrence of specific modules in gene regu-
lation – the identification of common network motifs.
9.6.1
Reconstructing Boolean Networks
Consider a set of experimental conditions. An interesting question is whether we
can reconstruct the qualitative interactions of the corresponding genes. Obviously,
the general answer is no, because the set of experimental conditions might be too
general. However, there are two setups where we might succeed: time-dependent
measurements and knockout experiments. In time-dependent measurements, the
conditions are dependent on each other in the sense that (theoretically) a strong ex-
pression of a transcription factor at a certain time point will lead to activation (or re-
pression) of the gene expression of its targets at the next time point. Similarly,
knocking out the transcription factor should point to expression changes of its tar-
gets.
The simplest models of gene regulatory networks are Boolean models (see Sec-
tion 3.5 and Section 8.2). Here, genes have only discrete states and the regulatory in-
teractions are described by Boolean functions (Kauffman 1993). Quite a number of
328
9 Analysis of Gene Expression Data
reverse engineering algorithms for Boolean networks have been proposed (Akutsu
et al. 1999, 2000; Liang et al. 1999). Recall that in a Boolean network, genes are re-
presented as nodes, some of which are connected by edges with weights equal to 1
or –1 depending on whether the nature of regulation is activation or repression, re-
spectively.
Reverse engineering methods are based on the following principle idea. Consider
a pair of consecutive time-dependent conditions (time t and t +1)onn gene probes.
Binarize the expression values and define a set of rules that allows the computation
of binarized expression levels at time t + 1 from those at time t. These rules repre-
sent the expression changes occurring from the consecutive time points. A second
pair of consecutive time points will generate a second set of rules that may overlap
with those rules reconstructed from the first pair. This procedure iterates over all
pairs of consecutive time points, and the resulting set of rules are those that are con-
sistent to explain the whole dataset.
One of the first algorithms to reconstruct a Boolean network was REVEAL (Liang
et al. 1999). The basic idea of REVEAL is to use mutual information measurement
from the state transition tables to extract the regulatory Boolean rules (for defini-
tions of the concepts of entropy and mutual information, see Section 11.3). Recall
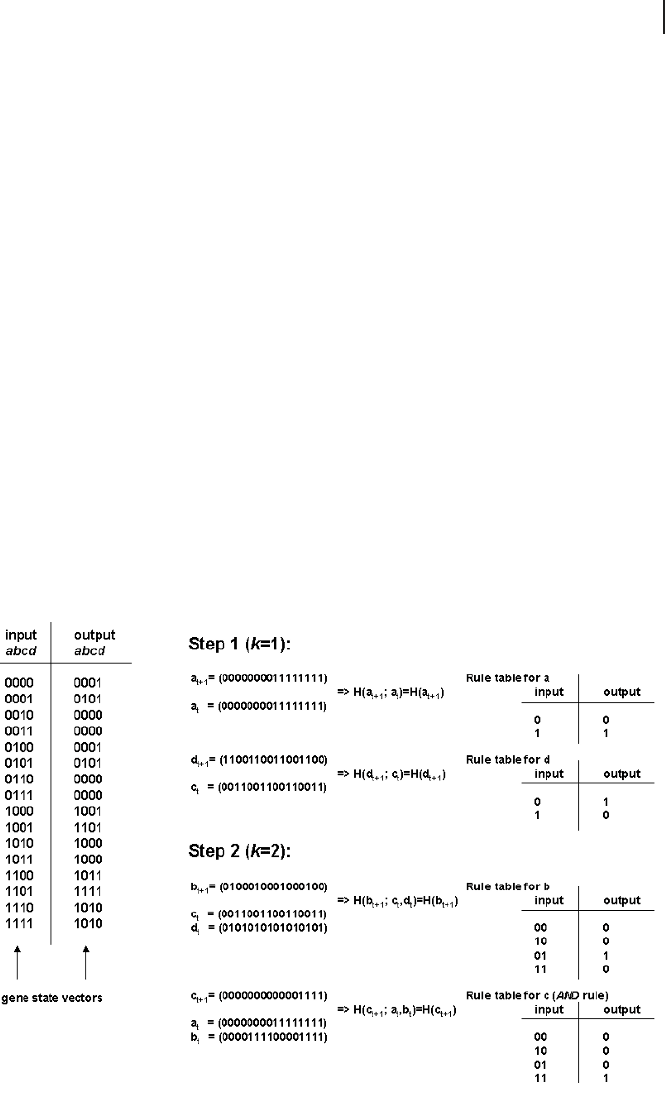
the state transition table (Tab. 8.1) for the four-gene network described in Fig. 8.2.
Denote by a
t
, b
t
, c
t
, d
t
the state vector of the genes across the 16 possible system
states at time t. Figure 9.10 describes the individual steps of the algorithm. A criti-
cal parameter of the algorithm is the connectivity, k, i. e., the number of inputs for
each gene.
329
9.6 Reverse Engineering Genetic Networks
Fig. 9.10 Analysis of a Boolean gene regulatory network with the REVEAL algorithm.
