Vaccari D.A., Strom P.F., Alleman J.E. Environmental Biology for Engineers and Scientists
Подождите немного. Документ загружается.

DNA can be specified either by any single nucleotide or by the sum of either A þ TorGþ
C. By convention, % G þ C is used as an indication of genetic relatedness. Organisms
with very different % G þ C compositions cannot be closely related. On the other
hand, % G þ C does not indicate the specific sequences of the bases, so that organisms
with similar % G þ C values are not necessarily related.
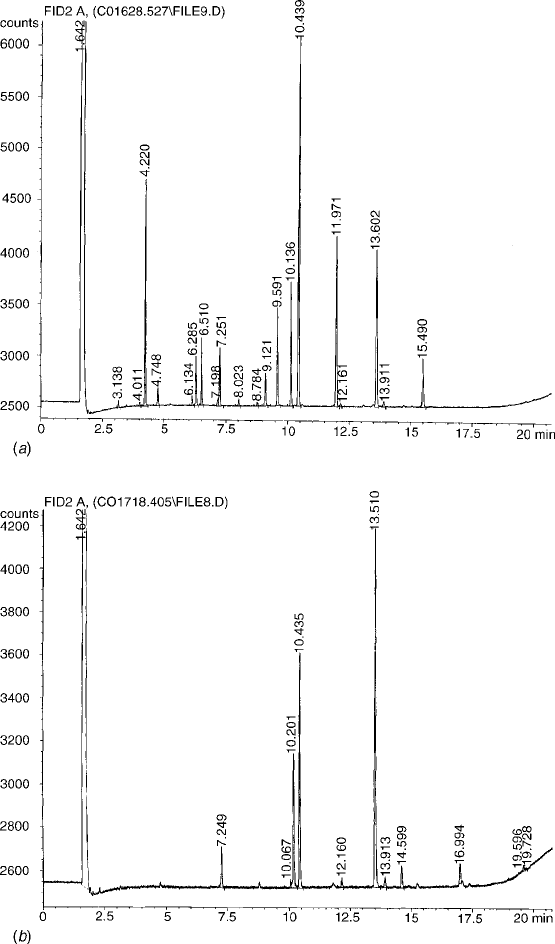
Figure 10.17 Fatty acid methyl ester (FAME) profiles showing different patterns for (a) Serratia
marcescens and (b) Tsukamurella paurometabolum. (Courtesy of Michael Fleming.)
238
MICROBIAL GROUPS
DNA Hybridization One means of genotyping cells is by comparing the similarity of
sequences of bases in their DNA. To compare two organisms, the DNA of one is first
labeled (often with radioactive
32
P). The DNA extracted from both organisms is then bro-
ken mechanically into small fragments and heated to separat e the two complemen tary
strands (denatured). The labeled DNA and an excess of the unlabeled DNA are next
mixed together and cooled to allow them to reanneal (re-form a double strand). The
extent to which the two different DNAs are able to combine with each other, or hybridize,
is a measure of how similar the base sequences are. This can be determined by the amount
of radioactivity present in the reannealed DNA.
Molecular Probes A large number of molecular probing techniques are now available or
under development. One group of methods of particular interest is referred to as fluores-
cent in situ hybridization (FISH). In a generalized FISH approach, an oligonucleotide
(short chain of nucleotides, typically 15 to 25 bases for FISH) containing a sequence char-
acteristic of a particular taxon of interest is labeled with a fluorescent dye. The labeled
oligonucleotide is then allowed to hybridize with the complementary sequence in the
organism in situ. With fluorescence microscopy, it is then possible to see the numbers
and location of the brightly colored target organism within a natural community. Depend-
ing on the degree of specificity of the sequence chosen, this approach potentially can be
used to identify organisms belonging to any taxonomic level ranging from domain (e.g.,
Bacteria) to a particular species or even strain.
Another type of probe is used in ribotyping. Restriction enzymes break DNA at points
where specific sequences of nucleotides occur. If DNA is treated with one or more such
enzymes, fragments of DNA are produced. These fragments are then separated by size on
a gel and probed with 16S rRNA genes. For treatme nt with a particular set of restriction
enzymes, the resulting pattern of DNA fragment sizes will be characteristic of the organ-
ism.
Denaturing gradient gel electrophoresis (DGGE) analysis offers a characterization of
the GC composition of DNA segments coding for rRNA or other genes. Following tagging
and amplification using PCR (Section 6.3.3), these fragments are drawn electrophoreti-
cally (i.e., attracted by their electrical charge) through gel tracks laden with a gradient
matrix of a denaturing agent (e.g., urea ranging from 30 to 55%). Since the G–C triple
bond is stronger than the A–T double bond, it does not denature until farther along in the
gradient. Fully denatured fragments stop migrating, resulting in a linear separation of the
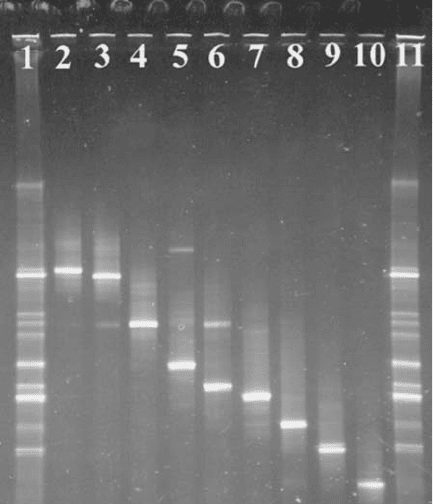
fragments into a banding pattern that is characteristic of the organism (Figure 10.18). A
similar approach is used in thermal gradient gel electrophoresis (TGGE), except that a
temperature gradient is used for denaturing the DNA.
16S rRNA Analysis The primary tool used today in determining phylogeny is 16S
rRNA analysis, developed largely by Carl Woese beginning in the early 1970s. Prokaryo-
tic ribosomal RNA (rRNA) is composed of three molecules, referred to by their sedimen-
tation coefficients measured in Svedberg units (S) as 5S (containing 120 bases), 16S
(1500), and 23S (2900). Woese initially worked with 5S fragments but found they
were too small to yield sufficient information. The emphasis then shifted to 16S rRNA
(and the comparable 18S rRNA found in eukaryotes) based on its larger, yet still manage-
able number of nucleotides.
One important advantage of using the nucleotide sequences of 16S (or 18S) rRNA for
determining relationships is that these molecules occur in all organisms. Also, because of
MICROBIAL TAXONOMY 239
their central role in the process of gene expression, there are strong selective pressures,
making it unlikely that they will undergo rapid changes. This leads to the presence of
extended regions of the molecule that are highly conserved (little changed) and which
therefore are useful for establishing distant phylogenetic relationships. On the other
hand, they also have an adequate number of variable regions that can be used to examine
closer relationships. Thus, 16S rRNA can be used as an evolutionary clock, or chron-
ometer, to measure the phylogenetic distance between taxa at a variety of levels based
on the number of nucleotide differences—representing stable mutations—that have
occurred over time.
One method to sequence 16S rRNA begins with extraction of a cell’s RNA. A small
DNA oligonucleotide primer (15 to 20 nucleotides in length) that is complementary in its
base sequence to a conserved region of the 16S rRNA molecule is added. Reverse tran-
scriptase can then be used to generate complemen tary DNA (cDNA), which in turn is
amplified using PCR. The nucleotide sequences of these cDNA are then determined,
and the original sequence of the rRNA is deduced from them. Another common approach
is to extract and sequence the DNA of the gene that codes for the 16S rRNA rather than
the RNA itself.
The 16S rR NA sequences of thousands of species have now been determin ed. Using
advanced computer algorithms, short (6 to 10 nucleotides) signature sequences have
been found that are highly consistent within groups of organisms. Also, phylogenetic
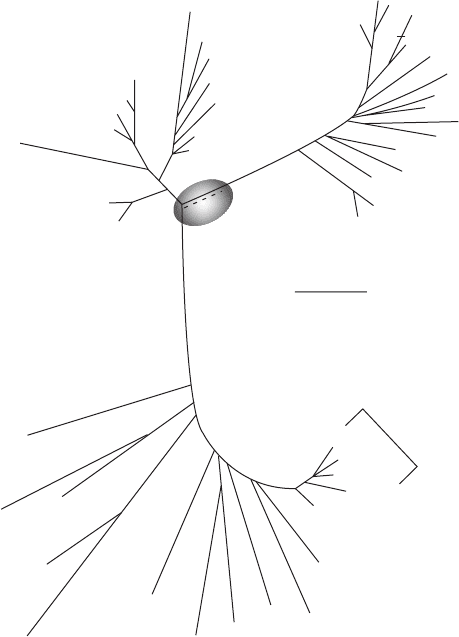
trees can be generated in which the evolutionary distance between two groups is indicated
by the length of the connecting lines (Figure 10.19).
Figure 10.18 Denaturing gradient gel extraction track profiles.
240
MICROBIAL GROUPS
10.5 BACTERIA
As a group, the domain Bacteria is extremely diverse, including phototrophs and chemo-
trophs, organotrophs and lithotrophs, heterotrophs and autotrophs, aerobes and anaerobes,
psychrophiles and mesophiles and therm ophiles and hyperthermophiles, halophiles and
nonhalophiles, acidophiles and neutrophiles and alkaliphiles, saprophytes and parasites.
They are able to utilize a vast array of organic compounds (i.e., all naturally occurring
and almost all synthetic organics) as carbon and energy sources, many reduced inorganics
as electron donors (for energy), and many oxidized inorganics as electron acceptors.
One way of discussing microorganisms is by function, grouping together all of those
with a particular ability or characteristic (e.g., phototrophs). However, sometimes organ-
isms that have similar functions are actually quite different from each other. (This is
origin
pJP 27
pJP 78
Marine
group I
Pyrodictium
Thermoproteus
Sulfolobus
Thermofilum
Methanopyrus
Thermococcus
Methanococcus
Methanobacterium
Methanosarcino
Methanospirillum
Holoferax
Chramatium
E. coli
Riftia symbiont
Agrobacterium
Mitochondria
Chlorobium
Cytophaga
Epulopiscium
Bacillus
Synechococcu
s
Thermus
Thermomicrobium
Thermologa
Aquifex
Chloroplast
EM 17
0.1 changes per nuclcotide (nt)
EUKARYA
m
a
croscopic
organism
s
Zea
Homo
Coprinus
Paramecium
Porphyra
Dictyostelium
Entamoeba
Naegleria
Euglena
Tr
y
panosoma
Physarum
Encephalitozoon
Vairimorpha
Hexamita
Giardia
Tritrichomonas
ARCHAEA
BACTERIA
Figure 10.19 Phylogenetic tree indicating evolutionary branching and distance between groups
based on on rRNA analysis. Fungi are represented by Coprinus (a mushroom), plants by Zea (corn),
and animals by Homo (humans). (From Atlas, 1997; all rights reserved.)
BACTE RIA 241
referred to as convergent evolution, since they have approached a similar form from dif-
ferent starting points.) Thus, microbiologists generally prefer, where possible, to group
bacteria based on phylogeny. That is the approach that is taken here, for the most part,
although it should be recognized that with our growing knowledge, these groupings con-
tinue to change.
Figure 10.19 showed the phylogeny of Bacteria using an approach based mainly on
16S rRNA analysis. The domain was broken into about eight known major groupings
(although other 16S rRNA sequences have been found in environmental samples, indicat-
ing that additional ‘‘unknown’’ groups also exist). What should these major groupings of
Bacteria (and Archaea) be called?
Previously, they may have been referred to as phyla, or divisions (reflecting bacterio-
logy’s roots in botany), or commonly, groups (reflecting phenotypic similarities rather
than phylogenetic relationships). However, in looking at phylogenetic trees, such as
that in Figure 10.19, som e microbiologists have come to believe that if the differences
between plants and animals (and fungi) warrant assignment to groupings at the level of
kingdoms, the much greater differences among bacteria should also.
Of course, not all microbiologists agree, and this has not yet been widely accepted
among other biologists, who traditionally have focused on Eukarya, and particularly,
plants and animals. One argument is that 16S (or 18S for eukaryotes) rRNA analysis is
not necessarily the ‘‘one true’’ measure of phylogeny and degree of evolutionary diver-
gence. Still, as more is learned about microbial diversity using a variety of tools, it seems
quite possible that some cat egorization scheme like this eventually will be accepted. Thus
for Chapters 10 through 13, where microbiology is emphasized, these groupings will be
referred to as ‘‘Kingdoms,’’ even though elsewhere we do not make this distinction.
Table 10.3 shows 14 major groupings of Bacteria and indicates some further subdivi-
sions into classes and orders. The newest (second) edition of Bergey’s Manual of Systema-
tic Bacteriology decided not to refer to the major groups as kingdoms, and instead, listed
23 phyla. (The term domain for Bacteria was retained; there had been a proposal to use
‘‘empire’’ instead.) The table also includes a number of representative, interesting, and
environmentally important genera. Comparison with the phylogenetic tree of
Figure 10.19 shows many similarities, but also some differences reflecting the changing
taxonomy of bacteria.
Some of the kingdoms have few genera, perhaps none of which are of known major
importance in environmental engineering and science. Other kingdoms have numerous
genera of great relevance to humans. The three largest are Proteobacteria, Firmicutes
(gram positives), and Cyanobacteria. It should also be noted that a number of bacteria
(generally not shown in the table) do not appear to fit in any of the kingdoms and are
thus of ‘‘uncertain’’ affiliation. (One extreme example is the genus Chrysiogenes, contain-
ing only one recognized species, which may merit its own kingdom!) Remember, these
divisions are based on similarities in the genetic code, not on any attempt to distribute
species equally among groups. One way to look at this unevenness (which also occurs
among plants and animals) is to realize that some approache s to survival lend themselves
to greater diversification because of the range of niches available.
It should also be recognized that our knowledge of bacterial groups is uneven. Natu-
rally, because more effort has gone into their study, we tend to know more about groups
that appear to be of greater importance to us. This includes bacteria used comme rcially, as
well as pathogens of humans, domestic animals, and crop plants. On the other hand, it is
not unusual to find that an organism isolated from the natural environment—or even a
242 MICROBIAL GROUPS

TABLE 10.3 The Bacteria: A Proposed Phylogeny Including
Some Representative, Interesting, and Environmentally
Important Genera
a
Kingdom 1. Aquaficae
Class 1. Aquaficae
Aquifex
Class 2. Thermotogae
Thermotoga
Class 3. Thermodesulfobacteria
Thermodesulfobacterium
Kingdom 2. Xenobacteria
Class 1. Deinococci
Deinococcus
Class 2. Thermi
Thermus
Nitrospira (or perhaps a separate Kingdom)
Leptospirillum
Magnetobacterium
Kingdom 3. Thermomicrobia
Class 1. Chloroflexi—green nonsulfur bacteria
Order 1. Chloroflexales
Chloroflexus
Order 2. Herpetosiphonales
Herpetosiphon
Class 2. Thermomicrobia
Thermomicrobium
Kingdom 4. Cyanobacteria—blue-green bacteria
(formerly, blue-green algae)
Class 1. Prochlorophyta
Order 1. Chroococcales
Chroococcus
Prochloron
Synechococcus
Order 2. Pleurocapsales
Pleurocapsa
Order 3. Oscillatoriales
Lyngbya
Oscillatoria
Spirulina
Order 4. Nostocales
Anabaena
Calothrix
Nostoc
Order 5. Stigonematales
Stigonema
Kingdom 5. Chlorobia—green sulfur bacteria
Chlorobium
Kingdom 6. Proteobacteria
b
Class 1. a-Proteobacteria (Rhodospirilli)
Class 2. b-Proteobacteria (Neisseriae)
(Continued )
BACTE RIA 243

sewage treatment plant—is not a previously known, or at least not well described, species.
Also, our knowledge of bacteria has been limited by our ability to culture (grow) them in
the laboratory; those that are not readily grown on laboratory media have been difficult to
study. Interestingly, we do know a fair amount about thermophilic bacteria. This is at least
in part because of the early efforts of Thomas D. Brock, who began studying hot springs
with the idea that the extreme conditions limited the microbia l diversity, making microbial
ecology studies simpler!
TABLE 10.3 (Continued )
Class 3. g-Proteobacteria (Zymobacteria)
Class 4. d-Proteobacteria (Predibacteria)
Class 5. e-Proteobacteria (Campylobacteres)
Kingdom 7. Firmicutes—Gram positives
b
Class 1. Clostridia
Class 2. Mollicutes
Class 3. Bacilli
Class 4. Actinobacteria
Kingdom 8. Planctomycetacia
Class 1. Planctomycetacia
Order 1. Planctomycetales
Planctomyces
Order 2. Chlamydiales
Chlamydia
Kingdom 9. Spirochetes
Borrelia
Leptospira
Spirochaeta
Treponema
Kingdom 10. Fibrobacteres
Fibrobacter
Kingdom 11. Bacteroidetes
Bacteroides
Kingdom 12. Flavobacteria
Flavobacterium
Kingdom 13. Sphingobacteria
Cytophaga
Flexibacter
Haliscomenobacter
Saprospira
Kingdom 14. Fusobacteria
Fusobacterium
Streptobacillus
Kingdom 15. Verrucomicrobia
Prosthecobacter
a
All classes listed if more than one. All orders listed if more than
one except for the two phyla Proteobacteria and Firmicutes. Families not
shown. (Based in part on outline available from Bergey’s Manual Trust.)
b
Large important group; additional breakdown and genera given in Tables
10.5 (Proteobacteria) and 10.7 (Firmicutes).
244 MICROBIAL GROUPS
Phylogenetic grouping, as in Table 10.3 (and Figure 10.19), does not always lend itself
to neat phenotypic groupings of organisms with similar characteristics. Still, we mainly
use this organization below for a brief description of the Bacteria. No attempt is made to
cover every subgroup or genus, and many details are omitted.
10.5.1 Aquaficae
This kingdom, which is considered to be deeply rooted (close to the original bacterial
forms), is so diverse that there are proposals to break it into three (or alternatively, to
lump it with the next). The known species are thermophilic or hyperthermophilic. Aquifex
is the most thermophilic known true bacteria (optimum 85
C, maximum 95
C) and is a
chemolithotrophic autotroph (growing on H
2
,S
0
,orS
2
O
2
3
). It requires oxygen or nitrate
as an elect ron acceptor but cannot tolerate high O
2
concentrations. Thermotoga is also a
hyperthermophile but is a fermentative chemoorganotroph. Thermodesulfobact erium is a
thermophilic sulfate reducer using fermentation products such as lactate and pyruvate as
its carbon and energy source.
Although they are undoubtedly of significance in the environments in which they are
found (mainly hot springs and marine thermal vents), these organisms are generally not of
concern to environmental engineers and scientists. However, their unique features make
them of special interest to microbiologists studying the evolution of life or biochemical
diversity.
10.5.2 Xenobacteria
This kingdom contains two classes, Deinoc occi and Thermi. Deinococcus radiodurans
was first isolated from food that had been irradiated to sterilize it. It is extraordinarily
radiation resistant, able to withstand more than 1.5 million rad of gamma irradiation
(3000 times the lethal dose to humans). Many deinococci, which are frequently reddish
in color due to caretenoid pigments, also are resistant to ultraviolet radiation and other
mutagens and are able to tolerate drying. Their resistance to genetic damage appears to
result in large part from an exceptional ability to repair damaged DNA rapidly. It is hoped
that these bacteria may prove useful in the future for certain types of hazardous waste site
cleanups.
Thermus aquaticus, first found in hot springs in Yellowstone National Park by a group
led by Thomas Brock, was one of the first extreme thermophilic organisms isolated. It has
also been found in hot-water heaters. Its heat-stable Taq DNA polymer ase is the enzyme
commonly used in PCR (Section 6.3.3).
Nitrospira is an autotroph that derives energy from the oxidation of nitrite to nitrate.
Thus, it is often grouped with the other nitrifying bacteria (Proteobacteria; see
Section 10.5.6). However, it also has been proposed to include it in the class Thermi,
or perhaps in its own kingdom, based on phylogeny.
10.5.3 Thermomicrobia (Including Green Nonsulfur Bacteria)
Although Thermomicrobium is a thermophilic (75
C optimum) chemoorganotroph, most
of the known members of this small kingdom are phototrophic green nonsulfur bacteria.
Chloroflexus is an anoxygenic (non-oxygen producing) phototroph that can grow autotro-
phically but grows better as a photoheterotroph (using organic carbon sources). It also can
BACTE RIA 245
grow aerobically as a chemoorganotroph. It is thought to be descended from the earliest
phototrophs. Table 10.4 compares several characteristics of the various groups of photo-
trophic bacteria.
10.5.4 Cyanobacteria: Blue-Green Bacteria (Formerly, Blue-Green Algae)
The Cyanobacteria is a large, diverse, and environmentally important bacterial group. As
discussed in Section 10.1, their early forms probably were responsible for producing the
oxygen of Earth’s atmosphere, allowing the development of advanced eukaryotic life
forms. Their present relevance to environmental engineers and scientists includes their
role in the eutrophication of lakes and the production of tastes and odors in drinking
waters drawn from surface supplies.
Because they are oxygen-producing photoautotrophs, cyanobacteria were long consid-
ered to be algae (Cyanophyta). However, once it was realized that they were prokaryotes,
they were renamed and reclassified. It is now believed that they probably shared a com-
mon ancestor with chloroplasts. They are compared with the other phototrophic bacteria
in Table 10.4. Their green color comes from chlorophyll a, while the blue color comes
from other pigments called phycocyanins.
Most Cyanobacteria are strictly photoautotrophic, but a few can utilize simple organics
as carbon sources (photoheterotrophs) or even grow in the dark as chemoorganotrophs.
Many (e.g., Oscillatoria) have gliding motility. Although they are usually thought of as
aquatic freshwater organisms, they are also found in the ocean and in soils. A number can
withstand extreme environments, such as hot springs with temperatures of 70
C, the sur-
face of deser t soils, bare rock, saline lakes, and shallow tidal seas (where they may form
large mats). Different species grow as individual cells, filaments, or various types of
aggregates.
Many Cyanobacteria have the unusual ability to be able to fix nitrogen (convert N
2
to a
combined form, usually as ammonium or an amine compound). Interestingly, nitrogen-
ase, the enzyme responsible for nitrogen fixation, is sensitive to oxygen; this means that
nitrogen fixation cannot readily occur in the light (when oxygen is being produced) in
most species. However, some filamentous Cyanobacteria (e.g., Anabaena, Figure 10.20)
form differentiated cells, known as heterocysts, for nitrogen fixation. These special struc-
tures do not photosynthesize and have thickened walls to minimize oxygen diffusion from
the outside. They receive nutrition from neighboring cells in the filament, and in return
make fixed nitrogen available to them.
The five major groups of Cyanobacteria of Table 10.3 correspond to general morpho-
logical forms: Chroococcales, single cells or small aggregates; Pleurocapsales, small
spherical cells formed through fission (for repr oduction); Oscillatoriales, filaments;
Nostocales, filaments with heterocysts; Stigonematales, branched filaments. Except for
the Chroococcales, which are very diverse, these groupings also hold up fairly well to
phylogenetic analysis.
10.5.5 Chlorobia: Green Sulfur Bacteria
Another distinct group of phototrophs included in Table 10.4 is the green sulfur bacteria.
Like the green nonsulfur bacteria, they are anoxygenic, and can use hydrogen sulfide
(H
2
S) as their electron donor. The sulfide is oxidized first to elemental sulfur, which pro-
duces granules outside the cell, and then to sulfate (SO
2
4
). Also, they contain unique
246 MICROBIAL GROUPS

TABLE 10.4 Characteristics of Phototrophic Bacteria
Green Sulfur Green Nonsulfur Purple Sulfur Purple Nonsulfur Blue-Green Heliobacteria
Kingdom Chlorobia Thermomicrobia g-Proteobacteria a -orb-Proteo-
bacteria
Cyanobacteria Firmicutes
Light Obligately Usually Obligately Usually Aerobic, Obligately
metabolism anaerobic, photo
autotroph (some
also photo
heterotophic)
anaerobic,
photoheterotroph
or photoautotroph
anaerobic,
photoautotroph
anaerobic, photo-
heterotroph or
photoautotroph
photoautotroph anaerobic,
photoheterotroph
Electron
donor for
lithotrophy
H
2
,H
2
S, S H
2
,H
2
SH
2
,H
2
S, S H
2
,H
2
SH
2
O
Dark None Chemoorgano- None Chemoorgano- None Chemoorgano-
metabolism heterotroph heterotroph (some are
chemoorgano-
heterotrophs)
heterotroph
Carbon
source
CO
2
CO
2
or organic
carbon
CO
2
CO
2
or organic
carbon
CO
2
(some can use
organic carbon)
Organic carbon
Sulfur Outside cell None Within cell None None None
deposition
N
2
Fixation Yes No Yes Yes Yes (many) Yes
Endospores No No No No No Yes
Chlorophyll Bact.-a, and
c, d,ore
Bact.-c Bact.-a, b Bact.-a, b Chlor.-a Bact.-g
247
